

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
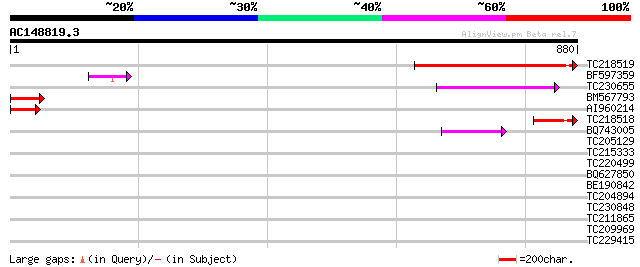
Query= AC148819.3 + phase: 0 /pseudo/partial
(880 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC218519 similar to UP|Q9LSJ0 (Q9LSJ0) Dbj|BAA20807.2, partial (... 387 e-107
BF597359 105 1e-22
TC230655 103 3e-22
BM567793 100 4e-21
AI960214 91 2e-18
TC218518 90 5e-18
BQ743005 54 4e-07
TC205129 homologue to UP|HT22_ARATH (P46604) Homeobox-leucine zi... 32 1.2
TC215333 similar to UP|P93344 (P93344) Aldehyde dehydrogenase (N... 32 1.2
TC220499 weakly similar to UP|Q751I1 (Q751I1) AGL275Wp, partial ... 32 1.2
BQ627850 31 2.7
BE190842 30 3.5
TC204894 similar to UP|Q94KA9 (Q94KA9) Importin alpha 2, partial... 30 4.6
TC230848 similar to UP|Q8LPT2 (Q8LPT2) At1g10410/F14N23_31, part... 30 6.0
TC211865 weakly similar to UP|Q7F266 (Q7F266) Nipped-B gene prod... 30 6.0
TC209969 similar to UP|Q9SBS1 (Q9SBS1) Ran GTPase activating pro... 29 7.9
TC229415 homologue to UP|Q9Y1J5 (Q9Y1J5) RNA binding protein Puf... 29 7.9
>TC218519 similar to UP|Q9LSJ0 (Q9LSJ0) Dbj|BAA20807.2, partial (24%)
Length = 1021
Score = 387 bits (993), Expect = e-107
Identities = 195/254 (76%), Positives = 215/254 (83%), Gaps = 2/254 (0%)
Frame = +2
Query: 629 KEPKCMLFPPRMFFSEEDIPEGSSFTAGERMHELVKVFVLLHQLQIFTHGRALPDQPLIY 688
KEPKC+LFP +M SEEDIPEGSSF AGE+MHELVKVFV+LHQLQIFT GR LP++PLIY
Sbjct: 8 KEPKCILFPSQMLSSEEDIPEGSSFAAGEKMHELVKVFVVLHQLQIFTLGRPLPEKPLIY 187
Query: 689 HPCDHRTNSRAHTSGL-AAVPKPGTEMNLVNAVPCRIAFERGKERHFCFLAISVGTSGWL 747
P D NSRA TSGL + PKPGTE++LVNAVPCRIAFERGKERHFCFLAIS GTSGWL
Sbjct: 188 PPGDLPANSRAQTSGLDVSGPKPGTEVSLVNAVPCRIAFERGKERHFCFLAISAGTSGWL 367
Query: 748 VLAEELPLKKPYGVVRVAAPLAGCNPRVDDKHSKWLHLRIRPSALPFLDPVKYNRHGKLK 807
VLAEELP+KK YGV+RVAAPLAGCNPR+DDKH +WLHLRIRPS+LP LDP K+N + KLK
Sbjct: 368 VLAEELPMKKLYGVIRVAAPLAGCNPRIDDKHPRWLHLRIRPSSLPVLDPAKFNPNRKLK 547
Query: 808 TKAFVDGRWILAFRDEESCKNAFLMIREEINHLCDEVDRRIKPLLKLETVLDISSSSAPV 867
TKAFVDGRW LAFRDEESCK+A MI EEIN L DEV RR+KPLL LET LD+S
Sbjct: 548 TKAFVDGRWTLAFRDEESCKSALSMILEEINFLSDEVHRRLKPLLNLETALDLSGP---- 715
Query: 868 SEDSSSH-TTPPDS 880
EDSSSH TTPP+S
Sbjct: 716 EEDSSSHSTTPPNS 757
>BF597359
Length = 406
Score = 105 bits (261), Expect = 1e-22
Identities = 58/102 (56%), Positives = 61/102 (58%), Gaps = 36/102 (35%)
Frame = -1
Query: 123 DEVVSFPLYVEAIRFAFHEESMIRAAVRAVTLNVYH------------------------ 158
DEVVSFPLYVEAIRFAFHEE+MIR AVR VTLNVYH
