

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148652.13 - phase: 0 /pseudo
(395 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
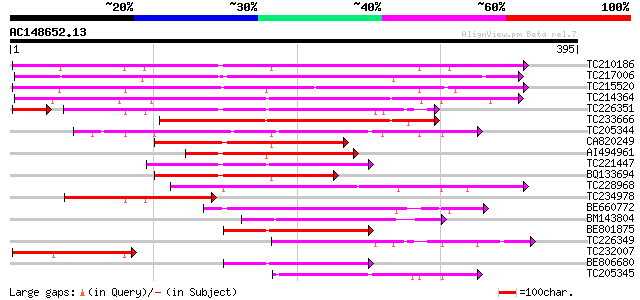
TC210186 UP|Q93WQ0 (Q93WQ0) Subtilisin-type protease, complete 295 3e-80
TC217006 UP|Q9ZTT3 (Q9ZTT3) Subtilisin-like protease C1, complete 285 3e-77
TC215520 GB|AAG38994.1|11611651|AF160513 subtilisin-type proteas... 269 1e-72
TC214364 UP|Q6WNU4 (Q6WNU4) Subtilisin-like protease, complete 253 1e-67
TC226351 similar to PIR|JC7519|JC7519 subtilisin-like serine pro... 187 2e-49
TC233666 weakly similar to UP|Q9FJF3 (Q9FJF3) Serine protease-li... 148 4e-36
TC205344 similar to UP|Q9SZV5 (Q9SZV5) Proteinase-like protein (... 133 2e-31
CA820249 similar to GP|13325079|gb| subtilisin-like protease C1 ... 129 2e-30
AI494961 similar to GP|14091078|gb subtilisin-like protein {Glyc... 129 3e-30
TC221447 similar to UP|Q75I27 (Q75I27) Putaive subtilisin-like p... 126 2e-29
BQ133694 weakly similar to GP|13325079|gb subtilisin-like protea... 125 3e-29
TC228968 similar to GB|BAB09208.1|9758669|AB012245 subtilisin-li... 121 5e-28
TC234978 similar to UP|P93204 (P93204) SBT1 protein (Subtilisin-... 115 4e-26
BE660772 106 2e-23
BM143804 similar to GP|29028287|gb| subtilase {Casuarina glauca}... 94 8e-20
BE801875 similar to PIR|T07172|T07 subtilisin-like proteinase (E... 90 2e-18
TC226349 similar to PIR|JC7519|JC7519 subtilisin-like serine pro... 82 4e-16
TC232007 similar to PIR|JC7519|JC7519 subtilisin-like serine pro... 81 7e-16
BE806680 80 2e-15
TC205345 similar to UP|Q9SZV5 (Q9SZV5) Proteinase-like protein (... 79 4e-15
>TC210186 UP|Q93WQ0 (Q93WQ0) Subtilisin-type protease, complete
Length = 2401
Score = 295 bits (754), Expect = 3e-80
Identities = 162/382 (42%), Positives = 229/382 (59%), Gaps = 23/382 (6%)
Frame = +1
Query: 3 GVISVFPSEEFHIQTTRSWDFLGLRQSIKRDQ----IIETDLVIGVIDIRIWPESESFND 58
GV+SVFP + TTRSWDFL + +K D + ++ VIG++D IWPE+ SF+D
Sbjct: 364 GVVSVFPGPVLKLHTTRSWDFLKYQTQVKIDTKPNAVSKSSSVIGILDTGIWPEAASFSD 543
Query: 59 KGLGPIPKKWRGVCAGGGNF---SCNNKIIGARFYGD----GDVSARYSFGHRTHTASTA 111
KG+GP+P +W+G C +F +CN K+IGAR+Y D GD +AR S GH TH A TA
Sbjct: 544 KGMGPVPSRWKGTCMKSQDFYSSNCNRKLIGARYYADPNDSGDNTARDSNGHGTHVAGTA 723
Query: 112 GGREVEDVSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAFDDAISDGVDV 171
G V + S+YG A G A+GG P SR+A Y++C N C G +ILAAFDDAI+DGVD+
Sbjct: 724 AGVMVTNASYYGVATGCAKGGSPESRLAVYRVCS---NFGCRGSSILAAFDDAIADGVDL 894
