

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148816.5 + phase: 0 /pseudo
(211 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
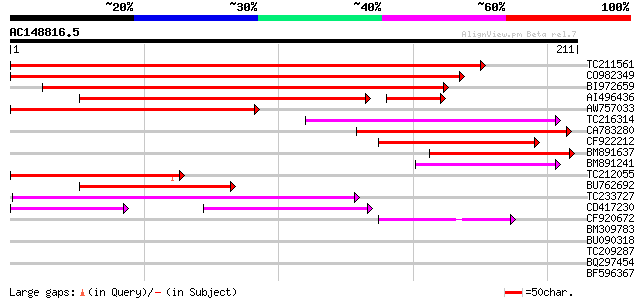
Score E
Sequences producing significant alignments: (bits) Value
TC211561 weakly similar to UP|O22675 (O22675) Reverse transcript... 165 2e-41
CO982349 151 2e-37
BI972659 134 3e-32
AI496436 108 7e-26
AW757033 102 1e-22
TC216314 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {A... 88 3e-18
CA783280 weakly similar to PIR|T01871|T01 RNA-directed DNA polym... 78 3e-15
CF922212 69 2e-12
BM891637 60 7e-10
BM891241 60 9e-10
TC212055 59 2e-09
BU762692 57 6e-09
TC233727 similar to UP|Q9FKH8 (Q9FKH8) Similarity to non-LTR ret... 48 4e-06
CD417230 weakly similar to GP|2244915|emb| reverse transcriptase... 36 2e-05
CF920672 40 6e-04
BM309783 39 0.002
BU090318 weakly similar to GP|9743345|gb| F21J9.30 {Arabidopsis ... 37 0.007
TC209287 similar to UP|HNM1_YEAST (P19807) Choline transport pro... 37 0.009
BQ297454 32 0.21
BF596367 29 1.4
>TC211561 weakly similar to UP|O22675 (O22675) Reverse transcriptase
(Fragment) , partial (59%)
Length = 1077
Score = 165 bits (417), Expect = 2e-41
Identities = 77/177 (43%), Positives = 117/177 (65%)
Frame = +1
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMAC 60
+S LQYADDT+ G+A+ +N+ +K ILR F++VS L+INF+KS V++ + A
Sbjct: 379 ISILQYADDTIFFGEAAKENVEAIKVILRSFELVSNLRINFAKSCFGVFGVTDQWKQEAA 558
Query: 61 DFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNSV 120
++L+CS + F YLG+P+GAN R W+ ++ +L+ W R+ISFGGR+ L+ SV
Sbjct: 559 NYLHCSLLAFPFIYLGIPIGANSRRCHMWDSIIRKCERKLSKWK*RHISFGGRVTLIKSV 738
Query: 121 LNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGV 177
L S+PI++ SF ++ V ++V+I R FLWGG KKISW+ W+ +C KE GG+
Sbjct: 739 LTSIPIYFFSFFRVPQSVVDKLVKI*RTFLWGGGPDQKKISWIRWEKLCLPKERGGI 909
>CO982349
Length = 795
Score = 151 bits (382), Expect = 2e-37
Identities = 72/169 (42%), Positives = 111/169 (65%)
Frame = +3
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMAC 60
VS LQYADDT+ +A+++N+ +K ILR F++ SGLKINF++S I + + A
Sbjct: 282 VSILQYADDTIFFEEATMENMKVIKTILRCFELASGLKINFAESRFGAIWKPDHWCKEAA 461
Query: 61 DFLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNSV 120
+FLNCS S+ F YLG+P+ AN R +P++ +L W R+IS GGR+ L+N++
Sbjct: 462 EFLNCSMLSMPFSYLGIPIEANPRCREI*DPIIRKCETKLARWKQRHISLGGRVTLINAI 641
Query: 121 LNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVC 169
L ++PI++ SF + V +++V IQR FLWGG ++I+WV W++VC
Sbjct: 642 LTALPIYFFSFFRAPSKVVQKLVNIQRKFLWGGGSXQRRIAWVKWETVC 788
>BI972659
Length = 453
Score = 134 bits (337), Expect = 3e-32
Identities = 62/151 (41%), Positives = 95/151 (62%)
Frame = +1
Query: 13 IGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMACDFLNCSAGSITF 72
+G+AS N+ +K++LRGF+MVSGL+IN++KS + ++ A LNC I F
