

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
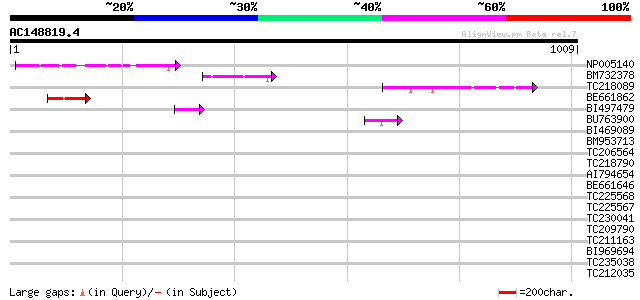
Query= AC148819.4 - phase: 0
(1009 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
NP005140 heat shock protein 82 9e-16
BM732378 65 1e-10
TC218089 54 4e-07
BE661862 50 5e-06
BI497479 43 6e-04
BU763900 43 8e-04
BI469089 41 0.003
BM953713 35 0.13
TC206564 homologue to UP|Q9LF37 (Q9LF37) ClpB heat shock protein... 35 0.13
TC218790 similar to UP|Q7XYX7 (Q7XYX7) AKIN beta2, partial (89%) 35 0.17
AI794654 34 0.37
BE661646 weakly similar to PIR|AG0096|AG00 phosphoenolpyruvate-p... 33 0.48
TC225568 homologue to UP|CLPA_PEA (P35100) ATP-dependent Clp pro... 33 0.82
TC225567 homologue to UP|CLPA_PEA (P35100) ATP-dependent Clp pro... 33 0.82
TC230041 similar to GB|AAB63625.1|2262117|ATAC002343 auxin induc... 32 1.4
TC209790 similar to UP|Q6NMR8 (Q6NMR8) At5g19260, partial (7%) 32 1.4
TC211163 similar to GB|AAR28015.1|39545904|AY463613 TAF12 {Arabi... 32 1.8
BI969694 31 3.1
TC235038 weakly similar to GB|AAB60452.1|555972|MMU15037 replica... 30 4.1
TC212035 similar to UP|NQRD_NEIMA (Q9JVQ1) Na(+)-translocating N... 30 4.1
>NP005140 heat shock protein
Length = 2736
Score = 82.4 bits (202), Expect = 9e-16
Identities = 84/308 (27%), Positives = 134/308 (43%), Gaps = 14/308 (4%)
Frame = +1
Query: 10 ETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSSLRNACLKSQQQNNHPLQC 69
E T + L + LA GHAQ+TP+H+A L+S P+ L
Sbjct: 10 EKFTHKTNEALASAHELAMSSGHAQLTPIHLAHALISDPNGIF---VLAINSAGGGEESA 180
Query: 70 RALELCFNVALNRLPTTTTSPLLQPQHVPSLSNALIAALKRAQAHQR-RGCIEQNQQQQQ 128
RA+E N AL +LP + P VP+ +N L+ A++RAQA Q+ RG
Sbjct: 181 RAVERVLNQALKKLPCQSP----PPDEVPASTN-LVRAIRRAQAAQKSRG---------- 315
Query: 129 QPLLSVKVELDQLILSILDDPSVSRVMREAGFSSPSVKNNLENSSTLINSSSVFHSSPSP 188
++ +DQLIL IL+D + +++EAG + V++ ++ L S S
Sbjct: 316 ----DTRLAVDQLILGILEDSQIGDLLKEAGVAVAKVESEVDK---LRGKEGKKVESASG 474
Query: 189 LSHNHFLSSYGYGSVLFSSQKKEQVVYHPFLKSSESNKEDINLVFDVLLRKKKKNTVIVG 248
++ L +YG ++ + K + V+ E+I V +L R+ K N V+VG
Sbjct: 475 DTNFQALKTYG-RDLVEQAGKLDPVI---------GRDEEIRRVVRILSRRTKNNPVLVG 624
Query: 249 DTVSLTEGLVSEIMKRFERGEVPDEMKTTHFV-----------KFHGLSSVSLKYMKK-- 295
+ +V + +R RG+VP + + K+ G LK + K
Sbjct: 625 EPGVGKTAVVEGLAQRIVRGDVPSNLADVRLIALDMGALVAGAKYRGEFEERLKAVLKEV 804
Query: 296 EEVEMNVI 303
EE E VI
Sbjct: 805 EEAEGKVI 828
