

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148918.4 - phase: 0
(468 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
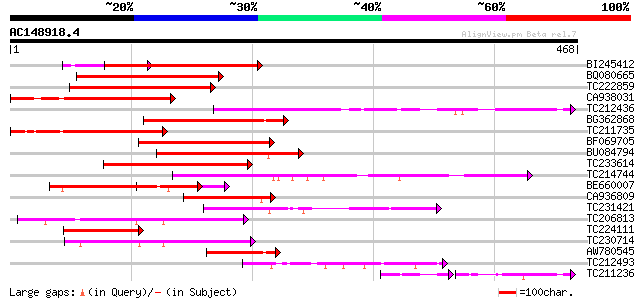
Score E
Sequences producing significant alignments: (bits) Value
BI245412 homologue to GP|29028906|gb At2g02080 {Arabidopsis thal... 265 2e-71
BQ080665 238 6e-63
TC222859 homologue to GB|AAO64832.1|29028906|BT005897 At2g02080 ... 233 2e-61
CA938031 218 6e-57
TC212436 218 6e-57
BG362868 homologue to PIR|T46147|T461 zinc finger protein - Arab... 212 3e-55
TC211735 208 5e-54
BF069705 similar to GP|16226322|gb AT3g45260/F18N11_20 {Arabidop... 201 4e-52
BU084794 191 6e-49
TC233614 186 2e-47
TC214744 homologue to UP|Q9LVQ7 (Q9LVQ7) Zinc finger protein, pa... 180 1e-45
BE660007 similar to GP|29028906|gb At2g02080 {Arabidopsis thalia... 148 6e-36
CA936809 homologue to PIR|T46147|T461 zinc finger protein - Arab... 139 2e-33
TC231421 homologue to GB|AAM10292.1|20147151|AY091693 AT3g13810/... 114 7e-26
TC206813 similar to UP|CW14_YEAST (O13547) Covalently-linked cel... 110 1e-24
TC224111 similar to UP|Q944L3 (Q944L3) AT3g45260/F18N11_20 (ID1-... 109 2e-24
TC230714 weakly similar to GB|AAO44076.1|28466935|BT004810 At5g2... 100 1e-21
AW780545 97 1e-20
TC212493 similar to GB|AAM10292.1|20147151|AY091693 AT3g13810/MC... 96 3e-20
TC211236 similar to PIR|S48856|S48856 finger protein pcp1 - pota... 72 2e-19
>BI245412 homologue to GP|29028906|gb At2g02080 {Arabidopsis thaliana},
partial (31%)
Length = 664
Score = 265 bits (678), Expect = 2e-71
Identities = 115/130 (88%), Positives = 126/130 (96%)
Frame = -1
Query: 79 ICNKGFQRDQNLQLHKRGHNLPWKLKQRTSNEIRKKVYVCPEPTCVHHDPSRALGDLTGI 138
+C KGFQR+QNLQLH+RGHNLPWKLKQ+T+ E ++KVY+CPEPTCVHHDPSRALGDLTGI
Sbjct: 547 VCTKGFQREQNLQLHRRGHNLPWKLKQKTTKEPKRKVYLCPEPTCVHHDPSRALGDLTGI 368
Query: 139 KKHFCRKHGEKKWKCDKCSKKYAVQSDWKAHSKTCGTREYRCDCGTLFSRRDSFITHRAF 198
KKH+ RKHGEKKWKCDKCSKKYAVQSDWKAHSKTCGTREYRCDCGTLFSRRDSFITHRAF
Sbjct: 367 KKHYSRKHGEKKWKCDKCSKKYAVQSDWKAHSKTCGTREYRCDCGTLFSRRDSFITHRAF 188
Query: 199 CDALAEESSR 208
CDALA+ES+R
Sbjct: 187 CDALAQESAR 158
Score = 49.3 bits (116), Expect = 4e-06
Identities = 30/75 (40%), Positives = 40/75 (53%)
Frame = -2
