

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148652.15 + phase: 0 /pseudo
(121 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
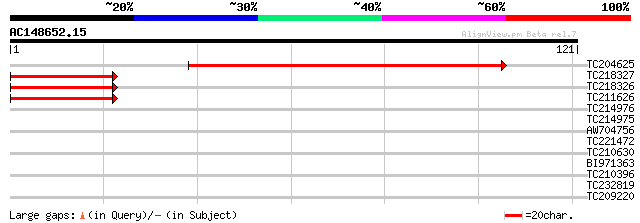
Score E
Sequences producing significant alignments: (bits) Value
TC204625 119 3e-28
TC218327 homologue to UP|Q9LH76 (Q9LH76) Arabidopsis thaliana ge... 43 2e-05
TC218326 homologue to UP|Q9LH76 (Q9LH76) Arabidopsis thaliana ge... 43 2e-05
TC211626 homologue to UP|Q9LH76 (Q9LH76) Arabidopsis thaliana ge... 43 2e-05
TC214976 similar to UP|Q9M0B6 (Q9M0B6) Nucleotide sugar epimeras... 36 0.003
TC214975 similar to UP|Q9M0B6 (Q9M0B6) Nucleotide sugar epimeras... 35 0.005
AW704756 30 0.16
TC221472 27 1.3
TC210630 similar to UP|Q8L819 (Q8L819) SET domain-containing pro... 25 5.1
BI971363 25 6.6
TC210396 similar to UP|Q75ZN6 (Q75ZN6) RNA-directed RNA Polymera... 25 6.6
TC232819 weakly similar to UP|Q9FZP2 (Q9FZP2) Receptor-like prot... 24 8.6
TC209220 similar to UP|Q9FKP4 (Q9FKP4) Similarity to kinesin hea... 24 8.6
>TC204625
Length = 2137
Score = 119 bits (297), Expect = 3e-28
Identities = 57/68 (83%), Positives = 62/68 (90%)
Frame = +1
Query: 39 QVKGFIHVSTDKFYGETDENAVVGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVI 98
Q+K FIHVSTD+ YGETDE+AVVGNHEASQLL TNPYSA K GAEMLVM+YGRSYGLPVI
Sbjct: 31 QIKRFIHVSTDEVYGETDEDAVVGNHEASQLLPTNPYSATKAGAEMLVMAYGRSYGLPVI 210
Query: 99 TTRGDRVY 106
TTRG+ VY
Sbjct: 211 TTRGNNVY 234
>TC218327 homologue to UP|Q9LH76 (Q9LH76) Arabidopsis thaliana genomic DNA,
chromosome 3, BAC clone: T21E2 (AT3g14790/T21E2_4),
partial (18%)
Length = 493
Score = 43.1 bits (100), Expect = 2e-05
Identities = 18/23 (78%), Positives = 21/23 (91%)
Frame = +1
Query: 1 MHFAAQTHVDNSYENSFEFTQTS 23
MHFAAQTHVDNS+ NSFEFT+ +
Sbjct: 394 MHFAAQTHVDNSFGNSFEFTKNN 462
>TC218326 homologue to UP|Q9LH76 (Q9LH76) Arabidopsis thaliana genomic DNA,
chromosome 3, BAC clone: T21E2 (AT3g14790/T21E2_4),
partial (18%)
Length = 854
Score = 43.1 bits (100), Expect = 2e-05
Identities = 18/23 (78%), Positives = 21/23 (91%)
Frame = +3
Query: 1 MHFAAQTHVDNSYENSFEFTQTS 23
MHFAAQTHVDNS+ NSFEFT+ +
Sbjct: 747 MHFAAQTHVDNSFGNSFEFTKNN 815
>TC211626 homologue to UP|Q9LH76 (Q9LH76) Arabidopsis thaliana genomic DNA,
chromosome 3, BAC clone: T21E2 (AT3g14790/T21E2_4),
partial (17%)
Length = 440
Score = 43.1 bits (100), Expect = 2e-05
Identities = 18/23 (78%), Positives = 21/23 (91%)
Frame = +1
Query: 1 MHFAAQTHVDNSYENSFEFTQTS 23
MHFAAQTHVDNS+ NSFEFT+ +
Sbjct: 355 MHFAAQTHVDNSFGNSFEFTKNN 423
>TC214976 similar to UP|Q9M0B6 (Q9M0B6) Nucleotide sugar epimerase-like
protein, partial (71%)
Length = 1356
Score = 35.8 bits (81), Expect = 0.003
Identities = 25/106 (23%), Positives = 43/106 (39%)
Frame = +1
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAV 60
MH AAQ V + EN + ++ L++ + N + S+ YG ++
Sbjct: 154 MHLAAQAGVRYAMENPHSYVHSNIAGLVTLLEACKTANPQPAIVWASSSSVYGLNEKVPF 333
Query: 61 VGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY 106
+ + Q + Y+A K E + +Y YGL + R VY
Sbjct: 334 SESDQTDQ--PASLYAATKKAGEEITHTYNHIYGLSITGLRFFTVY 465
>TC214975 similar to UP|Q9M0B6 (Q9M0B6) Nucleotide sugar epimerase-like
protein, partial (98%)
Length = 1828
Score = 35.0 bits (79), Expect = 0.005
Identities = 25/106 (23%), Positives = 43/106 (39%)
Frame = +3
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAV 60
MH AAQ V + EN + ++ L++ S N + S+ YG ++
Sbjct: 633 MHLAAQAGVRYAMENPHSYVHSNIAGLVTLLEACKSANPQPAVVWASSSSVYGLNEKVPF 812
Query: 61 VGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGLPVITTRGDRVY 106
