

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148817.2 - phase: 0
(608 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
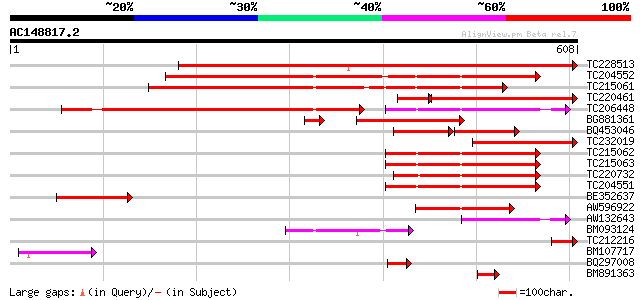
Score E
Sequences producing significant alignments: (bits) Value
TC228513 similar to UP|MAOM_SOLTU (P37221) NAD-dependent malic e... 585 e-167
TC204552 homologue to UP|Q9M4Q9 (Q9M4Q9) NADP-dependent malic pr... 322 3e-88
TC215061 similar to UP|O24550 (O24550) Malate dehydrogenase , p... 316 2e-86
TC220461 similar to UP|Q9LEN2 (Q9LEN2) NAD-dependent malic enzym... 277 3e-84
TC206448 homologue to GB|CAA56354.1|510876|PVME1G NADP dependent... 286 1e-77
BG881361 homologue to GP|8894554|emb NAD-dependent malic enzyme ... 214 7e-61
BQ453046 similar to GP|8894554|emb NAD-dependent malic enzyme (m... 127 2e-59
TC232019 homologue to UP|Q9LEN2 (Q9LEN2) NAD-dependent malic enz... 190 2e-48
TC215062 similar to UP|MAOX_VITVI (P51615) NADP-dependent malic ... 150 2e-36
TC215063 similar to UP|MAOX_VITVI (P51615) NADP-dependent malic ... 146 3e-35
TC220732 similar to UP|Q6PMI1 (Q6PMI1) NADP-dependent malic enzy... 134 9e-32
TC204551 homologue to UP|Q9M4Q9 (Q9M4Q9) NADP-dependent malic pr... 134 1e-31
BE352637 similar to SP|P37225|MAON NAD-dependent malic enzyme 59... 104 1e-22
AW596922 similar to GP|8118507|gb| NADP-dependent malic protein ... 79 6e-15
AW132643 similar to GP|8118507|gb| NADP-dependent malic protein ... 77 2e-14
BM093124 74 1e-13
TC212216 similar to UP|Q9LEN2 (Q9LEN2) NAD-dependent malic enzym... 54 2e-07
BM107717 similar to SP|P37225|MAON_ NAD-dependent malic enzyme 5... 53 4e-07
BQ297008 similar to GP|8894554|emb| NAD-dependent malic enzyme (... 47 3e-05
BM891363 homologue to GP|8894554|emb| NAD-dependent malic enzyme... 45 7e-05
>TC228513 similar to UP|MAOM_SOLTU (P37221) NAD-dependent malic enzyme 62 kDa
isoform, mitochondrial precursor (NAD-ME) , partial
(69%)
Length = 1451
Score = 585 bits (1508), Expect = e-167
Identities = 284/432 (65%), Positives = 363/432 (83%), Gaps = 5/432 (1%)
Frame = +2
Query: 182 LTDGSRILGLGDLGVQGIGIPVGKLDMYVAAAGINPQRILPIMLDVGTNNQKLLDDRLYL 241
+TDGSRILGLGDLGVQGIGI +GKLD+YVAAAGINPQR+LP+M+DVGTNN+KLL+D LYL
Sbjct: 2 VTDGSRILGLGDLGVQGIGIAIGKLDLYVAAAGINPQRVLPVMIDVGTNNEKLLEDPLYL 181
Query: 242 GLRQPRLEGEEYLSIVDEFMEAVHARWPKAIVQFEDFQMKWAFETLKRYRHKFCMFNDDI 301
GL+Q RL+G++YL++VDEFMEAV RWP IVQFEDFQ KWAF+ L+RYR+ + MFNDD+
Sbjct: 182 GLQQHRLDGDDYLAVVDEFMEAVFTRWPNVIVQFEDFQSKWAFKLLQRYRNTYRMFNDDV 361
Query: 302 QGTAGVALAGLLGTVRAQGRPLSDFAKQKIVIVGAGSAGLGVLNMAVQAVSRMSGCSETA 361
QGTAGVA+AGLLG VRAQGRP+ DF KQKIV+ GAGSAG+GVLN A + ++RM G +E A
Sbjct: 362 QGTAGVAIAGLLGAVRAQGRPMIDFPKQKIVVAGAGSAGIGVLNAARKTMARMLGNNEVA 541