Query: 44 PAKKRRNLPGNPDPQAEVIALSPKTLMATNRFICEICNKGFQRDQNLQLHKRGHNLPWKL 103
P +KR +L N AEVIALSPKTLMATNRFIC KG + + + L
Sbjct: 645 PKEKRTHL--NTISNAEVIALSPKTLMATNRFICRCAPKGSKESKTYSYTEEDTTCLGSL 472
Query: 104 KQRTSNEIRKKVYVC 118
+R R++ ++C
Sbjct: 471 SRRLQKSQRER-FIC 430
>BQ080665
Length = 425
Score = 238 bits (606), Expect = 6e-63
Identities = 103/121 (85%), Positives = 115/121 (94%)
Frame = -2
Query: 56 DPQAEVIALSPKTLMATNRFICEICNKGFQRDQNLQLHKRGHNLPWKLKQRTSNEIRKKV 115
DP AEVIALSP TL+ATNRF+CEICNKGFQRDQNLQLH+RGHNLPWKLK RT+ ++RK+V
Sbjct: 364 DPSAEVIALSPNTLVATNRFVCEICNKGFQRDQNLQLHRRGHNLPWKLKLRTTTDVRKRV 185
Query: 116 YVCPEPTCVHHDPSRALGDLTGIKKHFCRKHGEKKWKCDKCSKKYAVQSDWKAHSKTCGT 175
YVCPEP+CVHH+P+RALGDLTGIKKHF RKHGEKKWKC+KCSKKYAVQSDWKAHSK CGT
Sbjct: 184 YVCPEPSCVHHNPARALGDLTGIKKHFSRKHGEKKWKCEKCSKKYAVQSDWKAHSKICGT 5
Query: 176 R 176
+
Sbjct: 4 K 2
>TC222859 homologue to GB|AAO64832.1|29028906|BT005897 At2g02080 {Arabidopsis
thaliana;} , partial (22%)
Length = 454
Score = 233 bits (593), Expect = 2e-61
Identities = 102/121 (84%), Positives = 113/121 (93%)
Frame = +2
Query: 50 NLPGNPDPQAEVIALSPKTLMATNRFICEICNKGFQRDQNLQLHKRGHNLPWKLKQRTSN 109
N+ DP AEVIALSPKTLMATNRF+CE+CNKGFQR+QNLQLH+RGHNLPWKLKQ+T+
Sbjct: 92 NICAYADPDAEVIALSPKTLMATNRFLCEVCNKGFQREQNLQLHRRGHNLPWKLKQKTNK 271
Query: 110 EIRKKVYVCPEPTCVHHDPSRALGDLTGIKKHFCRKHGEKKWKCDKCSKKYAVQSDWKAH 169
E ++KVY+CPEPTCVHHDPSRALGDLTGIKKH+ RKHGEKKWKCDKCSKKYAVQSDWKAH
Sbjct: 272 EPKRKVYLCPEPTCVHHDPSRALGDLTGIKKHYSRKHGEKKWKCDKCSKKYAVQSDWKAH 451
Query: 170 S 170
S
Sbjct: 452 S 454
>CA938031
Length = 416
Score = 218 bits (554), Expect = 6e-57
Identities = 107/137 (78%), Positives = 116/137 (84%)
Frame = +1
Query: 1 MSNLTSASGEASANSSGNRTHEVDAKFSQQYFASSQTQTHDETPAKKRRNLPGNPDPQAE 60
MSNLTSASGEASA SSGNRT + Y A +QT +KK+RNLPGNPDP AE
Sbjct: 28 MSNLTSASGEASAASSGNRT-----EIGTSYMAPPPSQTQQ---SKKKRNLPGNPDPDAE 183
Query: 61 VIALSPKTLMATNRFICEICNKGFQRDQNLQLHKRGHNLPWKLKQRTSNEIRKKVYVCPE 120
VIALSPK+L+ATNRFICEICNKGFQRDQNLQLH+RGHNLPWKLKQRTS E+RKKVYVCPE
Sbjct: 184 VIALSPKSLLATNRFICEICNKGFQRDQNLQLHRRGHNLPWKLKQRTSKEVRKKVYVCPE 363
Query: 121 PTCVHHDPSRALGDLTG 137
P+CVHHDPSRALGDLTG
Sbjct: 364 PSCVHHDPSRALGDLTG 414
>TC212436
Length = 920
Score = 218 bits (554), Expect = 6e-57