Query: 172 ITVSLGPEHA--SDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAPWLVSVAA 229
++VSLG D +DPI++G+FHAME GIL +AGN GP ++ + APW+++VAA
Sbjct: 895 LSVSLGASTGFRPDLTSDPISLGAFHAMEHGILVVCSAGNDGPSSYTLVNDAPWILTVAA 1074
Query: 230 TSIDRQFIDKVILGNGKTFVGKSINITP-SNGKKIPIAVRNAQACLAGGNASPEMC--DC 286
++IDR F+ ++LG+ K GK+IN++P SN K P+ + + C +
Sbjct: 1075STIDRNFLSNIVLGDNKIIKGKAINLSPLSNSPKYPLIYGESAKANSTSLVEARQCHPNS 1254
Query: 287 IDENMVKGKLVLCGSRNGK-------DLAYANGAIGSIHNVTKSQLGASFVTPRPSLNLK 339
+D N VKGK+V+C +N K A G IG +H +++ AS P+ +
Sbjct: 1255LDGNKVKGKIVVCDDKNDKYSTRKKVATVKAVGGIGLVHITDQNEAIASNYGDFPATVIS 1434
Query: 340 TNDFVHIQSYTNSSKYPVADFL 361
+ D V I Y NS+ PVA L
Sbjct: 1435SKDGVTILQYINSTSNPVATIL 1500
>TC217006 UP|Q9ZTT3 (Q9ZTT3) Subtilisin-like protease C1, complete
Length = 2422
Score = 285 bits (728), Expect = 3e-77
Identities = 165/367 (44%), Positives = 216/367 (57%), Gaps = 12/367 (3%)
Frame = +3
Query: 4 VISVFPSEEFHIQTTRSWDFLGLRQSIKRDQIIETDLVIGVIDIRIWPESESFNDKGLGP 63
V++VFP+++ + TTRSWDF+G R E+D++I V D IWPESESFNDKG GP
Sbjct: 312 VVAVFPNKKKQLHTTRSWDFIGFPLQANRAPA-ESDVIIAVFDSGIWPESESFNDKGFGP 488
Query: 64 IPKKWRGVCAGGGNFSCNNKIIGARFYG-------DGDVSARYSFGHRTHTASTAGGREV 116
P KW+G C NF+CNNKIIGA+ Y D S R GH TH ASTA G V
Sbjct: 489 PPSKWKGTCQTSKNFTCNNKIIGAKIYKVDGFFSKDDPKSVRDIDGHGTHVASTAAGNPV 668
Query: 117 EDVSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAFDDAISDGVDVITVSL 176
S G +GT+RGGV +RIA YK+C D C+ ILAAFDDAI+DGVD+ITVSL
Sbjct: 669 STASMLGLGQGTSRGGVTKARIAVYKVCW-FDG--CTDADILAAFDDAIADGVDIITVSL 839
Query: 177 GPEHASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAPWLVSVAATSIDRQF 236
G ++ D IAIG+FHA+ G+LT +AGN GP PSS+ + +PW +SVAA++IDR+F
Sbjct: 840 GGFSDENYFRDGIAIGAFHAVRNGVLTVTSAGNSGPRPSSLSNFSPWSISVAASTIDRKF 1019
Query: 237 IDKVILGNGKTFVGKSINITPSNGKKIPIA----VRNAQACLAGGNASPEMCDCIDENMV 292
+ KV LGN T+ G SIN G+ PI N + G ++ +D+ +V
Sbjct: 1020VTKVELGNKITYEGTSINTFDLKGELYPIIYGGDAPNKGEGIDGSSSRYCSSGSLDKKLV 1199
Query: 293 KGKLVLCGSRNGKDLAYANGAIGS-IHNVTKSQLGASFVTPRPSLNLKTNDFVHIQSYTN 351
KGK+VLC SR+ + GA+G+ I L S P P L D + Y N
Sbjct: 1200KGKIVLCESRSKALGPFDAGAVGALIQGQGFRDLPPSL--PLPGSYLALQDGASVYDYIN 1373
Query: 352 SSKYPVA 358
S++ P+A
Sbjct: 1374STRTPIA 1394
>TC215520 GB|AAG38994.1|11611651|AF160513 subtilisin-type protease precursor
{Glycine max;} , complete
Length = 2392
Score = 269 bits (688), Expect = 1e-72
Identities = 160/388 (41%), Positives = 216/388 (55%), Gaps = 29/388 (7%)