Sbjct: 394 DEVVSFPLYVEAIRFAFHEENMIRTAVRTVTLNVYHGKIYYFTWPPFFFLPELSYVMMQK 215
Query: 159 ------------VGDDSVNRYITSEPHTDYFSNLVSFFRKQC 188
VGD+ VNRYITS PHT+YFSNLVSFFR QC
Sbjct: 214 YLIF*QLAVHFSVGDECVNRYITSAPHTEYFSNLVSFFRNQC 89
>TC230655
Length = 960
Score = 103 bits (258), Expect = 3e-22
Identities = 61/191 (31%), Positives = 95/191 (48%)
Frame = +1
Query: 663 VKVFVLLHQLQIFTHGRALPDQPLIYHPCDHRTNSRAHTSGLAAVPKPGTEMNLVNAVPC 722
++ F+L QL+ F L D+PL+ +SR + + G+ ++L + +PC
Sbjct: 85 IQAFILHFQLKTFILKGVLVDKPLLNMISSSTNDSRVIRASDVSSASFGSNVSLESGIPC 264
Query: 723 RIAFERGKERHFCFLAISVGTSGWLVLAEELPLKKPYGVVRVAAPLAGCNPRVDDKHSKW 782
IAF + R + ++ G G L+LAE+ P + +GVV APLAG P++D+ H W
Sbjct: 265 GIAFSNSEIRDIYVIPVASGIIGKLLLAEKHPFRSRHGVVIAIAPLAGLFPKIDEDHPSW 444
Query: 783 LHLRIRPSALPFLDPVKYNRHGKLKTKAFVDGRWILAFRDEESCKNAFLMIREEINHLCD 842
LHL+IR F H + DGRW L F + +C+ A L+I EI
Sbjct: 445 LHLQIREFDPQFYSTKARGNHLSMPDH-LADGRWTLGFPNARACEEAHLVILNEITKQRS 621
Query: 843 EVDRRIKPLLK 853
V+ + PLL+
Sbjct: 622 AVEYMLAPLLQ 654
>BM567793
Length = 428
Score = 100 bits (248), Expect = 4e-21
Identities = 47/53 (88%), Positives = 52/53 (97%)
Frame = +1
Query: 1 DFVIEALRSIAELVTYGDQHDPTFFEFFMEKQVVGDFVRILKLSRTISIPLQL 53
DFVIEALRSIAEL+TYGDQHDP+FFEFFMEKQVV +FVR+LKLSRT+SIPLQL
Sbjct: 268 DFVIEALRSIAELITYGDQHDPSFFEFFMEKQVVAEFVRVLKLSRTVSIPLQL 426
>AI960214
Length = 399
Score = 90.9 bits (224), Expect = 2e-18
Identities = 42/48 (87%), Positives = 47/48 (97%)
Frame = +2
Query: 1 DFVIEALRSIAELVTYGDQHDPTFFEFFMEKQVVGDFVRILKLSRTIS 48
DFVIEALRSIAEL+TYGDQHDP+FFEFFMEKQVV +FVR+LKLSRT+S
Sbjct: 254 DFVIEALRSIAELITYGDQHDPSFFEFFMEKQVVAEFVRVLKLSRTVS 397
>TC218518
Length = 489
Score = 89.7 bits (221), Expect = 5e-18
Identities = 48/68 (70%), Positives = 52/68 (75%), Gaps = 1/68 (1%)
Frame = +1
Query: 814 GRWILAFRDEESCKNAFLMIREEINHLCDEVDRRIKPLLKLETVLDISSSSAPVSEDSSS 873
GRW LAFRDEESCK+A MI EEIN L DEV RR+KPLL LET LD+ S P EDSSS
Sbjct: 1 GRWTLAFRDEESCKSALSMILEEINFLSDEVHRRLKPLLNLETALDL---SGPEEEDSSS 171
Query: 874 H-TTPPDS 880
H TTPP+S
Sbjct: 172 HSTTPPNS 195
>BQ743005
Length = 422
Score = 53.5 bits (127), Expect = 4e-07
Identities = 32/100 (32%), Positives = 51/100 (51%)
Frame = +2
Query: 671 QLQIFTHGRALPDQPLIYHPCDHRTNSRAHTSGLAAVPKPGTEMNLVNAVPCRIAFERGK 730
QL+ F L D+PL+ +SR + + G+ ++L + +PC IAF +
Sbjct: 8 QLKTFILKGVLVDKPLLNMISSSTNDSRVIRASDVSSASFGSNVSLESGIPCGIAFSNSE 187
Query: 731 ERHFCFLAISVGTSGWLVLAEELPLKKPYGVVRVAAPLAG 770