Identities = 145/328 (44%), Positives = 170/328 (51%), Gaps = 29/328 (8%)
Frame = +2
Query: 169 HSKTCGTREYRCDCGTLFSRRDSFITHRAFCDALAEESSRTVIPQPTQPNSHHNMNNLQT 228
HSKTCGTR+YRCDCGTLFSRRDSFITHRAFCDALAEESSR+V NS
Sbjct: 2 HSKTCGTRDYRCDCGTLFSRRDSFITHRAFCDALAEESSRSVTGIGIVANSTSTQPTAAA 181
Query: 229 QDIQGFTLKKEHQSFNMLRPEQEVQIPSWLCQSSIDLSSNYSSLDQDLHLYENPNPRNGP 288
Q + +F++ + +Q P W+ Q S +S++ Q ENPNPR G
Sbjct: 182 ASHQQDIIHGNSNNFSLKKEQQAGFRPPWIGQPSPSSASSFLVSHQ-----ENPNPRGGG 346
Query: 289 TSTLPSYQPSSAASPHMSATALLQKAAQMGATSSCSSQSMMSGTH-QQGHVSIVDSATNN 347
P+ P +PHMSATALLQKA+QMGAT S M GTH QQ HVS
Sbjct: 347 PG--PTLLPPYQTAPHMSATALLQKASQMGAT--MSKTGSMIGTHQQQAHVS-------- 490
Query: 348 MINSNGNFSLNLSSCEDQM-----------------INNSFS-----------SSGFHGT 379
N +LNLSS + QM + N S SS F GT
Sbjct: 491 -----ANAALNLSSRDHQMTPTLHGLVPFGNKAVPAVGNGVSPSLLHHIIDSFSSPFEGT 655
Query: 380 SFEDTFAGNILHSNQDHNINHDGDNDIPKTTTNDDDVAAGGNNAFTRDFLGLKPLSDSDI 439
SFEDTF G G + + KTTT DD N A TRDFLGL+PLS +DI
Sbjct: 656 SFEDTFGG-------------AGGDAMTKTTTADDGARGNNNEALTRDFLGLRPLSHTDI 796
Query: 440 LTIAGMGSCMNPSNSNHQENHSQKPWEG 467
L IAGMGSC+N S Q N + PW+G
Sbjct: 797 LNIAGMGSCINSS----QHNQTPNPWQG 868
>BG362868 homologue to PIR|T46147|T461 zinc finger protein - Arabidopsis
thaliana, partial (21%)
Length = 482
Score = 212 bits (540), Expect = 3e-55
Identities = 97/120 (80%), Positives = 104/120 (85%)
Frame = +1
Query: 111 IRKKVYVCPEPTCVHHDPSRALGDLTGIKKHFCRKHGEKKWKCDKCSKKYAVQSDWKAHS 170
+RK+VYVCPEPTCVHHDPSRALGDLTGIKKHFCRKHGEKKWKCDKCSKKYAV SDWKAHS
Sbjct: 16 VRKRVYVCPEPTCVHHDPSRALGDLTGIKKHFCRKHGEKKWKCDKCSKKYAVHSDWKAHS 195
Query: 171 KTCGTREYRCDCGTLFSRRDSFITHRAFCDALAEESSRTVIPQPTQPNSHHNMNNLQTQD 230
K CGTREY+CDCGTLFSRRDSFITHRAFCDALAEES+R+ PQ S + + T D
Sbjct: 196 KICGTREYKCDCGTLFSRRDSFITHRAFCDALAEESARSQ-PQTVAKASSESDSKAVTGD 372
>TC211735
Length = 689
Score = 208 bits (529), Expect = 5e-54
Identities = 105/131 (80%), Positives = 115/131 (87%), Gaps = 1/131 (0%)
Frame = +1
Query: 1 MSNLTSASGEASANSSGNRTHEVDAKFSQQYFASSQTQTHDETPAKKRRNLPGNPDPQAE 60
MSNLTSASGEASA SSGNRT E+ +SQQYFA +Q P KK+RNLPGNPDP+AE
Sbjct: 304 MSNLTSASGEASA-SSGNRT-EIGTDYSQQYFAPPLSQAQPP-PLKKKRNLPGNPDPEAE 474
Query: 61 VIALSPKTLMATNRFICEICNKGFQRDQNLQLHKRGHNLPWKLKQRTSNE-IRKKVYVCP 119