Query: 362 ---AKSQFFLIDKNGLVTTERSNLDPAAVPFAKNPRDLE--GLAEGASIIEVVKKVKPHV 416
AKSQF+++D GL+T R N+DP A+PFA+N ++++ GL EGAS++EVVK+VKP V
Sbjct: 542 FESAKSQFWVVDAQGLITEGRENIDPDALPFARNLKEMDRQGLREGASLVEVVKQVKPDV 721
Query: 417 LLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMNAECTAIDAFNHAGENIVFASGS 476
LLGLS VGG+F+++VL+A++ S ST+PAIFAMSNPT NAECTA +AF+ G+NI+FASGS
Sbjct: 722 LLGLSAVGGLFSKEVLEALKGSTSTRPAIFAMSNPTKNAECTAEEAFSILGDNIIFASGS 901
Query: 477 PFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAASECLASYMTEEDI 536
PF NVDLGNG GH NQ NNMYLFPGIGLG+LLSGA ++DGMLQAA+E LA+YM+EE++
Sbjct: 902 PFSNVDLGNGHIGHCNQGNNMYLFPGIGLGTLLSGARIVSDGMLQAAAERLATYMSEEEV 1081
Query: 537 QKGILYPSIDCIRNVTAEVGAAVLRAAIAEGLAEGHGGVGSKELEHMSEDDTVEYVRGNM 596
KGI++PS IR++T +V AV++ A+ E LAEG+ G+ ++EL+ +SED+ EYV+ NM
Sbjct: 1082LKGIIFPSTSRIRDITKQVATAVIKEAVEEDLAEGYHGMDARELQKLSEDEIAEYVQNNM 1261
Query: 597 WYPEYCPLVHEK 608
W PEY LV++K
Sbjct: 1262WSPEYPTLVYKK 1297
>TC204552 homologue to UP|Q9M4Q9 (Q9M4Q9) NADP-dependent malic protein ,
partial (66%)
Length = 1640
Score = 322 bits (825), Expect = 3e-88
Identities = 171/404 (42%), Positives = 250/404 (61%), Gaps = 2/404 (0%)
Frame = +3
Query: 168 MIYNWPAPQVDMIVLTDGSRILGLGDLGVQGIGIPVGKLDMYVAAAGINPQRILPIMLDV 227
++ NWP + +IV+TDG RILGLGDLG QG+GIPVGKL +Y A G+ P LPI +DV
Sbjct: 6 VLRNWPEKNIQVIVVTDGERILGLGDLGCQGMGIPVGKLSLYTALGGVRPSACLPITIDV 185
Query: 228 GTNNQKLLDDRLYLGLRQPRLEGEEYLSIVDEFMEAVHARW-PKAIVQFEDFQMKWAFET 286
GTNN+KLL+D LY+GL+Q R G+EY ++ EFM AV + K ++QFEDF AF+
Sbjct: 186 GTNNEKLLNDELYIGLKQRRATGQEYAELMHEFMTAVKQTYGEKVLIQFEDFANHNAFDL 365
Query: 287 LKRYRHKFCMFNDDIQGTAGVALAGLLGTVRAQGRPLSDFAKQKIVIVGAGSAGLGVLNM 346
L++YR +FNDDIQGTA V LAGL+ +++ G L+D + + +GAG AG G+ +
Sbjct: 366 LEKYRSTHLVFNDDIQGTASVVLAGLVASLKLVGGNLAD---HRFLFLGAGEAGTGIAEL 536
Query: 347 AVQAVSRMSGCSETAAKSQFFLIDKNGLVTTER-SNLDPAAVPFAKNPRDLEGLAEGASI 405
S+ + + +L+D GL+ + R +L P+A ++ L
Sbjct: 537 IALETSKQTNAPLEEVRKNIWLVDSKGLIVSSRKDSLQHFKKPWAHEHEPVKNL------ 698
Query: 406 IEVVKKVKPHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMNAECTAIDAFNH 465
++ V K+KP VL+G SG G F ++V++AM S++ +P I ++SNPT +ECTA +A+
Sbjct: 699 LDAVNKIKPTVLIGTSGQGRTFTKEVIEAM-ASINERPIILSLSNPTSQSECTAEEAYKW 875
Query: 466 AGENIVFASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAASE 525
+ +FASGSPF V+ G+ QANN Y+FPG GLG ++SG + D +L AASE
Sbjct: 876 SQGRAIFASGSPFPPVEY-EGKVFVPGQANNAYIFPGFGLGLIMSGTIRVHDDLLLAASE 1052
Query: 526 CLASYMTEEDIQKGILYPSIDCIRNVTAEVGAAVLRAAIAEGLA 569
LA+ +++E+ KG++YP IR ++A + A V A GLA
Sbjct: 1053ALAAQVSQENFDKGLIYPPFTNIRKISAHIAANVAAKAYELGLA 1184
>TC215061 similar to UP|O24550 (O24550) Malate dehydrogenase , partial (58%)