Score = 32.7 bits (73), Expect = 0.82
Identities = 20/61 (32%), Positives = 35/61 (56%), Gaps = 5/61 (8%)
Frame = +1
Query: 643 ETVSSIAEALVDSKSS-----KECATWLFLQGNDSVGKKRLALAIAESVFGSVEMFSHVD 697
+ V+++AEA++ S++ + ++LFL G VGK LA A+AE +F + +D
Sbjct: 1726 QAVNAVAEAVLRSRAGLGRPQQPTGSFLFL-GPTGVGKTELAKALAEQLFDNENQLVRID 1902
Query: 698 M 698
M
Sbjct: 1903 M 1905
>BM732378
Length = 434
Score = 65.5 bits (158), Expect = 1e-10
Identities = 41/138 (29%), Positives = 70/138 (50%), Gaps = 6/138 (4%)
Frame = +1
Query: 344 VDYVVEEIGKLFGEEGNKNGKIWLVATASYQSYMRCQMRVPTFENQWCLQAVPVPSGGLG 403
V+++V E+ KL G ++ ++WL+ +++++YM+C++ P+ E W L +P G L
Sbjct: 22 VEHMVMELKKLVCGSG-ESSRLWLMGISTFKTYMKCKICHPSLETIWELHPFTIPVGILI 198
Query: 404 LSLHSSSVHDSKMSISQNPSPMLESKFFSNKEEHEKLNCCEECVSNYEKEAQLF------ 457
LSL+ S ++ +N + F L CC +C N+EKEAQ
Sbjct: 199 LSLNLDSDFQAQ---ERNKVFFKDVAFEDRAGVRNHLTCCRDCTINFEKEAQSITSTISK 369
Query: 458 KPDQKNLLPSWLQSHSTE 475
K + LP+WLQ+ E
Sbjct: 370 KACTTSSLPTWLQNCKEE 423
>TC218089
Length = 1393
Score = 53.5 bits (127), Expect = 4e-07
Identities = 65/298 (21%), Positives = 126/298 (41%), Gaps = 23/298 (7%)
Frame = +2
Query: 664 WLFLQGNDSVGKKRLALAIAESVFGSVEMFSHVDMMKRE--NSETPFSEK-----VVGPL 716
W+ G+D +GKK++A+++AE ++GS E F VD+ E + F K +VG
Sbjct: 68 WMNFVGHDRLGKKKIAVSLAELLYGSRESFIFVDLSSEEMKGCDVKFRGKTALDFIVGEC 247
Query: 717 KNNEKFVVLVENADFGDTLIRKMLADEFEIAKFG-------TLGQKIFILSNG---GSMV 766
VV +EN + D L + L+ + K ++ +F+ S S++
Sbjct: 248 CKKPLSVVFLENVEKADILAQNSLSLAIKTGKISDSHGREVSVNNTMFVFSFSDYQNSLM 427
Query: 767 SEDQKKDSVMKLVLKISETEKKPTFELSPSSSSSSKSPCLGNKRSA--ELDLFSKIKIPR 824
+ + + +L+ K E S N A L+ SK K+
Sbjct: 428 PRGEPSNYSEERILRAKGGGIKIKVEHVIGDIRSQSISLTNNSIDAIPNLNFLSKRKL-- 601
Query: 825 IEENEGNKKREFSFSRQ---SSFNNTLDLNMKADEEDNEDYDEGENSPISSDLTRETLGE 881
I +NE + S + + ++ N LDLN+ A+E + + ++G +SD T +
Sbjct: 602 IGDNEFHDPHLLSDTAKRAHTTSNWLLDLNLPAEENEQKQTNDG-----NSDHVVLTENQ 766
Query: 882 HLISNESLDSIENLFEFNQSPAKNKEMTQMFMSRVKESFEEVLGN-VKFSVQDKVIEE 938
L + D ++ F P + + ++ +F ++LG+ +Q +V+++
Sbjct: 767 KLWLQDLCDLVDETVVF--KPYDFDALADRVLKVIRSNFNKILGSKCALQIQTEVMDQ 934
>BE661862
Length = 597
Score = 50.1 bits (118), Expect = 5e-06
Identities = 35/81 (43%), Positives = 52/81 (63%), Gaps = 4/81 (4%)
Frame = +1
Query: 67 LQCRALELCFNVALNRLPTTTTSPLLQPQHVPSLSNALIAALKRAQAHQRRGCIEQNQ-Q 125