Frame = +2
Query: 3 GVISVFPSEEFHIQTTRSWDFLGLRQSIKRD---------QIIETDLVIGVIDIRIWPES 53
GV+SVFP + TTRSWDFL + + D +D+++GV+D IWPE+
Sbjct: 332 GVVSVFPDPILKLHTTRSWDFLKSQTRVNIDTKPNTLSGSSFSSSDVILGVLDTGIWPEA 511
Query: 54 ESFNDKGLGPIPKKWRGVCAGGGNFS---CNNKIIGARFYGDGDVSARYSF-GHRTHTAS 109
SF+DKG GP+P +W+G C +F+ CN KIIGARFY + + F GH TH +S
Sbjct: 512 ASFSDKGFGPVPSRWKGTCMTSKDFNSSCCNRKIIGARFYPNPEEKTARDFNGHGTHVSS 691
Query: 110 TAGGREVEDVSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAFDDAISDGV 169
TA G V SFYG A GTARGG P SR+A YK+CG G C G AILA FDDAI DGV
Sbjct: 692 TAVGVPVSGASFYGLAAGTARGGSPESRLAVYKVCGAF--GSCPGSAILAGFDDAIHDGV 865
Query: 170 DVITVSLG--PEHASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAPWLVSV 227
D++++SLG +D DPIAIG+FH++++GIL AAGN G P +V + APW+++V
Sbjct: 866 DILSLSLGGFGGTKTDLTTDPIAIGAFHSVQRGILVVCAAGNDGE-PFTVLNDAPWILTV 1042
Query: 228 AATSIDRQFIDKVILGNGKTFVGKSINITP-SNGKKIP-IAVRNAQACLAGGNASPEMC- 284
AA++IDR V+LGN + G++IN +P N P I +A C
Sbjct: 1043AASTIDRDLQSDVVLGNNQVVKGRAINFSPLLNSPDYPMIYAESAARANISNITDARQCH 1222
Query: 285 -DCIDENMVKGKLVLCGSRNGKDLAY----------ANGAIGSIHNVTKSQLGASFVTPR 333
D +D V GK+V+C +N D+ Y A G IG +H +S A +
Sbjct: 1223PDSLDPKKVIGKIVVCDGKN--DIYYSTDEKIVIVKALGGIGLVHITDQSGSVAFYYVDF 1396
Query: 334 PSLNLKTNDFVHIQSYTNSSKYPVADFL 361
P +K+ I Y NS+ +PV L
Sbjct: 1397PVTEVKSKHGDAILQYINSTSHPVGTIL 1480
>TC214364 UP|Q6WNU4 (Q6WNU4) Subtilisin-like protease, complete
Length = 2667
Score = 253 bits (645), Expect = 1e-67
Identities = 158/386 (40%), Positives = 220/386 (56%), Gaps = 31/386 (8%)
Frame = +1
Query: 4 VISVFPSEEFHIQTTRSWDFLGLRQ-------SIKRDQIIETDLVIGVIDIRIWPESESF 56
V+SVF + + TTRSWDF+ L SI + ++IG +D +WPES+SF
Sbjct: 343 VLSVFENRGRKLHTTRSWDFMELEHNGVIQSSSIWKKARFGEGVIIGNLDTGVWPESKSF 522
Query: 57 NDKGLGPIPKKWRGVCAGG--GNFSCNNKIIGARFYGDGDVSA-----------RYSFGH 103
+++GLGPIP KWRG+C G F CN K+IGAR++ G S R + GH
Sbjct: 523 SEQGLGPIPSKWRGICHNGIDHTFHCNRKLIGARYFNKGYASVAGPLNSSFDSPRDNEGH 702
Query: 104 RTHTASTAGGREVEDVSFYGFAKGTARGGVPSSRIAAYKICGD-IDNGRCSGDAILAAFD 162
THT STAGG V VS +G GTA+GG P +R+AAYK+C + C ILAAFD
Sbjct: 703 GTHTLSTAGGNMVARVSVFGQGHGTAKGGSPMARVAAYKVCWPPVAGDECFDADILAAFD 882
Query: 163 DAISDGVDVITVSLGPEHASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAP 222
AI DGVDV++VSLG +S F D +AIGSFHA ++G++ +AGN GP ++ + AP
Sbjct: 883 LAIHDGVDVLSVSLGGS-SSTFFKDSVAIGSFHAAKRGVVVVCSAGNSGPAEATAENLAP 1059
Query: 223 WLVSVAATSIDRQFIDKVILGNGKTFVGKSINITPSNGKKIPIAVRNAQACLAGGNASPE 282