V+ALSPKTL+ATNRFICEICNKGFQRDQNLQLH+RGHNLPWKLKQR+SNE IRKKVYVCP
Sbjct: 475 VVALSPKTLLATNRFICEICNKGFQRDQNLQLHRRGHNLPWKLKQRSSNEIIRKKVYVCP 654
Query: 120 EPTCVHHDPSR 130
E +CVHHDPSR
Sbjct: 655 EASCVHHDPSR 687
>BF069705 similar to GP|16226322|gb AT3g45260/F18N11_20 {Arabidopsis
thaliana}, partial (22%)
Length = 406
Score = 201 bits (512), Expect = 4e-52
Identities = 88/112 (78%), Positives = 99/112 (87%)
Frame = +1
Query: 107 TSNEIRKKVYVCPEPTCVHHDPSRALGDLTGIKKHFCRKHGEKKWKCDKCSKKYAVQSDW 166
++ E+RKKVY+CPE TCVHHD +RALGDLTGIKKH+ RKHGEKKWKC+KCSKKYAVQSDW
Sbjct: 4 SNKEVRKKVYICPEKTCVHHDAARALGDLTGIKKHYSRKHGEKKWKCEKCSKKYAVQSDW 183
Query: 167 KAHSKTCGTREYRCDCGTLFSRRDSFITHRAFCDALAEESSRTVIPQPTQPN 218
KAH+KTCGTREY+CDCG LFSR+DSFITHRAFCDALA+ESSR T N
Sbjct: 184 KAHTKTCGTREYKCDCGNLFSRKDSFITHRAFCDALADESSRLTSVASTSLN 339
>BU084794
Length = 420
Score = 191 bits (485), Expect = 6e-49
Identities = 89/125 (71%), Positives = 99/125 (79%), Gaps = 4/125 (3%)
Frame = +2
Query: 122 TCVHHDPSRALGDLTGIKKHFCRKHGEKKWKCDKCSKKYAVQSDWKAHSKTCGTREYRCD 181
+CVHHDPSRALGDLTGIKKH+ RKHGEKKWKCDKCSKKYAVQSDWKAHSK CGTREY+CD
Sbjct: 2 SCVHHDPSRALGDLTGIKKHYSRKHGEKKWKCDKCSKKYAVQSDWKAHSKICGTREYKCD 181
Query: 182 CGTLFSRRDSFITHRAFCDALAEESSR-TVIP---QPTQPNSHHNMNNLQTQDIQGFTLK 237
CGTLFSR+DSFITHRAFCDALAEES+R T +P + + HH++ N Q I
Sbjct: 182 CGTLFSRKDSFITHRAFCDALAEESARVTTVPAALSNLRNDHHHHLTNAQASRIPQINFS 361
Query: 238 KEHQS 242
H S
Sbjct: 362 GFHSS 376
>TC233614
Length = 460
Score = 186 bits (472), Expect = 2e-47
Identities = 80/124 (64%), Positives = 101/124 (80%), Gaps = 1/124 (0%)
Frame = +1
Query: 78 EICNKGFQRDQNLQLHKRGHNLPWKLKQRTSNEIRKKVYVCPEPTCVHHDPSRALGDLTG 137
EICN+GFQRDQNLQ+H+R H +PWKL +R + ++K+V+VCPEP+C+HHDP ALGDL G
Sbjct: 1 EICNQGFQRDQNLQMHRRRHKVPWKLLKRETPVVKKRVFVCPEPSCLHHDPCHALGDLVG 180
Query: 138 IKKHFCRKH-GEKKWKCDKCSKKYAVQSDWKAHSKTCGTREYRCDCGTLFSRRDSFITHR 196
IKKHF RKH K+W C++CSK YAVQSD+KAH KTCGTR + CDCG +FSR +SFI H+
Sbjct: 181 IKKHFRRKHNNHKQWVCERCSKGYAVQSDYKAHLKTCGTRGHSCDCGRVFSRVESFIEHQ 360
Query: 197 AFCD 200
C+
Sbjct: 361 DACN 372
>TC214744 homologue to UP|Q9LVQ7 (Q9LVQ7) Zinc finger protein, partial (19%)
Length = 1045
Score = 180 bits (457), Expect = 1e-45
Identities = 129/352 (36%), Positives = 167/352 (46%), Gaps = 55/352 (15%)