Length = 1146
Score = 316 bits (809), Expect = 2e-86
Identities = 168/387 (43%), Positives = 243/387 (62%), Gaps = 1/387 (0%)
Frame = +1
Query: 149 YRRPRGMYFSAKDKGEMMSMIYNWPAPQVDMIVLTDGSRILGLGDLGVQGIGIPVGKLDM 208
++RP+G++ S K+KG+++ ++ NWP + +IV+TDG RILGLGDLG QG+GIPVGKL +
Sbjct: 1 FKRPQGLFISLKEKGKVLEVLKNWPEKSIQVIVVTDGERILGLGDLGCQGMGIPVGKLAL 180
Query: 209 YVAAAGINPQRILPIMLDVGTNNQKLLDDRLYLGLRQPRLEGEEYLSIVDEFMEAVHARW 268
Y A G+ P LPI +DVGTNN+KLL+D Y+GLRQ R G+EY ++ EFM AV +
Sbjct: 181 YTALGGVRPSACLPITIDVGTNNEKLLNDEFYIGLRQKRATGQEYSELLQEFMTAVKQNY 360
Query: 269 -PKAIVQFEDFQMKWAFETLKRYRHKFCMFNDDIQGTAGVALAGLLGTVRAQGRPLSDFA 327
K ++QFEDF AFE L +Y +FNDDIQGTA V LAG++ ++ G L+D
Sbjct: 361 GEKVLIQFEDFANHNAFELLAKYGTTHLVFNDDIQGTASVVLAGVVAALKLIGGNLAD-- 534
Query: 328 KQKIVIVGAGSAGLGVLNMAVQAVSRMSGCSETAAKSQFFLIDKNGLVTTERSNLDPAAV 387
+ +GAG AG G+ + +S+ + + + +L+D GL+ R A++
Sbjct: 535 -HTFLFLGAGEAGTGIAELIALEMSKQTKTPIEETRKKIWLVDSKGLIVGSRK----ASL 699
Query: 388 PFAKNPRDLEGLAEGASIIEVVKKVKPHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFA 447
K P E G S++E VK +KP VL+G SGVG F ++V++A+ S++ KP + A
Sbjct: 700 QHFKQPWAHEHEPVG-SLLEAVKVIKPTVLIGSSGVGKTFTKEVIEAV-TSINEKPLVLA 873
Query: 448 MSNPTMNAECTAIDAFNHAGENIVFASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGS 507
+SNPT +ECTA +A+ + +FASGSPF+ V+ G+ + QANN Y+FPG GLG
Sbjct: 874 LSNPTSQSECTAEEAYQWSEGRAIFASGSPFDPVEY-KGKVYYSGQANNAYIFPGFGLGL 1050
Query: 508 LLSGAHHITDGMLQAASECLASYMTEE 534
++SGA + D ML AS LA + E
Sbjct: 1051VISGAIRVHDDMLSTASVALAKLVYNE 1131
>TC220461 similar to UP|Q9LEN2 (Q9LEN2) NAD-dependent malic enzyme (Malate
oxidoreductase) (Fragment) , partial (59%)
Length = 1101
Score = 277 bits (709), Expect(2) = 3e-84
Identities = 131/156 (83%), Positives = 147/156 (93%)
Frame = +1
Query: 453 MNAECTAIDAFNHAGENIVFASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGA 512
+ AECTAI+AF+HAGENIVFASGSPFENVDLGNG GHVNQANNMYLFPGIGLG+LLSGA
Sbjct: 346 VTAECTAIEAFSHAGENIVFASGSPFENVDLGNGEVGHVNQANNMYLFPGIGLGTLLSGA 525
Query: 513 HHITDGMLQAASECLASYMTEEDIQKGILYPSIDCIRNVTAEVGAAVLRAAIAEGLAEGH 572
HITDGML+AA+ECLASYMTE+D++KGILYPSIDCIR+VTAEVGAAV+ AA+AE AEGH
Sbjct: 526 RHITDGMLRAAAECLASYMTEDDVRKGILYPSIDCIRDVTAEVGAAVVCAAVAEKQAEGH 705
Query: 573 GGVGSKELEHMSEDDTVEYVRGNMWYPEYCPLVHEK 608
G VG KELE+MS+++TVEYVRGNMWYPEYCPLVHEK
Sbjct: 706 GDVGFKELENMSKEETVEYVRGNMWYPEYCPLVHEK 813
Score = 53.9 bits (128), Expect(2) = 3e-84
Identities = 27/39 (69%), Positives = 31/39 (79%)
Frame = +2
Query: 416 VLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMN 454
+++G G + QVLKAMRESVSTKPAIFAMSNPTMN
Sbjct: 143 IVIGFLHFNG*DSTQVLKAMRESVSTKPAIFAMSNPTMN 259