W V+VAA+++DRQF V+LGN TF G+S++ T K PI ++ A LA A
Sbjct: 1060WHVTVAASTMDRQFPTYVVLGNDITFKGESLSATKLAHKFYPI-IKATDAKLASARAEDA 1236
Query: 283 -MCD--CIDENMVKGKLVLC----GSRNGK-DLAYANGAIGSIHNVTKSQLGASFVTPR- 333
+C +D N KGK+V+C +R K + A+ GA+G + K+ P
Sbjct: 1237VLCQNGTLDPNKAKGKIVVCLRGINARVDKGEQAFLAGAVGMVLANDKTTGNEIIADPHV 1416
Query: 334 -PSLNLKTNDFVHIQSYTNSSKYPVA 358
P+ ++ D + Y NS+K+PVA
Sbjct: 1417LPASHINFTDGSAVFKYINSTKFPVA 1494
>TC226351 similar to PIR|JC7519|JC7519 subtilisin-like serine proteinase -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(48%)
Length = 1490
Score = 187 bits (474), Expect(2) = 2e-49
Identities = 117/288 (40%), Positives = 158/288 (54%), Gaps = 26/288 (9%)
Frame = +3
Query: 38 TDLVIGVIDI-RIWPESESFNDKGLGPIPKKWRGVCAGGGNFS---CNNKIIGARFYGDG 93
+D++IGV++ R+ ++ + + GLGP+P W+G C G N + CN K+IGARF+ G
Sbjct: 588 SDVIIGVLNTWRLAGKAGASTNTGLGPVPSTWKGACETGTNLTASNCNRKLIGARFFSKG 767
Query: 94 -------------DVSARYSFGHRTHTASTAGGREVEDVSFYGFAKGTARGGVPSSRIAA 140
SAR GH THTASTA G V D S +G+A GTARG +R+AA
Sbjct: 768 VEAILGPINETEESRSARDDDGHGTHTASTAAGSVVSDASLFGYASGTARGMATRARVAA 947
Query: 141 YKICGDIDNGRCSGDAILAAFDDAISDGVDVITVSLGPEHASDFLNDPIAIGSFHAMEKG 200
YK+C G C ILAA + AI D V+V+++SLG SD+ D +AIG+F AME G
Sbjct: 948 YKVCW---KGGCFSSDILAAIERAILDNVNVLSLSLGGG-MSDYYRDSVAIGAFSAMENG 1115
Query: 201 ILTTQAAGNFGPIPSSVCSGAPWLVSVAATSIDRQFIDKVILGNGKTFVGKSI---NITP 257
IL + +AGN GP P S+ + APW+ +V A ++DR F V LGNG F G S+ N P
Sbjct: 1116ILVSCSAGNAGPSPYSLSNVAPWITTVGAGTLDRDFPAYVALGNGLNFSGVSLYRGNAVP 1295
Query: 258 SN------GKKIPIAVRNAQACLAGGNASPEMCDCIDENMVKGKLVLC 299
+ + N C+ G SPE V GK+VLC
Sbjct: 1296DSPLPFVYAGNVSNGAMNGNLCIT-GTLSPE--------KVAGKIVLC 1412
Score = 27.3 bits (59), Expect(2) = 2e-49
Identities = 11/27 (40%), Positives = 18/27 (65%)
Frame = +2
Query: 3 GVISVFPSEEFHIQTTRSWDFLGLRQS 29
G+++V P + + TTR+ FLGL +S
Sbjct: 476 GILAVLPETRYELHTTRTPMFLGLDKS 556
>TC233666 weakly similar to UP|Q9FJF3 (Q9FJF3) Serine protease-like protein,
partial (30%)
Length = 678
Score = 148 bits (374), Expect = 4e-36
Identities = 84/199 (42%), Positives = 123/199 (61%), Gaps = 4/199 (2%)
Frame = +1
Query: 105 THTASTAGGREVEDVSFYGFAKGTARGGVPSSRIAAYKICGD-IDNGRCSGDAILAAFDD 163
+HT ST GG V + +G GTA GG P +R+A YK+C ID C I+AAFD
Sbjct: 7 SHTLSTIGGTFVPGANVFGLGNGTAEGGSPRARVATYKVCWPPIDGNECFDADIMAAFDM 186
Query: 164 AISDGVDVITVSLGPEHASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAPW 223
AI DGVDV+++SLG +A+D+ +D ++IG+FHA KGI +AGN+GP P++V + APW