Frame = +1
Query: 135 LTGIKKHFCRKHGEKKWKCDKCSKKYAVQSDWKAHSKTCGTREYRCDCGTLFSRRDSFIT 194
LTGIKKHFCRKHGEKKWKCDKCSKKYAVQSDWKAHSK CGTREY+CDCGT+FSRRDSFIT
Sbjct: 1 LTGIKKHFCRKHGEKKWKCDKCSKKYAVQSDWKAHSKICGTREYKCDCGTIFSRRDSFIT 180
Query: 195 HRAFCDALAEESSRTVIPQPTQ--PNSH------------HNMNNLQTQDIQ--GFTLKK 238
HRAFCDALAEE+++ Q + PN N N++ Q G T +
Sbjct: 181 HRAFCDALAEENNKANEGQLPKIGPNLQCQQIPNLVSSLPINTNSIVPNPAQMGGTTSEF 360
Query: 239 EHQSFN--MLRPEQEVQIPSW------LCQSSIDLSSNYSSLDQDLHLYENPNPRNGPTS 290
H + P + + +P+ + ++ S S+ L L N NG
Sbjct: 361 NHADHKHPLSLPHELMPMPAQKPFNNNMAAGTVFTRSLSSTSSPSLQLSSNMFDENG--- 531
Query: 291 TLPSYQPSSAASPHMSATALLQKAAQMGAT------------------------------ 320
+A SPHMSATALLQKAAQMGAT
Sbjct: 532 -----LHLAAGSPHMSATALLQKAAQMGATLTEKSTFATNMAPPSFGVLQQHHHHQQPNG 696
Query: 321 SSCSSQSMMSG-THQQGHVSIVDSATNNMINSNGNFSLNLSSCEDQMINNSFSSSGFHGT 379
++Q M SG HQQ V+I + N G S+ ++ + F+
Sbjct: 697 QPFTNQYMHSGHHHQQQEVNISTQYSFGSANGMGGGSVGMNGVD-----------MFNAI 843
Query: 380 SFEDTFAGNILHSNQDHNINHDGDNDIPKTTTNDDDVAAGGNNAFTRDFLGL 431
+ I+ N + + + N + N+ + G + T DFLG+
Sbjct: 844 LDQSKALSKIIEQNNNRSSSGGTTNGGSSSAINNVAGSKGSGDVMTLDFLGI 999
>BE660007 similar to GP|29028906|gb At2g02080 {Arabidopsis thaliana}, partial
(25%)
Length = 667
Score = 148 bits (373), Expect = 6e-36
Identities = 76/135 (56%), Positives = 92/135 (67%), Gaps = 9/135 (6%)
Frame = +2
Query: 34 SSQTQTHDE--------TPAKKRRNLPGNPDPQAEVIALSPKTLMATNRFICEICNKGFQ 85
S Q TH + P KKRRN PG P P AEVI LSPKTLMATNRFICE+CNKGFQ
Sbjct: 197 SPQITTHHQPSTVSPTTAPQKKRRNQPGTPYPDAEVIKLSPKTLMATNRFICEVCNKGFQ 376
Query: 86 RDQNLQLHKRGHNLPWKLKQR-TSNEIRKKVYVCPEPTCVHHDPSRALGDLTGIKKHFCR 144
R+QNLQLH+RGHNLPWKLKQ+ T+ E ++KVY+CP P + L + + +
Sbjct: 377 REQNLQLHRRGHNLPWKLKQKSTTKEPKRKVYLCPSPP-ASIMILQGLWETSLGSRSLLS 553
Query: 145 KHGEKKWKCDKCSKK 159
+ EKKWKC+KCSK+
Sbjct: 554 QAXEKKWKCEKCSKE 598
Score = 45.4 bits (106), Expect = 5e-05
Identities = 26/80 (32%), Positives = 39/80 (48%), Gaps = 3/80 (3%)
Frame = +3
Query: 105 QRTSNEIRKKVYVCPEPTCVHHDPS---RALGDLTGIKKHFCRKHGEKKWKCDKCSKKYA 161
+R + +K+ ++C H PS + G ++ + RK + +KYA
Sbjct: 435 RRVQQKSQKERFICVR---AHLRPS*SFKGFGRPHWDQEAYYRKPARRNGSVKSAQRKYA 605
Query: 162 VQSDWKAHSKTCGTREYRCD 181
QSDWKAHSKT RE+ CD