LQ RALEL V+L+RLP++ ++ + + P +SN+L+AA+KR+QA+QRR + Q
Sbjct: 355 LQFRALELSVGVSLDRLPSSKSTSAGEEE--PPVSNSLMAAIKRSQANQRRHPESFHMFQ 528
Query: 126 QQQQPLLS---VKVELDQLIL 143
Q QQ S +KVEL +L
Sbjct: 529 QSQQGTASTSFLKVELKHFVL 591
>BI497479
Length = 311
Score = 43.1 bits (100), Expect = 6e-04
Identities = 25/54 (46%), Positives = 32/54 (58%)
Frame = -3
Query: 294 KKEEVEMNVIRVLKRKVSDYVALGVGAIFYVGDLKWIVDDNDGSLNEKEVVDYV 347
KK E EM+ + LKRKV D +A G G IFYVGDL+W V E ++ Y+
Sbjct: 213 KKNEEEMS-LSALKRKV-DCIASGRGTIFYVGDLEWAVKATSSQKEE*KICCYI 58
>BU763900
Length = 424
Score = 42.7 bits (99), Expect = 8e-04
Identities = 22/75 (29%), Positives = 39/75 (51%), Gaps = 8/75 (10%)
Frame = +2
Query: 632 KVLQENIPWHCETVSSIAEALVDSKSSK--------ECATWLFLQGNDSVGKKRLALAIA 683
++L E + W + + +I++ L KS WL G D +GK+++A A+A
Sbjct: 182 RLLNEKVGWQDQAIRAISQTLSLCKSGAGKRRGSHGRADIWLAFLGPDRLGKRKIASALA 361
Query: 684 ESVFGSVEMFSHVDM 698
E++FG+ E VD+
Sbjct: 362 ETIFGNPESLISVDL 406
>BI469089
Length = 386
Score = 40.8 bits (94), Expect = 0.003
Identities = 38/137 (27%), Positives = 57/137 (40%), Gaps = 3/137 (2%)
Frame = +3
Query: 141 LILSILDDPSVSRVMREAGFSSPSVKNNLENSSTLINSSSVFHSSPSPLSHNHFLSSYGY 200
LILSILDDP VSRV EAGF S +K + PL
Sbjct: 3 LILSILDDPVVSRVFAEAGFRSSDIKLAILR----------------PLRPR-------- 110
Query: 201 GSVLFSSQKKEQVVYHPFL---KSSESNKEDINLVFDVLLRKKKKNTVIVGDTVSLTEGL 257
GS +F E PF + E+ + +VL+R + KN +++G +
Sbjct: 111 GSPIFLCNLSESPRRFPFFFGCGDEDFGGENFRRIGEVLVRSRGKNPLLLGACANDALRG 290
Query: 258 VSEIMKRFERGEVPDEM 274
+E +++ G +P E+
Sbjct: 291 FAEAVEKRREGALPVEL 341
>BM953713
Length = 395
Score = 35.4 bits (80), Expect = 0.13
Identities = 16/47 (34%), Positives = 24/47 (51%)
Frame = +3
Query: 493 CQCLHQNKQPQNHWSNNHSSNAKIYPYNSSYPYWPNQGSSILPDTSS 539
C C +N Q W +N +S + + YN S+P + G+S P SS
Sbjct: 12 CSCQ*ENSSKQQQWRSNATSQSWAFVYNLSFPLSLSNGTSPFPICSS 152
>TC206564 homologue to UP|Q9LF37 (Q9LF37) ClpB heat shock protein-like,
partial (40%)
Length = 1497
Score = 35.4 bits (80), Expect = 0.13
Identities = 23/75 (30%), Positives = 40/75 (52%), Gaps = 5/75 (6%)
Frame = +2
Query: 629 HIYKVLQENIPWHCETVSSIAEALVDSKSS-----KECATWLFLQGNDSVGKKRLALAIA 683
H+ +VL + + V +IAEA+ S++ + A+++F+ G VGK LA A+A
Sbjct: 248 HLEEVLHKRVVGQDPAVKAIAEAIQRSRAGLSDPHRPIASFMFM-GPTGVGKTELAKALA 424
Query: 684 ESVFGSVEMFSHVDM 698
+F + E +DM
Sbjct: 425 AYLFNTEEALVRIDM 469