+ + + + Y+A K E + +Y YGL + R VY
Sbjct: 813 SESDQTDR--PASLYAATKKAGEEITHTYNHIYGLSITGLRFFTVY 944
>AW704756
Length = 425
Score = 30.0 bits (66), Expect = 0.16
Identities = 22/95 (23%), Positives = 36/95 (37%)
Frame = +3
Query: 1 MHFAAQTHVDNSYENSFEFTQTSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAV 60
MH AAQ V + EN + ++ ++ + S N + S+ YG +
Sbjct: 105 MHLAAQAGVRYAMENPGSYVHSNIAGFVNLLEVCKSVNPQPAIVWASSSSVYGLNTKVPF 284
Query: 61 VGNHEASQLLETNPYSAMKIGAEMLVMSYGRSYGL 95
Q + Y+A K E + +Y YGL
Sbjct: 285 SERDRTDQ--PASLYAATKKAGEEIAHTYNHIYGL 383
>TC221472
Length = 640
Score = 26.9 bits (58), Expect = 1.3
Identities = 18/58 (31%), Positives = 30/58 (51%), Gaps = 1/58 (1%)
Frame = +2
Query: 5 AQTHVDNSYENSFEFTQTSFMELM-SFRSLQVSKNQVKGFIHVSTDKFYGETDENAVV 61
+QT VD+ T SF+ +M S + + +S +KGF+H G TD N+++
Sbjct: 374 SQTPVDD--------TAISFISMMPSLKDVDLSNTNIKGFLH------QGRTDVNSLL 505
>TC210630 similar to UP|Q8L819 (Q8L819) SET domain-containing protein SET102,
partial (14%)
Length = 460
Score = 25.0 bits (53), Expect = 5.1
Identities = 12/26 (46%), Positives = 17/26 (65%), Gaps = 2/26 (7%)
Frame = +3
Query: 84 MLVMSYGRSYGLPVITTRG--DRVYP 107
+L++ Y YGLP+ +TRG VYP
Sbjct: 108 LLLIPYTMKYGLPLKSTRGIAKTVYP 185
>BI971363
Length = 610
Score = 24.6 bits (52), Expect = 6.6
Identities = 13/29 (44%), Positives = 16/29 (54%)
Frame = -3
Query: 31 RSLQVSKNQVKGFIHVSTDKFYGETDENA 59
RSL+ N +GFIH+ST G D A
Sbjct: 128 RSLKTGPND*QGFIHLSTQYGNGS*DSVA 42
>TC210396 similar to UP|Q75ZN6 (Q75ZN6) RNA-directed RNA Polymerase
(Fragment), partial (82%)
Length = 919
Score = 24.6 bits (52), Expect = 6.6
Identities = 22/75 (29%), Positives = 39/75 (51%), Gaps = 3/75 (4%)
Frame = +3
Query: 22 TSFMELMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAVVGNHEASQLLETNPYSAMKIG 81
TSF +L++ R + +V GF+ D FY +T+ + +GN L++ Y +K
Sbjct: 228 TSFTKLVA-RDSYDHEMEVDGFMDYVDDAFYHKTNYDYKLGN-----LMD---YYGIKTE 380
Query: 82 AEML---VMSYGRSY 93
AE+L +M +S+
Sbjct: 381 AEILGGNIMKMSKSF 425
>TC232819 weakly similar to UP|Q9FZP2 (Q9FZP2) Receptor-like protein kinase,
partial (10%)
Length = 650
Score = 24.3 bits (51), Expect = 8.6
Identities = 14/32 (43%), Positives = 21/32 (64%), Gaps = 2/32 (6%)
Frame = -1
Query: 7 THVDN-SYENSFE-FTQTSFMELMSFRSLQVS 36
T DN + +N F+ F + MELMSFR++Q +
Sbjct: 401 TETDNLTLQNMFKVFLLWALMELMSFRNVQAA 306
>TC209220 similar to UP|Q9FKP4 (Q9FKP4) Similarity to kinesin heavy chain,
partial (19%)
Length = 961
Score = 24.3 bits (51), Expect = 8.6
Identities = 26/97 (26%), Positives = 39/97 (39%), Gaps = 5/97 (5%)
Frame = +3
Query: 27 LMSFRSLQVSKNQVKGFIHVSTDKFYGETDENAVVGN-----HEASQLLETNPYSAMKIG 81
L S SL QVK +ST ++ + V+G+ + S LLE + A +
Sbjct: 123 LASLISLDRILKQVKDISRLSTVNTIEKSKKRTVLGSLDKLTEQMSSLLEIDHPCARRYI 302
Query: 82 AEMLVMSYGRSYGLPVITTRGDRVYPFSHERKESIDS 118
A+ + I DR+ SH RK S D+
Sbjct: 303 ADA-------RRKVESIPEEDDRIQNLSHSRKPSTDT 392
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.317 0.130 0.380
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,271,869
Number of Sequences: 63676
Number of extensions: 38031
Number of successful extensions: 154
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 154
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 154
length of query: 121
length of database: 12,639,632
effective HSP length: 97
effective length of query: 24
effective length of database: 6,463,060
effective search space: 155113440
effective search space used: 155113440
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 51 (24.3 bits)
Medicago: description of AC148652.15