Sbjct: 606 GQSDWKAHSKTLWPREFXCD 665
>CA936809 homologue to PIR|T46147|T461 zinc finger protein - Arabidopsis
thaliana, partial (14%)
Length = 423
Score = 139 bits (351), Expect = 2e-33
Identities = 63/78 (80%), Positives = 70/78 (88%), Gaps = 2/78 (2%)
Frame = +3
Query: 144 RKHGEKKWKCDKCSKKYAVQSDWKAHSKTCGTREYRCDCGTLFSRRDSFITHRAFCDALA 203
RKHGEKKWKCDKCSKKYAVQSDWKAHSK CGTREY+CDCGT+FSRRDSFITHRAFCDALA
Sbjct: 3 RKHGEKKWKCDKCSKKYAVQSDWKAHSKVCGTREYKCDCGTVFSRRDSFITHRAFCDALA 182
Query: 204 EES--SRTVIPQPTQPNS 219
EE+ S TV+ ++ +S
Sbjct: 183 EENFLSHTVVKDISENDS 236
>TC231421 homologue to GB|AAM10292.1|20147151|AY091693 AT3g13810/MCP4_2
{Arabidopsis thaliana;} , partial (12%)
Length = 1164
Score = 114 bits (286), Expect = 7e-26
Identities = 79/208 (37%), Positives = 104/208 (49%), Gaps = 12/208 (5%)
Frame = +3
Query: 161 AVQSDWKAHSKTCGTREYRCDCGTLFSRRDSFITHRAFCDALAEESSRTVIPQ------- 213
AVQSDWKAHSKTCGTREYRCDCGTLFSR+DSFITHRAFCDALAEES+R Q
Sbjct: 3 AVQSDWKAHSKTCGTREYRCDCGTLFSRKDSFITHRAFCDALAEESARLSANQLAAAAGA 182
Query: 214 ---PTQPNSHHNMNNLQTQDIQGFTLKKEHQ--SFNMLRPEQEVQIPSWLCQSSIDLSSN 268
T N ++ QTQ Q F + HQ SFN Q
Sbjct: 183 TVTTTTTNPFQSLYLFQTQQ-QNF---QNHQMTSFNQWDSSQ------------------ 296
Query: 269 YSSLDQDLHLYENPNPRNGPTSTLPSYQPSSAASPHMSATALLQKAAQMGATSSCSSQSM 328
ENPNP N +T +P S + + + ++ LQ+ Q G + +++ M
Sbjct: 297 -----------ENPNPSNNIATTSLHIKPESQSFHNPTLSSFLQQ-QQQGQQPNNNNRGM 440
Query: 329 MSGTHQQGHVSIVDSATNNMINSNGNFS 356
++ HV+ AT++ +++ S
Sbjct: 441 IASPFGNLHVAAAAPATSSYMSATALLS 524
>TC206813 similar to UP|CW14_YEAST (O13547) Covalently-linked cell wall
protein 14 precursor (Inner cell wall protein), partial
(6%)
Length = 1214
Score = 110 bits (276), Expect = 1e-24
Identities = 66/210 (31%), Positives = 101/210 (47%), Gaps = 19/210 (9%)
Frame = +1
Query: 7 ASGEASANSSGNRTHEVDAKFS-------QQYFASSQTQTHDETPAKKRRNLPGNPDPQA 59
+SG N+SGN+ + + +Q + + + DE A + NLP
Sbjct: 550 SSGIRPQNNSGNKLFDQSTQNDLPNKLEMEQNYNMEEHEPKDEEDADEGENLPPG---SY 720
Query: 60 EVIALSPKTLMATNRFICEICNKGFQRDQNLQLHKRGHNLPWKL--------KQRTSNEI 111
E++ L + ++A + C IC KGF+RD NL++H RGH +K K+ S
Sbjct: 721 EILQLEKEEILAPHTHFCTICGKGFKRDANLRMHMRGHGDKYKTPAALAKPHKETGSEPK 900
Query: 112 RKKVYVCPEPTCVH---HDPSRALGDLTGIKKHFCRKHGEKKWKCDKCS-KKYAVQSDWK 167