>TC218790 similar to UP|Q7XYX7 (Q7XYX7) AKIN beta2, partial (89%)
Length = 1533
Score = 35.0 bits (79), Expect = 0.17
Identities = 17/62 (27%), Positives = 32/62 (51%)
Frame = +1
Query: 293 MKKEEVEMNVIRVLKRKVSDYVALGVGAIFYVGDLKWIVDDNDGSLNEKEVVDYVVEEIG 352
+++ + +++VL V + + G Y DL W DD+ + N ++ DYV E+IG
Sbjct: 709 LQRSGKDFTIMKVLPSGVYQFRFIVDGQWRYAPDLPWAQDDSGNAYNVLDLQDYVPEDIG 888
Query: 353 KL 354
+
Sbjct: 889 SI 894
>AI794654
Length = 452
Score = 33.9 bits (76), Expect = 0.37
Identities = 30/130 (23%), Positives = 57/130 (43%), Gaps = 3/130 (2%)
Frame = +3
Query: 796 SSSSSSKSPCLGNKRSAELDLFSKIKIPRIEENEGNKKREFSFSRQSSFNNTLDLNMKAD 855
SS S + N S + +L ++K + + K + S SR + + ++ D
Sbjct: 63 SSGSELQDQGEDNVNSGD-ELQDQVKDGGLSDQSDLKDDDDSISRSPENQSQHNSDVSGD 239
Query: 856 EEDNEDYDEGENSPISSDLTRETL---GEHLISNESLDSIENLFEFNQSPAKNKEMTQMF 912
E N+ + E + + S T T G+H + S+DS +NL + + + NK+ +
Sbjct: 240 NEFNQMSVDSEKNILLSKSTNNTSSGSGDHEFNQMSVDSEKNLLSKSNNTSSNKDEYSVH 419
Query: 913 MSRVKESFEE 922
M+ S +E
Sbjct: 420 MNTQDVSIDE 449
>BE661646 weakly similar to PIR|AG0096|AG00 phosphoenolpyruvate-protein
phosphotransferase (EC 2.7.3.9) [imported] - Yersinia
pestis, partial (2%)
Length = 813
Score = 33.5 bits (75), Expect = 0.48
Identities = 38/144 (26%), Positives = 61/144 (41%), Gaps = 11/144 (7%)
Frame = +2
Query: 98 PSLSNALIAALKRAQAHQRRGCIE------QNQQQQQQPLLSVKVELDQLILSILDDPSV 151
PSL+ IA H + C + +NQQ QQQPL + + Q + S +D +
Sbjct: 32 PSLAEKKIAV---CDIHAEKKCTQVQLVEDENQQPQQQPLAEEEPRIQQDLNSKIDTLT- 199
Query: 152 SRVMREAGFSSPSVKNNLENSSTLINSSSVFHSSPSPLSHNHFLSSYGYGSVLFSSQKKE 211
G S S ++L SS + SSV S + + + G +Q+K
Sbjct: 200 ------KGSSEISQDSSLVFSS---DRSSVTDDDSSKVEEEKHIEATGKKGFNEINQEKL 352
Query: 212 QV-----VYHPFLKSSESNKEDIN 230
+ +YH L + S+K+D+N
Sbjct: 353 ETCSLPSLYHNLLITKASHKDDLN 424
>TC225568 homologue to UP|CLPA_PEA (P35100) ATP-dependent Clp protease
ATP-binding subunit clpA homolog, chloroplast precursor
, partial (36%)
Length = 1287
Score = 32.7 bits (73), Expect = 0.82
Identities = 40/161 (24%), Positives = 69/161 (42%), Gaps = 9/161 (5%)
Frame = +2
Query: 632 KVLQENIPWHCETVSSIAEALVDSK-----SSKECATWLFLQGNDSVGKKRLALAIAESV 686
+ L + + E V +I+ A+ ++ ++ A+++F G VGK LA A+A
Sbjct: 50 ETLHKRVIGQDEAVKAISRAIRRARVGLKNPNRPIASFIF-SGPTGVGKSELAKALAAYY 226
Query: 687 FGSVEMFSHVDMMKRENSETPFSEKVVGPLKNNEKFVVLVENADFGDTLIRK----MLAD 742