Sbjct: 1 VGEASWDNIIVLKSMLRGFEMVSGLRINYAKSQFGIVGFQPNWAHDAAQLLNCRQLDIPF 180
Query: 73 KYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNSVLNSMPIFYLSFL 132
YLG+P+ S WEPL+ +L+ W+ +Y+S GGR+ L+ SVLN++PI+ LSF
Sbjct: 181 HYLGMPIAVKASSRMVWEPLINKFKAKLSKWNQKYLSMGGRVTLIKSVLNALPIYLLSFF 360
Query: 133 KMLVGVWKRIVRIQR*FLWGGVGGGKKISWV 163
K+ + ++V +QR F+WGG +ISWV
Sbjct: 361 KIPQRIVDKLVSLQRTFMWGGNQHHNRISWV 453
>AI496436
Length = 414
Score = 108 bits (270), Expect(2) = 7e-26
Identities = 52/108 (48%), Positives = 73/108 (67%)
Frame = +1
Query: 27 ILRGFQMVSGLKINFSKSSLVGINVSEDFMAMACDFLNCSAGSITFKYLGLPVGANMRSM 86
ILR F++ SGLKINF+KS I S+ + A D+LN S S+ F YLG+P+GAN R
Sbjct: 4 ILRCFELSSGLKINFAKSQFGTICKSDIWRKEAADYLNSSVLSMPFVYLGIPIGANSRHS 183
Query: 87 STWEPLVETIGGRLNTWSTRYISFGGRIVLLNSVLNSMPIFYLSFLKM 134
WEP+V +L +W +Y+SFGGR+ L+NSVL ++PI+ LSF ++
Sbjct: 184 DVWEPVVRKCERKLASWKQKYVSFGGRVTLINSVLTALPIYLLSFFRI 327
Score = 25.4 bits (54), Expect(2) = 7e-26
Identities = 11/22 (50%), Positives = 15/22 (68%)
Frame = +2
Query: 141 RIVRIQR*FLWGGVGGGKKISW 162
++ IQR FLWGG +KI+W
Sbjct: 347 KLTIIQRRFLWGGDLE*RKIAW 412
>AW757033
Length = 441
Score = 102 bits (255), Expect = 1e-22
Identities = 47/93 (50%), Positives = 68/93 (72%)
Frame = -2
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMAC 60
VS LQYADDT+ G+A+++N+ +K ILRGF++ SGLKINF+KS I+V + + A
Sbjct: 284 VSILQYADDTIFFGEATMENVRVIKTILRGFELASGLKINFAKSRFGAISVPD*WRKEAA 105
Query: 61 DFLNCSAGSITFKYLGLPVGANMRSMSTWEPLV 93
+F+NCS S+ F YLG+P+GAN R TW+P++
Sbjct: 104 EFMNCSLLSLPFSYLGIPIGANPRRRETWDPII 6
>TC216314 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {Arabidopsis
thaliana;} , partial (18%)
Length = 759
Score = 87.8 bits (216), Expect = 3e-18
Identities = 38/95 (40%), Positives = 56/95 (58%)
Frame = +2
Query: 111 GGRIVLLNSVLNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQ 170
GGR+ L+ SVL+++PI LSF K+ + ++V +QR FLWGG +I WV W +C
Sbjct: 5 GGRVTLIKSVLSALPIXXLSFFKIPQRIVDKLVTLQRQFLWGGTQHHNRIPWVKWADICN 184
Query: 171 QKENGGVRVKDIRVMNISLLAKWRWRLIDGREALW 205
K +GG+ +KD+ N +L +W W L LW
Sbjct: 185 PKIDGGLGIKDLSNFNAALRGRWIWGLASNHNQLW 289
>CA783280 weakly similar to PIR|T01871|T01 RNA-directed DNA polymerase
homolog T24M8.8 - Arabidopsis thaliana, partial (4%)
Length = 456
Score = 78.2 bits (191), Expect = 3e-15
Identities = 35/80 (43%), Positives = 49/80 (60%)
Frame = +2
Query: 130 SFLKMLVGVWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVRVKDIRVMNISL 189
SF + + ++ +QR FLWGG +KI+WVNWK+VC K GG+ +KD+R N +L
Sbjct: 5 SFFSLP*KIIAKLESLQRRFLWGGEADSRKIAWVNWKTVCLPKAKGGLGIKDLRTFNTTL 184
Query: 190 LAKWRWRLIDGREALWK*VL 209
L KWRW L ++ W VL
Sbjct: 185 LGKWRWDLFYIQQEPWAKVL 244
>CF922212
Length = 445
Score = 68.9 bits (167), Expect = 2e-12
Identities = 32/60 (53%), Positives = 41/60 (68%)