K Y CP C H + L + +K H+ R H +K + C +C+ KK++V +D K
Sbjct: 901 LIKRYSCPYAGCKRNKDHKKFQPLKTILCVKNHYKRTHCDKSYTCSRCNTKKFSVMADLK 1080
Query: 168 AHSKTCGTREYRCDCGTLFSRRDSFITHRA 197
H K CG ++ C CGT FSR+D H A
Sbjct: 1081THEKHCGKDKWLCSCGTTFSRKDKLFGHIA 1170
>TC224111 similar to UP|Q944L3 (Q944L3) AT3g45260/F18N11_20 (ID1-like zinc
finger protein 1), partial (17%)
Length = 688
Score = 109 bits (273), Expect = 2e-24
Identities = 47/66 (71%), Positives = 59/66 (89%)
Frame = +2
Query: 45 AKKRRNLPGNPDPQAEVIALSPKTLMATNRFICEICNKGFQRDQNLQLHKRGHNLPWKLK 104
AK+RR+LP PDP AEV+ALSPK LMATNRF+ E+C++G QRDQNLQLH+RGHNLPW+L+
Sbjct: 476 AKRRRSLPRTPDPDAEVVALSPKCLMATNRFLWEVCSEGVQRDQNLQLHRRGHNLPWRLR 655
Query: 105 QRTSNE 110
+RT N+
Sbjct: 656 KRTDND 673
>TC230714 weakly similar to GB|AAO44076.1|28466935|BT004810 At5g22890
{Arabidopsis thaliana;} , partial (58%)
Length = 1049
Score = 100 bits (250), Expect = 1e-21
Identities = 58/173 (33%), Positives = 88/173 (50%), Gaps = 15/173 (8%)
Frame = +1
Query: 46 KKRRNLPGNPDP----QAEVIALSPKTLMATNRFICEICNKGFQRDQNLQLHKRGHNLPW 101
K ++ L +P +E++ L ++A + CEIC KGF+RD NL++H R H +
Sbjct: 220 KAKQTLDSKLEPLEGDDSEIVELDAVEILAEHMHFCEICGKGFRRDSNLRMHMRAHGEQF 399
Query: 102 KLKQ------RTSNEIRKKVYVCPEPTCVH---HDPSRALGDLTGIKKHFCRKHGEKKWK 152
K + T+ + R + CP C H R L + +K HF R H K +
Sbjct: 400 KTVEALAKPSETTAQRRATRFSCPFEGCNRNKLHRRFRPLKSVICVKNHFKRSHCPKMYT 579
Query: 153 CDKCSKK-YAVQSDWKAHSKTC-GTREYRCDCGTLFSRRDSFITHRAFCDALA 203
C++C KK ++V SD ++H+K C G ++C CGT FSR+D H A D A
Sbjct: 580 CERCRKKHFSVLSDLRSHAKHCGGEARWKCTCGTTFSRKDKLFGHIALFDGHA 738
>AW780545
Length = 409
Score = 97.4 bits (241), Expect = 1e-20
Identities = 44/61 (72%), Positives = 50/61 (81%)
Frame = +2
Query: 163 QSDWKAHSKTCGTREYRCDCGTLFSRRDSFITHRAFCDALAEESSRTVIPQPTQPNSHHN 222
++DWKAHSKTCGTREYRCDCGTLFSRRDSFITHRAFCDALA +S+R P P H+
Sbjct: 2 RADWKAHSKTCGTREYRCDCGTLFSRRDSFITHRAFCDALAHDSARH--PSSLNPLGTHH 175
Query: 223 M 223
+
Sbjct: 176 L 178
>TC212493 similar to GB|AAM10292.1|20147151|AY091693 AT3g13810/MCP4_2
{Arabidopsis thaliana;} , partial (5%)
Length = 912
Score = 96.3 bits (238), Expect = 3e-20
Identities = 82/205 (40%), Positives = 105/205 (51%), Gaps = 36/205 (17%)
Frame = +2