FGS E +DM E E K++G + +V E + + R+ +L D
Sbjct: 227 FGSEEAMIRLDM--SEFMERHTVSKLIG---SPPGYVGYTEGGQLTEAVRRRPYTVVLFD 391
Query: 743 EFEIAKFGTLGQKIFILSNGGSMVSEDQKKDSVMKLVLKIS 783
E E A + IL +G S+ + D L++ S
Sbjct: 392 EIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTLLIMTS 514
>TC225567 homologue to UP|CLPA_PEA (P35100) ATP-dependent Clp protease
ATP-binding subunit clpA homolog, chloroplast precursor
, partial (61%)
Length = 2077
Score = 32.7 bits (73), Expect = 0.82
Identities = 40/161 (24%), Positives = 69/161 (42%), Gaps = 9/161 (5%)
Frame = +3
Query: 632 KVLQENIPWHCETVSSIAEALVDSK-----SSKECATWLFLQGNDSVGKKRLALAIAESV 686
+ L + + E V +I+ A+ ++ ++ A+++F G VGK LA A+A
Sbjct: 750 ETLHKRVIGQDEAVKAISRAIRRARVGLKNPNRPIASFIF-SGPTGVGKSELAKALAAYY 926
Query: 687 FGSVEMFSHVDMMKRENSETPFSEKVVGPLKNNEKFVVLVENADFGDTLIRK----MLAD 742
FGS E +DM E E K++G + +V E + + R+ +L D
Sbjct: 927 FGSEEAMIRLDM--SEFMERHTVSKLIG---SPPGYVGYTEGGQLTEAVRRRPYTVVLFD 1091
Query: 743 EFEIAKFGTLGQKIFILSNGGSMVSEDQKKDSVMKLVLKIS 783
E E A + IL +G S+ + D L++ S
Sbjct: 1092EIEKAHPDVFNMMLQILEDGRLTDSKGRTVDFKNTLLIMTS 1214
>TC230041 similar to GB|AAB63625.1|2262117|ATAC002343 auxin inducible protein
isolog {Arabidopsis thaliana;} , partial (18%)
Length = 763
Score = 32.0 bits (71), Expect = 1.4
Identities = 16/52 (30%), Positives = 28/52 (53%), Gaps = 4/52 (7%)
Frame = +3
Query: 416 MSISQNPSP----MLESKFFSNKEEHEKLNCCEECVSNYEKEAQLFKPDQKN 463
M + +P P M++ F ++E+ +K+ CC+ S+ E E L PD +N
Sbjct: 312 MLVGDDPWPEFCNMVKRIFICSREDLKKMKCCKLPASSSEVEEVLLSPDSQN 467
>TC209790 similar to UP|Q6NMR8 (Q6NMR8) At5g19260, partial (7%)
Length = 675
Score = 32.0 bits (71), Expect = 1.4
Identities = 34/111 (30%), Positives = 41/111 (36%), Gaps = 10/111 (9%)
Frame = +3
Query: 815 DLFSKIKIPRIEENEGNKKREFSFSRQSSFNNTLDLNMKADEEDNEDYDEGEN------- 867
D S IK P EEN NKK + SF L K ++ Y +
Sbjct: 303 DSSSTIK-PHSEENNNNKKSTPANPNSWSFREALSNVTKEPSQNQTTYVHPQQKRFSLSL 479
Query: 868 SPISSDLTRETLGEHLISNESLDSIENLFEFNQSPAKN---KEMTQMFMSR 915
SP S L E LG S+ DS EN + S N +E TQ R
Sbjct: 480 SPKSLQLCTENLGNESGSDSGSDSDENSIDMFSSVNGNSGTREQTQQRQPR 632
>TC211163 similar to GB|AAR28015.1|39545904|AY463613 TAF12 {Arabidopsis
thaliana;} , partial (17%)
Length = 724
Score = 31.6 bits (70), Expect = 1.8
Identities = 33/132 (25%), Positives = 47/132 (35%), Gaps = 5/132 (3%)
Frame = +3