>TC206448 homologue to GB|CAA56354.1|510876|PVME1G NADP dependent malic
enzyme {Phaseolus vulgaris;} , partial (97%)
Length = 2208
Score = 286 bits (733), Expect = 1e-77
Identities = 143/326 (43%), Positives = 205/326 (62%), Gaps = 1/326 (0%)
Frame = +3
Query: 56 ERDRLGLRGLLPPRIISFQEQYDRFMSSYRSLEKNTHGQSDKIVSLSKWRILNRLHDRNE 115
ERD LRGLLPP I + + Q R M + R E V L ++ + L +RNE
Sbjct: 300 ERDAHYLRGLLPPAIFNQELQEKRLMHNLRQYE----------VPLHRYMAMMDLQERNE 449
Query: 116 TLYYRALIDNIKEFAPIIYTPTVGLVCQNYCGLYRRPRGMYFSAKDKGEMMSMIYNWPAP 175
L+Y+ LI+N++E P++YTPTVG CQ Y ++RRP+G+Y S K+KG+++ ++ NWP
Sbjct: 450 RLFYKLLINNVEELLPVVYTPTVGEACQKYGSIFRRPQGLYISLKEKGKILEVLKNWPEK 629
Query: 176 QVDMIVLTDGSRILGLGDLGVQGIGIPVGKLDMYVAAAGINPQRILPIMLDVGTNNQKLL 235
+ +IV+TDG RILGLGDLG QG+GIPVGKL +Y A G+ P LPI +DVGTNN+KLL
Sbjct: 630 SIQVIVVTDGERILGLGDLGCQGMGIPVGKLSLYTALGGVRPSSCLPITIDVGTNNEKLL 809
Query: 236 DDRLYLGLRQPRLEGEEYLSIVDEFMEAVHARW-PKAIVQFEDFQMKWAFETLKRYRHKF 294
+D Y+GL+Q R G+EY +++EFM AV + K ++QFEDF AF+ L++Y
Sbjct: 810 NDEFYIGLKQKRATGQEYAELLEEFMHAVKQNYGEKVLIQFEDFANHNAFDLLEKYSSSH 989
Query: 295 CMFNDDIQGTAGVALAGLLGTVRAQGRPLSDFAKQKIVIVGAGSAGLGVLNMAVQAVSRM 354
+FNDDIQGTA V LAGLL +++ G L D + +GAG AG G+ + +S+
Sbjct: 990 LVFNDDIQGTASVVLAGLLASLKLVGGTLVD---HTFLFLGAGEAGTGIAELIALEISKQ 1160
Query: 355 SGCSETAAKSQFFLIDKNGLVTTERS 380
+ + + +L+D G + + RS
Sbjct: 1161TKAPVEETRKKIWLVDSKGKIASSRS 1238
Score = 142 bits (359), Expect = 3e-34
Identities = 81/199 (40%), Positives = 118/199 (58%), Gaps = 1/199 (0%)
Frame = +2
Query: 404 SIIEVVKKVKPHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMNAECTAIDAF 463
S+++ VK +KP VL+G SG G F ++V++ M S++ KP I A+SNPT +ECTA +A+
Sbjct: 1394 SLLDAVKAIKPTVLIGSSGAGKTFTKEVVETMA-SLNEKPLILALSNPTSQSECTAEEAY 1570
Query: 464 NHAGENIVFASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAA 523
+ +FASGSPF+ V+ G+ QANN Y+FPG GLG ++SGA + D ML AA
Sbjct: 1571 TWSKGKAIFASGSPFDPVEY-EGKLFVPGQANNAYIFPGFGLGLIISGAIRVRDEMLLAA 1747
Query: 524 SECLASYMTEEDIQKGILYPSIDCIRNVTAEVGAAVLRAAIAEGLAEGHGGVGSKELEHM 583
SE LA+ +++E+ KG++YP IR ++A + A V A GLA H+
Sbjct: 1748 SEALAAQVSQENYDKGLIYPPFSNIRKISAHIAAKVAAKAYDLGLA-----------SHL 1894
Query: 584 SE-DDTVEYVRGNMWYPEY 601
D V+Y M+ P Y
Sbjct: 1895 PRPKDLVKYAESCMYSPGY 1951
>BG881361 homologue to GP|8894554|emb NAD-dependent malic enzyme (malate
oxidoreductase) {Cicer arietinum}, partial (44%)
Length = 412
Score = 214 bits (544), Expect(2) = 7e-61
Identities = 103/115 (89%), Positives = 112/115 (96%)
Frame = +2
Query: 373 GLVTTERSNLDPAAVPFAKNPRDLEGLAEGASIIEVVKKVKPHVLLGLSGVGGVFNEQVL 432
GLVTTER++LDPAA PFAKNPRD+EGL EGASIIEVVKK++PHVLLGLSGVGG+FNE+VL
Sbjct: 68 GLVTTERNSLDPAAAPFAKNPRDIEGLTEGASIIEVVKKIRPHVLLGLSGVGGIFNEEVL 247