Query: 193 ITHRAFCDALAEESSRTVIPQP---TQPNSHHNMNNLQTQDIQGFTLKKEHQSFNMLRPE 249
ITHRAFCDALA+ESSR V P P TQ SH LQ Q LK+EH FN+L E
Sbjct: 2 ITHRAFCDALAQESSRVVNPHPLLSTQFRSH----GLQLQ--APSLLKREHDHFNLLTSE 163
Query: 250 QEVQIPSWLC------------QSSIDLSSNYSSLDQ-----------DLHLYENPNPRN 286
IPSWL +I +S+Y S Q L +NPNP
Sbjct: 164 ----IPSWLTSPTVVEEAILLNNQTIRTTSDYFSTPQLFPTAHVNNNHSLLHDQNPNPNT 331
Query: 287 GPTST-------LPSYQPSSAASPHMSATALLQKAAQMGAT-SSCSSQSMMSGTH--QQG 336
T+T P+Y SS++SPHMSA ALLQKA+Q+G T SS SQ+M+ H Q
Sbjct: 332 TTTTTFLSSLSSFPNYSTSSSSSPHMSA-ALLQKASQIGETVSSAPSQAMLVRPHLLLQQ 508
Query: 337 HVSIVDSATNNMINSNGNFSLNLSS 361
V + + T I + G +++N++S
Sbjct: 509 QVHVPECTTTTAIATTG-YNINMAS 580
>TC211236 similar to PIR|S48856|S48856 finger protein pcp1 - potato {Solanum
tuberosum;} , partial (6%)
Length = 684
Score = 72.4 bits (176), Expect(2) = 2e-19
Identities = 48/101 (47%), Positives = 55/101 (53%), Gaps = 2/101 (1%)
Frame = +1
Query: 369 NSFSSSGFHGTSFEDTFAGNILHSNQDHNINHDGDNDIPKTTTNDDDVAAGGNN--AFTR 426
NSFSSS F GT FEDTF G GD T AGGNN TR
Sbjct: 235 NSFSSSPFEGT-FEDTFGG--------------GD---AMTADEGGGGGAGGNNNEGLTR 360
Query: 427 DFLGLKPLSDSDILTIAGMGSCMNPSNSNHQENHSQKPWEG 467
DFLGL+ LS +DIL IAG+G+CMN S Q N ++ PW+G
Sbjct: 361 DFLGLRHLSHTDILNIAGVGNCMNSS----QHNQTRNPWQG 471
Score = 41.6 bits (96), Expect(2) = 2e-19
Identities = 30/60 (50%), Positives = 34/60 (56%)
Frame = +3
Query: 307 ATALLQKAAQMGATSSCSSQSMMSGTHQQGHVSIVDSATNNMINSNGNFSLNLSSCEDQM 366
ATALLQKAAQMGAT S + SM+ QQ HVS N +LNLSS + QM
Sbjct: 3 ATALLQKAAQMGATMS-KTGSMIRTHQQQAHVS-------------ANAALNLSSRDHQM 140
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.313 0.126 0.381
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 24,467,944
Number of Sequences: 63676
Number of extensions: 408078
Number of successful extensions: 2250
Number of sequences better than 10.0: 111
Number of HSP's better than 10.0 without gapping: 2182
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2216
length of query: 468
length of database: 12,639,632
effective HSP length: 101
effective length of query: 367
effective length of database: 6,208,356
effective search space: 2278466652
effective search space used: 2278466652
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC148918.4