R + ++ G G L+LAE+ P + +GVV APLAG
Sbjct: 188 IRDIYVIPVASGIIGKLLLAEKHPFRSRHGVVIAIAPLAG 307
>TC205129 homologue to UP|HT22_ARATH (P46604) Homeobox-leucine zipper protein
HAT22 (HD-ZIP protein 22), partial (47%)
Length = 793
Score = 32.0 bits (71), Expect = 1.2
Identities = 28/79 (35%), Positives = 38/79 (47%), Gaps = 7/79 (8%)
Frame = -1
Query: 330 NLAENSAKCLVVNVPC----SSSSSDCHPQSITI---LNNGSSSNVALREVLLEYVTGGA 382
N + CL + V C SSSS+DC S++ L SSS++ L E L T +
Sbjct: 238 NCLAKACFCLGLRVLCCLKLSSSSADCSLVSLSFFLALVPSSSSSLTLEETLSSASTSTS 59
Query: 383 DVQVLGSLSVLATLLQTKE 401
L SLS+L TL K+
Sbjct: 58 MSSQLRSLSLLMTLPVEKD 2
>TC215333 similar to UP|P93344 (P93344) Aldehyde dehydrogenase (NAD+) ,
partial (73%)
Length = 1405
Score = 32.0 bits (71), Expect = 1.2
Identities = 16/39 (41%), Positives = 23/39 (58%), Gaps = 3/39 (7%)
Frame = -1
Query: 649 EGSSFTAGERMHELVKVFV---LLHQLQIFTHGRALPDQ 684
EGSS +R H+LV ++HQL++ HG LP+Q
Sbjct: 199 EGSSKE*SDRQHQLVHAMFAHDMVHQLELLDHGSYLPNQ 83
>TC220499 weakly similar to UP|Q751I1 (Q751I1) AGL275Wp, partial (4%)
Length = 1075
Score = 32.0 bits (71), Expect = 1.2
Identities = 16/50 (32%), Positives = 27/50 (54%)
Frame = +3
Query: 619 KRAMESSSPPKEPKCMLFPPRMFFSEEDIPEGSSFTAGERMHELVKVFVL 668
K+ +++ SPPK PK +L P +PE + + +LVK+F+L
Sbjct: 114 KKMLDTFSPPKNPKLLLLPESRLPCNTKLPEDCHY----QPEDLVKLFLL 251
>BQ627850
Length = 428
Score = 30.8 bits (68), Expect = 2.7
Identities = 24/91 (26%), Positives = 42/91 (45%)
Frame = +3
Query: 53 LLQTVSIMIQNLQSEHAIYYMFSNEHMNYLITYSFDFRNEELLSYYISFLRAIGGKLNKN 112
LLQT S ++Q L + Y FSN+++N + ++F +Y++ + I L +
Sbjct: 105 LLQTKSKVLQVLHFPAFVLYFFSNQYINIXVVLKYNFDTSS*F-FYLNLVS*I*KVLLYS 281
Query: 113 TISLLVKTRNDEVVSFPLYVEAIRFAFHEES 143
+ L N F L+ + I F H+ S
Sbjct: 282 LVKYLYVILNWFFYQF-LFTDGINFG*HDIS 371
>BE190842
Length = 450
Score = 30.4 bits (67), Expect = 3.5
Identities = 12/34 (35%), Positives = 21/34 (61%)
Frame = -2
Query: 288 ANTIVAALYYPLESFTKCSGGQVNGYIPDHGFTS 321
++T+VA + + SFT+C Q Y+ HGF++
Sbjct: 302 SSTLVAPMMIKISSFTECPVSQKPSYV*SHGFSA 201
>TC204894 similar to UP|Q94KA9 (Q94KA9) Importin alpha 2, partial (95%)
Length = 2332
Score = 30.0 bits (66), Expect = 4.6
Identities = 24/70 (34%), Positives = 39/70 (55%), Gaps = 8/70 (11%)
Frame = +2
Query: 221 LYYFSDIISAGIPDV-ERLITDSILMLLIFP---VLLPSLRTVAD----NDMQSGVVTSL 272
L Y SD + I V E + ++ LL+ P VL+P+LRTV + +DMQ+ V+ +
Sbjct: 962 LSYLSDGTNDKIQAVIEAGVCPRLVELLLHPSPSVLIPALRTVGNIVTGDDMQTQVIINH 1141