Query: 433 KAMRESVSTKPAIFAMSNPTMNAECTAIDAFNHAGENIVFASGSPFENVDLGNGR 487
KAMRESVSTKPAIFAMSNPTMNAECT+IDAF HAGENIVFASGSPFENVDLGNG+
Sbjct: 248 KAMRESVSTKPAIFAMSNPTMNAECTSIDAFKHAGENIVFASGSPFENVDLGNGK 412
Score = 39.3 bits (90), Expect(2) = 7e-61
Identities = 17/21 (80%), Positives = 19/21 (89%)
Frame = +3
Query: 317 RAQGRPLSDFAKQKIVIVGAG 337
RAQGRPL+DF QKIV+VGAG
Sbjct: 3 RAQGRPLTDFVNQKIVVVGAG 65
>BQ453046 similar to GP|8894554|emb NAD-dependent malic enzyme (malate
oxidoreductase) {Cicer arietinum}, partial (44%)
Length = 407
Score = 127 bits (320), Expect(2) = 2e-59
Identities = 60/69 (86%), Positives = 65/69 (93%)
Frame = +1
Query: 478 FENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAASECLASYMTEEDIQ 537
FENVDLGNG HVNQANNMYLFPGIGLG+LLSGA HITDGML+AA+ECLASYMT+ED+Q
Sbjct: 199 FENVDLGNGEFFHVNQANNMYLFPGIGLGTLLSGARHITDGMLRAAAECLASYMTDEDVQ 378
Query: 538 KGILYPSID 546
KGILYPSID
Sbjct: 379 KGILYPSID 405
Score = 120 bits (301), Expect(2) = 2e-59
Identities = 59/65 (90%), Positives = 62/65 (94%)
Frame = +2
Query: 412 VKPHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMNAECTAIDAFNHAGENIV 471
VKPHVLLGLSGVGGVFN +VLKAMRESVSTKPAIFAMSNPTMNAECTAI+AF+HAGENI
Sbjct: 2 VKPHVLLGLSGVGGVFNAEVLKAMRESVSTKPAIFAMSNPTMNAECTAIEAFSHAGENIA 181
Query: 472 FASGS 476
FA GS
Sbjct: 182 FAIGS 196
>TC232019 homologue to UP|Q9LEN2 (Q9LEN2) NAD-dependent malic enzyme (Malate
oxidoreductase) (Fragment) , partial (37%)
Length = 616
Score = 190 bits (482), Expect = 2e-48
Identities = 93/112 (83%), Positives = 99/112 (88%)
Frame = +2
Query: 497 MYLFPGIGLGSLLSGAHHITDGMLQAASECLASYMTEEDIQKGILYPSIDCIRNVTAEVG 556
MYLFPGIGLG+LLSGAH ITDGMLQAASECLASYM EEDI KGILYPS+D IR+VTAEVG
Sbjct: 2 MYLFPGIGLGTLLSGAHLITDGMLQAASECLASYMAEEDILKGILYPSVDSIRDVTAEVG 181
Query: 557 AAVLRAAIAEGLAEGHGGVGSKELEHMSEDDTVEYVRGNMWYPEYCPLVHEK 608
AAVLRAA+ E LAEGHG VG KEL HMS+D+ VEYVR NMWYP Y PLVHEK
Sbjct: 182 AAVLRAAVEEELAEGHGDVGPKELSHMSKDEAVEYVRSNMWYPVYSPLVHEK 337
>TC215062 similar to UP|MAOX_VITVI (P51615) NADP-dependent malic enzyme
(NADP-ME) , partial (36%)
Length = 971
Score = 150 bits (378), Expect = 2e-36
Identities = 77/166 (46%), Positives = 112/166 (67%)
Frame = +2
Query: 404 SIIEVVKKVKPHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMNAECTAIDAF 463
S++E VK +KP VL+G SGVG F ++V++A+ S++ KP + A+SNPT +ECTA +A+
Sbjct: 62 SLLEAVKVIKPTVLIGSSGVGKTFTKEVIEAVT-SINEKPLVLALSNPTSQSECTAEEAY 238
Query: 464 NHAGENIVFASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAA 523
+ +FASGSPF+ V+ G+ + QANN Y+FPG GLG ++SGA + D ML AA
Sbjct: 239 EWSEGRAIFASGSPFDPVEY-KGKVYYSGQANNAYIFPGFGLGLVISGAIRVHDDMLLAA 415
Query: 524 SECLASYMTEEDIQKGILYPSIDCIRNVTAEVGAAVLRAAIAEGLA 569