Frame = -2
Query: 138 VWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVRVKDIRVMNISLLAKWRWRL 197
V ++VR+Q+ FLWGG G KKI+WVNW SVC KE G+ +D+R N +LL K RW L
Sbjct: 429 VLDKLVRLQQWFLWGGGLGQKKIAWVNWDSVCLPKEKEGLGNRDLRKFNYALLGKRRWNL 250
>BM891637
Length = 427
Score = 60.1 bits (144), Expect = 7e-10
Identities = 21/54 (38%), Positives = 36/54 (65%)
Frame = -1
Query: 157 GKKISWVNWKSVCQQKENGGVRVKDIRVMNISLLAKWRWRLIDGREALWK*VLV 210
GKKI+W++W+ C + GG+ +KDI+++N +LL KW+W + LW +L+
Sbjct: 421 GKKIAWISWRQCCASGDVGGLGIKDIKILNNALLIKWKWLMFHQPHQLWNRILI 260
>BM891241
Length = 407
Score = 59.7 bits (143), Expect = 9e-10
Identities = 26/54 (48%), Positives = 32/54 (59%)
Frame = -2
Query: 152 GGVGGGKKISWVNWKSVCQQKENGGVRVKDIRVMNISLLAKWRWRLIDGREALW 205
GG KKI WV W+ VC K GG+ +KDI NI+LL KW W L ++ LW
Sbjct: 406 GGDHDHKKIPWVKWEVVCLPKTKGGLGIKDISKFNIALLGKWIWALASDQQQLW 245
>TC212055
Length = 776
Score = 58.5 bits (140), Expect = 2e-09
Identities = 30/66 (45%), Positives = 47/66 (70%), Gaps = 1/66 (1%)
Frame = +1
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMA- 59
V+ LQYADDT+ +G+A++QN+ T+K++LR F++ SGLKI+F+KSS I + + A
Sbjct: 100 VNILQYADDTIFLGEATMQNVMTIKSMLRVFELASGLKIHFAKSSFGAIGKPDHWQKEAG 279
Query: 60 CDFLNC 65
+ NC
Sbjct: 280 LNTFNC 297
>BU762692
Length = 423
Score = 57.0 bits (136), Expect = 6e-09
Identities = 28/58 (48%), Positives = 38/58 (65%)
Frame = -3
Query: 27 ILRGFQMVSGLKINFSKSSLVGINVSEDFMAMACDFLNCSAGSITFKYLGLPVGANMR 84
ILR F++VSGLKINF+KS +++ + A LNCS +I F Y G+P+GAN R
Sbjct: 181 ILRTFELVSGLKINFAKSCFGAFGMTDRWKTEAASCLNCSLLAIPFVYQGIPIGANPR 8
>TC233727 similar to UP|Q9FKH8 (Q9FKH8) Similarity to non-LTR retroelement
reverse transcriptase, partial (7%)
Length = 956
Score = 47.8 bits (112), Expect = 4e-06
Identities = 31/129 (24%), Positives = 60/129 (46%)
Frame = -2
Query: 2 SHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKSSLVGINVSEDFMAMACD 61
SH+ +ADD + K + +N+ + + ++ G I+ KS + ++ + + D
Sbjct: 451 SHVMHADDIMIFCKGTSRNVHNLINFFQLYKDSFGQVISKHKSKIFIGSIPGQRLRVISD 272
Query: 62 FLNCSAGSITFKYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNSVL 121
L I F Y G+P+ + + + I +L +W +S GRI L+NSV+
Sbjct: 271 LLQYFHSDIPFNYFGVPIFKGKPTRRHLYCVADKIKCKLASWKGSILSIMGRIQLVNSVI 92
Query: 122 NSMPIFYLS 130
+ M ++ S
Sbjct: 91 HGMLLYSFS 65
>CD417230 weakly similar to GP|2244915|emb| reverse transcriptase like
protein {Arabidopsis thaliana}, partial (7%)
Length = 688
Score = 36.2 bits (82), Expect(2) = 2e-05
Identities = 16/63 (25%), Positives = 34/63 (53%)
Frame = -2
Query: 73 KYLGLPVGANMRSMSTWEPLVETIGGRLNTWSTRYISFGGRIVLLNSVLNSMPIFYLSFL 132
KYLG+P+ + T++ +++ + R + W + +S GR+ L V+ ++P + F
Sbjct: 246 KYLGVPIFHKNPTSHTFDFIIDKVNKRFSRWKSNLLSMVGRLTLTK*VV*ALPSHIMQFD 67
Query: 133 KML 135
+L
Sbjct: 66 NLL 58