Query: 273 YLLCCILRIV 282
L C+L ++
Sbjct: 1142 QALPCLLNLL 1171
>TC230848 similar to UP|Q8LPT2 (Q8LPT2) At1g10410/F14N23_31, partial (24%)
Length = 1400
Score = 29.6 bits (65), Expect = 6.0
Identities = 17/48 (35%), Positives = 23/48 (47%), Gaps = 3/48 (6%)
Frame = -3
Query: 691 CDHRTNSRA---HTSGLAAVPKPGTEMNLVNAVPCRIAFERGKERHFC 735
CDH T +R+ H+S + P G+ V A R + RGK R C
Sbjct: 363 CDHPTRTRSEPSHSSKASRFPAEGSSRASVRAPTVRASSSRGKARTPC 220
>TC211865 weakly similar to UP|Q7F266 (Q7F266) Nipped-B gene product-like
protein, partial (6%)
Length = 871
Score = 29.6 bits (65), Expect = 6.0
Identities = 26/117 (22%), Positives = 49/117 (41%), Gaps = 12/117 (10%)
Frame = +2
Query: 274 LLCCILRIVKIKDLANTIVAALYYPLESFTKCSGGQVNGYIPDHG----FTSKSDGIDND 329
L+C + ++DL N AA ++P ++ C GG+V HG F + GI
Sbjct: 59 LVCSQEKFWILQDLLNQDAAAQHHPKDTCCVCLGGRVENLFICHGCQRLFHADCLGIKEH 238
Query: 330 NLAENSAKC--------LVVNVPCSSSSSDCHPQSITILNNGSSSNVALREVLLEYV 378
++ + C L+V C +S + ++ S ++++LL Y+
Sbjct: 239 EVSSRNWSCQTCICHKKLLVLQSCCNSQQKNDVKKNCNTDSEVSKQEIVQQLLLNYL 409
>TC209969 similar to UP|Q9SBS1 (Q9SBS1) Ran GTPase activating protein, partial
(64%)
Length = 1143
Score = 29.3 bits (64), Expect = 7.9
Identities = 14/24 (58%), Positives = 17/24 (70%)
Frame = -1
Query: 857 VLDISSSSAPVSEDSSSHTTPPDS 880
+LD SSS+ P S DSSS +TP S
Sbjct: 1047 ILDSSSSAPPSSSDSSSSSTPSGS 976
>TC229415 homologue to UP|Q9Y1J5 (Q9Y1J5) RNA binding protein PufA, partial
(3%)
Length = 1099
Score = 29.3 bits (64), Expect = 7.9
Identities = 17/50 (34%), Positives = 29/50 (58%), Gaps = 1/50 (2%)
Frame = +2
Query: 317 HGFTSKSDGIDNDNLAENSAKCLVVNVPCSSS-SSDCHPQSITILNNGSS 365
HG KS G + + A + VN+ C+SS +S+ HP+ ++L++ SS
Sbjct: 353 HG--GKSSGSSSPSTAGKRLRLFGVNMECASSTTSEDHPKCFSLLSSSSS 496
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.323 0.139 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 41,692,829
Number of Sequences: 63676
Number of extensions: 632395
Number of successful extensions: 3169
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 3122
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3160
length of query: 880
length of database: 12,639,632
effective HSP length: 106
effective length of query: 774
effective length of database: 5,889,976
effective search space: 4558841424
effective search space used: 4558841424
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 63 (28.9 bits)
Medicago: description of AC148819.3