SE LA +TEE+ +KG++YP IR ++A + A+V A GLA
Sbjct: 416 SEALAKLVTEENYEKGLIYPPFSNIRKISANIAASVAAKAYELGLA 553
>TC215063 similar to UP|MAOX_VITVI (P51615) NADP-dependent malic enzyme
(NADP-ME) , partial (32%)
Length = 918
Score = 146 bits (368), Expect = 3e-35
Identities = 75/166 (45%), Positives = 111/166 (66%)
Frame = +2
Query: 404 SIIEVVKKVKPHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMNAECTAIDAF 463
S++E VK +KP VL+G SGVG F ++V++A+ S++ KP + A+SNPT +ECTA +A+
Sbjct: 8 SLLEAVKVIKPTVLIGSSGVGKTFTKEVIEAVT-SINEKPLVLALSNPTSQSECTAEEAY 184
Query: 464 NHAGENIVFASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAA 523
+ +FASGSPF+ V+ G+ + QANN Y+FPG GLG ++SGA + D ML AA
Sbjct: 185 QWSEGRAIFASGSPFDPVEY-KGKVYYSGQANNAYIFPGFGLGLVISGAIRVHDDMLLAA 361
Query: 524 SECLASYMTEEDIQKGILYPSIDCIRNVTAEVGAAVLRAAIAEGLA 569
SE LA ++ E+ +KG++YP IR ++A + A+V A GLA
Sbjct: 362 SEALAKLVSNENYEKGLIYPPFSNIREISANIAASVAGKAYELGLA 499
>TC220732 similar to UP|Q6PMI1 (Q6PMI1) NADP-dependent malic enzyme 3 ,
partial (32%)
Length = 662
Score = 134 bits (338), Expect = 9e-32
Identities = 72/158 (45%), Positives = 102/158 (63%)
Frame = +1
Query: 412 VKPHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMNAECTAIDAFNHAGENIV 471
+KP VL+G SGVG F ++V++AM + + KP I A+SNPT +ECTA +A+ + +
Sbjct: 40 IKPTVLIGSSGVGRTFTKEVVEAMTSN-NDKPLILALSNPTSQSECTAEEAYQWSEGRAI 216
Query: 472 FASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAASECLASYM 531
FASGSPF+ V+ G+ QANN Y+FPG GLG ++SGA + D ML AASE LA +
Sbjct: 217 FASGSPFDPVEY-KGKVYASGQANNAYIFPGFGLGLVISGAIRVHDDMLLAASESLAKQV 393
Query: 532 TEEDIQKGILYPSIDCIRNVTAEVGAAVLRAAIAEGLA 569
+EE+ + G++YP IR ++A + A V A GLA
Sbjct: 394 SEENYKNGLIYPPFSNIRRISANIAANVAAKAYELGLA 507
>TC204551 homologue to UP|Q9M4Q9 (Q9M4Q9) NADP-dependent malic protein ,
partial (30%)
Length = 856
Score = 134 bits (337), Expect = 1e-31
Identities = 69/166 (41%), Positives = 106/166 (63%)
Frame = +2
Query: 404 SIIEVVKKVKPHVLLGLSGVGGVFNEQVLKAMRESVSTKPAIFAMSNPTMNAECTAIDAF 463
++++ V K+KP L+G SG G F ++V++AM S++ +P I ++SNPT +ECTA +A+
Sbjct: 14 NLLDAVNKIKPTELIGTSGQGRTFTKEVIEAMA-SINKRPIILSLSNPTSQSECTAEEAY 190
Query: 464 NHAGENIVFASGSPFENVDLGNGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAA 523
+ +FASGSPF V+ G+ QANN Y+FPG GLG ++SG + D +L AA
Sbjct: 191 KWSQGRAIFASGSPFPPVEY-EGKVFVPGQANNAYIFPGFGLGLIMSGTIRVHDDLLLAA 367
Query: 524 SECLASYMTEEDIQKGILYPSIDCIRNVTAEVGAAVLRAAIAEGLA 569
SE LA+ +++E+ KG++YP IR ++A + A V A GLA
Sbjct: 368 SEALAAQVSQENFDKGLIYPPFTNIRKISAHIAANVAAKAYELGLA 505
>BE352637 similar to SP|P37225|MAON NAD-dependent malic enzyme 59 kDa isoform
mitochondrial precursor (EC 1.1.1.39) (NAD-ME).