Score = 28.5 bits (62), Expect(2) = 2e-05
Identities = 13/44 (29%), Positives = 23/44 (51%)
Frame = -3
Query: 1 VSHLQYADDTLCIGKASVQNLWTMKAILRGFQMVSGLKINFSKS 44
+SHL + DD + +A++ + + L F SG K+N K+
Sbjct: 467 ISHLAFVDDLILFVEANMDQVDVINLTLELFSDSSGAKVNVDKT 336
>CF920672
Length = 792
Score = 40.4 bits (93), Expect = 6e-04
Identities = 23/51 (45%), Positives = 30/51 (58%)
Frame = -2
Query: 138 VWKRIVRIQR*FLWGGVGGGKKISWVNWKSVCQQKENGGVRVKDIRVMNIS 188
V +++ +IQR F WGG KKI+WV W C E +RVK R+ NIS
Sbjct: 782 VAEKLTQIQRRFXWGGGLDQKKIAWVKWD--C*LMEQEPLRVKYPRLYNIS 636
>BM309783
Length = 433
Score = 38.5 bits (88), Expect = 0.002
Identities = 21/58 (36%), Positives = 30/58 (51%)
Frame = +2
Query: 152 GGVGGGKKISWVNWKSVCQQKENGGVRVKDIRVMNISLLAKWRWRLIDGREALWK*VL 209
G GG+K V WK++ +GGV + D+ +MN + L K W G +LW VL
Sbjct: 8 GDNDGGRKHHVVGWKAMIMP*HSGGVGMCDLTIMNQACLMKIGWAFRMGDGSLWADVL 181
>BU090318 weakly similar to GP|9743345|gb| F21J9.30 {Arabidopsis thaliana},
partial (1%)
Length = 422
Score = 37.0 bits (84), Expect = 0.007
Identities = 14/34 (41%), Positives = 24/34 (70%)
Frame = +3
Query: 159 KISWVNWKSVCQQKENGGVRVKDIRVMNISLLAK 192
K+ W +WK++C K++GG+ ++ + MN SLL K
Sbjct: 3 KVHWTSWKALCSPKKSGGLGLRLLSHMNKSLLMK 104
>TC209287 similar to UP|HNM1_YEAST (P19807) Choline transport protein,
partial (4%)
Length = 1046
Score = 36.6 bits (83), Expect = 0.009
Identities = 16/32 (50%), Positives = 22/32 (68%)
Frame = +3
Query: 108 ISFGGRIVLLNSVLNSMPIFYLSFLKMLVGVW 139
I+F G I+ L+ V+NS+PI LSFL L +W
Sbjct: 729 IAFHGGILFLHGVINSLPISLLSFLGQLAAIW 824
>BQ297454
Length = 135
Score = 32.0 bits (71), Expect = 0.21
Identities = 17/42 (40%), Positives = 26/42 (61%), Gaps = 1/42 (2%)
Frame = -1
Query: 113 RIVLLNS-VLNSMPIFYLSFLKMLVGVWKRIVRIQR*FLWGG 153
R+ L+ S L S+ I++ SF ++ V ++ RIQR FLW G
Sbjct: 135 RVTLIPSWXLTSISIYFFSFFRVPKKVVDKLARIQRKFLWEG 10
>BF596367
Length = 419
Score = 29.3 bits (64), Expect = 1.4
Identities = 14/28 (50%), Positives = 20/28 (71%)
Frame = +1
Query: 182 IRVMNISLLAKWRWRLIDGREALWK*VL 209
I+ + ++LAK WRLID ++AL *VL
Sbjct: 64 IKAFHKAMLAKQMWRLIDNKDALVT*VL 147
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.330 0.142 0.465
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,292,133
Number of Sequences: 63676
Number of extensions: 168291
Number of successful extensions: 1135
Number of sequences better than 10.0: 47
Number of HSP's better than 10.0 without gapping: 1128
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1132
length of query: 211
length of database: 12,639,632
effective HSP length: 93
effective length of query: 118
effective length of database: 6,717,764
effective search space: 792696152
effective search space used: 792696152
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 57 (26.6 bits)
Medicago: description of AC148816.5