Query: 7 TLQETLTAEAASILKHSLGLARRRGHAQITPLHVAATLLSLPSSS-----LRNACLKSQQ 61
++Q +L ++ G +GH + P V +T PSSS L L S
Sbjct: 327 SVQSSLRPPTSAPNNQPAGSQSFQGHGLMRPSSVGSTATPSPSSSQSMQSLNQPWLSSGS 506
Query: 62 QNNHPLQCRALELCFNVALNRLPTTTTSPLLQPQHVPSLSNALIAALKRAQAHQRRGCIE 121
Q PL A N + Q H+P + + +
Sbjct: 507 QGKPPLPSAAYRQQLN----------PQSMQQRSHIPPMQQSTPTS-------------- 614
Query: 122 QNQQQQQQPLLS 133
+QQQQQQPLLS
Sbjct: 615 -SQQQQQQPLLS 647
>BI969694
Length = 556
Score = 30.8 bits (68), Expect = 3.1
Identities = 19/52 (36%), Positives = 27/52 (51%), Gaps = 5/52 (9%)
Frame = -1
Query: 917 KESFEEVLGNVKFSVQDKVIEE-----IGVGCGSFTNNMFEKWLKGIFQTSL 963
KE E +LG V FS+ K+++E I VGC S F++ L +F L
Sbjct: 418 KEGVEPILGVVSFSIPFKIVKEEKGWVIPVGCLSKPMKRFQRELACVFTPPL 263
>TC235038 weakly similar to GB|AAB60452.1|555972|MMU15037 replication factor
C large subunit {Mus musculus;} , partial (3%)
Length = 507
Score = 30.4 bits (67), Expect = 4.1
Identities = 14/52 (26%), Positives = 25/52 (47%)
Frame = -3
Query: 455 QLFKPDQKNLLPSWLQSHSTEARQKDELTQLNKKWNRLCQCLHQNKQPQNHW 506
+L + D + P W +E+ + E+ +L KK +C C N++P W
Sbjct: 274 ELMRYDTVKVSPLWWFPSDSESEEVAEVNELKKKSEAVCLCC--NREPNGIW 125
>TC212035 similar to UP|NQRD_NEIMA (Q9JVQ1) Na(+)-translocating NADH-quinone
reductase subunit D (Na(+)-translocating NQR subunit D)
(Na(+)-NQR subunit D) (NQR complex subunit D) (NQR-1
subunit D) , partial (8%)
Length = 336
Score = 30.4 bits (67), Expect = 4.1
Identities = 12/41 (29%), Positives = 24/41 (58%)
Frame = +3
Query: 80 LNRLPTTTTSPLLQPQHVPSLSNALIAALKRAQAHQRRGCI 120
++ +PTT SPL P + P+ + +L+R+++ RR +
Sbjct: 93 ISTVPTTLPSPLQPPSNAPNAPTLSLQSLRRSRSSSRRRAV 215
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.313 0.129 0.367
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 43,579,450
Number of Sequences: 63676
Number of extensions: 666023
Number of successful extensions: 3851
Number of sequences better than 10.0: 55
Number of HSP's better than 10.0 without gapping: 3750
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3827
length of query: 1009
length of database: 12,639,632
effective HSP length: 107
effective length of query: 902
effective length of database: 5,826,300
effective search space: 5255322600
effective search space used: 5255322600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 64 (29.3 bits)
Medicago: description of AC148819.4