[Potato], partial (18%)
Length = 557
Score = 104 bits (259), Expect = 1e-22
Identities = 54/82 (65%), Positives = 61/82 (73%), Gaps = 1/82 (1%)
Frame = +2
Query: 51 GFPLTERDRLGLRGLLPPRIISFQEQYDRFMSSYRSLEKNTHGQSDKIVSLSKWRILNRL 110
GFPLTERDRLGLRGLLPP +ISF++QYDRFM+S+RSLE NT G DK+VS
Sbjct: 191 GFPLTERDRLGLRGLLPPXVISFEQQYDRFMNSFRSLENNTQGLPDKVVSWQNGGS*XDC 370
Query: 111 HDRNETLYYRALID-NIKEFAP 131
RNETLYYR LI +KEFAP
Sbjct: 371 MTRNETLYYRVLIX*YLKEFAP 436
>AW596922 similar to GP|8118507|gb| NADP-dependent malic protein {Ricinus
communis}, partial (16%)
Length = 460
Score = 79.0 bits (193), Expect = 6e-15
Identities = 40/106 (37%), Positives = 66/106 (61%)
Frame = +2
Query: 436 RESVSTKPAIFAMSNPTMNAECTAIDAFNHAGENIVFASGSPFENVDLGNGRAGHVNQAN 495
R S++ +P I ++SNPT +ECTA +A + +FASGSPF V+ G+ QAN
Sbjct: 5 RASINERPIILSLSNPT*QSECTAEEANEWSHGRAIFASGSPFPPVEY-EGKVFVPGQAN 181
Query: 496 NMYLFPGIGLGSLLSGAHHITDGMLQAASECLASYMTEEDIQKGIL 541
N Y+FPG GLG ++SG + + ++AA E A+ +++E + + ++
Sbjct: 182 NAYIFPGFGLGLIMSGTILVQEDQVEAACEA*AAQLSQETMDQELI 319
>AW132643 similar to GP|8118507|gb| NADP-dependent malic protein {Ricinus
communis}, partial (17%)
Length = 381
Score = 77.0 bits (188), Expect = 2e-14
Identities = 43/117 (36%), Positives = 62/117 (52%)
Frame = +3
Query: 485 NGRAGHVNQANNMYLFPGIGLGSLLSGAHHITDGMLQAASECLASYMTEEDIQKGILYPS 544
+G+ QANN Y+FPG GLG ++SG + D +L AASE LAS +T++D KG++YP
Sbjct: 15 DGKVFVPGQANNAYIFPGFGLGLIMSGTIRVHDDLLLAASEALASQVTQKDYDKGLIYPP 194
Query: 545 IDCIRNVTAEVGAAVLRAAIAEGLAEGHGGVGSKELEHMSEDDTVEYVRGNMWYPEY 601
IR ++A + A V A GLA D V++ +M+ P Y
Sbjct: 195 FSNIRKISARIAANVAAKAYELGLA----------TSLPQPKDLVKFAESSMYTPLY 335
>BM093124
Length = 420
Score = 74.3 bits (181), Expect = 1e-13
Identities = 45/144 (31%), Positives = 77/144 (53%), Gaps = 6/144 (4%)
Frame = +2
Query: 296 MFNDDIQGTAGVALAGLLGTVRAQGRPLSDFAKQKIVIVGAGSAGLGVLNMAVQAVSRMS 355
+FNDDIQGTA V LAGLL +++ G L+D + +GAG AG G+ + +S+ +
Sbjct: 14 VFNDDIQGTASVVLAGLLASLKLVGGTLAD---HTFLFLGAGEAGTGIAELIALEISKQT 184
Query: 356 GCSETAAKSQFFLIDK-----NGLVTTER-SNLDPAAVPFAKNPRDLEGLAEGASIIEVV 409
+ + +L+D GL+ + R +L P+A ++ ++++ V
Sbjct: 185 KAPVEKTRKKIWLVDSKVKMPQGLIVSSRLESLQHFKKPWAHEHEPVK------TLLDAV 346
Query: 410 KKVKPHVLLGLSGVGGVFNEQVLK 433
K +KP VL+G SG G F ++V++
Sbjct: 347 KAIKPTVLIGSSGAGKTFTKEVVE 418
>TC212216 similar to UP|Q9LEN2 (Q9LEN2) NAD-dependent malic enzyme (Malate
oxidoreductase) (Fragment) , partial (9%)
Length = 415
Score = 54.3 bits (129), Expect = 2e-07
Identities = 22/27 (81%), Positives = 24/27 (88%)
Frame = +1
Query: 582 HMSEDDTVEYVRGNMWYPEYCPLVHEK 608
HMS+D+TVEYVR NMWYP Y PLVHEK
Sbjct: 1 HMSKDETVEYVRSNMWYPVYSPLVHEK 81
>BM107717 similar to SP|P37225|MAON_ NAD-dependent malic enzyme 59 kDa
isoform mitochondrial precursor (EC 1.1.1.39) (NAD-ME).
[Potato], partial (13%)
Length = 408
Score = 52.8 bits (125), Expect = 4e-07
Identities = 37/89 (41%), Positives = 47/89 (52%), Gaps = 5/89 (5%)
Frame = +3
Query: 10 CGRLRDSPH---HRDRGCSRRRYLHRVWFINVVLIFFMILGS--IRGFPLTERDRLGLRG 64
CG D P DRG S R+L R F + V + M GS I+GF + G
Sbjct: 135 CGNSCDMPPPPTSPDRGGSPPRFLVRALFTSAVPTYSMTPGSTRIQGFL*LKGIGWGFGA 314
Query: 65 LLPPRIISFQEQYDRFMSSYRSLEKNTHG 93
R+ISF++QYDRFM+S+RSL NT G
Sbjct: 315 SYLLRVISFEQQYDRFMNSFRSLXNNTPG 401
>BQ297008 similar to GP|8894554|emb| NAD-dependent malic enzyme (malate
oxidoreductase) {Cicer arietinum}, partial (14%)
Length = 169
Score = 46.6 bits (109), Expect = 3e-05
Identities = 20/25 (80%), Positives = 24/25 (96%)
Frame = +1
Query: 406 IEVVKKVKPHVLLGLSGVGGVFNEQ 430
IEVVKK++ HVLLGLSGVGG+FNE+
Sbjct: 1 IEVVKKIRSHVLLGLSGVGGIFNEE 75
Score = 35.0 bits (79), Expect = 0.097
Identities = 16/20 (80%), Positives = 17/20 (85%)
Frame = +2
Query: 527 LASYMTEEDIQKGILYPSID 546
LASYM EEDI GILYPS+D
Sbjct: 77 LASYMAEEDILIGILYPSLD 136
>BM891363 homologue to GP|8894554|emb| NAD-dependent malic enzyme (malate
oxidoreductase) {Cicer arietinum}, partial (7%)
Length = 427
Score = 45.4 bits (106), Expect = 7e-05
Identities = 22/24 (91%), Positives = 23/24 (95%)
Frame = +1
Query: 502 GIGLGSLLSGAHHITDGMLQAASE 525
GIGLG+LLSGAH ITDGMLQAASE
Sbjct: 1 GIGLGTLLSGAHLITDGMLQAASE 72
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.322 0.139 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,167,589
Number of Sequences: 63676
Number of extensions: 321897
Number of successful extensions: 1466
Number of sequences better than 10.0: 46
Number of HSP's better than 10.0 without gapping: 1432
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1441
length of query: 608
length of database: 12,639,632
effective HSP length: 103
effective length of query: 505
effective length of database: 6,081,004
effective search space: 3070907020
effective search space used: 3070907020
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 62 (28.5 bits)
Medicago: description of AC148817.2