Sbjct: 187 AIHDGVDVLSLSLGG-NATDYFDDGLSIGAFHANMKGIPVICSAGNYGPTPATVFNVAPW 363
Query: 224 LVSVAATSIDRQFIDKVILGNGKTFVGKSINITPSNGKKIPIAVRNAQACLAG---GNAS 280
+++V A+++DRQF V L NG+ F+G S++ K P+ + A A A NA+
Sbjct: 364 ILTVGASTLDRQFDSVVELHNGQRFMGASLSKAMPEDKLYPL-INAADAKAANKPVENAT 540
Query: 281 PEMCDCIDENMVKGKLVLC 299
M ID +GK+++C
Sbjct: 541 LCMRGTIDPEKARGKILVC 597
>TC205344 similar to UP|Q9SZV5 (Q9SZV5) Proteinase-like protein (AT4g30020
protein) (AT4g30020/F6G3_50), partial (78%)
Length = 2192
Score = 133 bits (334), Expect = 2e-31
Identities = 101/325 (31%), Positives = 156/325 (47%), Gaps = 40/325 (12%)
Frame = +2
Query: 45 IDIRIWPESESF---NDKGLGPIPKKWRGVCAGGGNFS---CNNKIIGARFYGDGDVSAR 98
+D I+P SF N + GP+ ++RG C + CN KI+GA+ + ++A
Sbjct: 8 VDSGIYPHHPSFTTHNTEPYGPV-SRYRGKCEVDPDTKKSFCNGKIVGAQHFAQAAIAAG 184
Query: 99 Y------------SFGHRTHTASTAGGREVEDVSFYGFAKGTARGGVPSSRIAAYKICGD 146
GH +HTAS A GR V +G G A G P +RIA YK
Sbjct: 185 AFNPFIVFDSPLDGDGHGSHTASIAAGRNGIPVRMHGHEFGKASGMAPRARIAVYKALYR 364
Query: 147 IDNGRCSGDAILAAFDDAISDGVDVITVSLGPEHA-----SDFLNDPIAIGSFHAMEKGI 201
+ G + ++AA D A+ DGVD++++S+GP + FLN P A++ G+
Sbjct: 365 LFGGFIAD--VVAAIDQAVHDGVDILSLSVGPNSPPSNTKTTFLN-PFDATLLGAVKAGV 535
Query: 202 LTTQAAGNFGPIPSSVCSGAPWLVSVAATSIDRQFIDKVILGNGKTFVGKSINITPS--- 258
QAAGN GP P S+ S +PW+ +VAA DR++ + +ILGNGK G + ++PS
Sbjct: 536 FVAQAAGNGGPFPKSLVSYSPWIATVAAAIDDRRYKNHLILGNGKILAG--LGLSPSTRL 709
Query: 259 NGKKIPIAVRNAQACLAGGNASPEMC---DCIDENMVKGKLVLCG-----------SRNG 304
N +A + + SP C + +++N++KG ++LCG +
Sbjct: 710 NQTYTLVAATDVLLDSSVTKYSPTDCQRPELLNKNLIKGNILLCGYSYNFVIGSASIKQV 889
Query: 305 KDLAYANGAIGSIHNVTKSQLGASF 329
+ A A GA+G + V G F
Sbjct: 890 SETAKALGAVGFVLYVENVSPGTKF 964
>CA820249 similar to GP|13325079|gb| subtilisin-like protease C1 {Glycine
max}, partial (15%)
Length = 421
Score = 129 bits (325), Expect = 2e-30
Identities = 70/138 (50%), Positives = 91/138 (65%), Gaps = 3/138 (2%)
Frame = +2
Query: 102 GHRTHTASTAGGREVEDVSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAF 161
GH +H ASTA G V S GF GTARGGVPS+RIA YK+C C IL A+
Sbjct: 14 GHGSHCASTAAGNPVRSASLLGFGSGTARGGVPSARIAVYKVCWATG---CDTTDILKAY 184
Query: 162 DDAISDGVDVITVSLGPEHA--SDFLNDPIAIGSFHAMEKGILTTQAAGNFGPI-PSSVC 218
D AI+DGVD+++VS+G + + D AIG+FHAM+KGILT+ +A N G + P S
Sbjct: 185 DAAIADGVDILSVSVGATQLTHNKYFKDVHAIGAFHAMKKGILTSTSADNLGQLGPYSTS 364
Query: 219 SGAPWLVSVAATSIDRQF 236
APWL+SVAA++ID++F
Sbjct: 365 KFAPWLLSVAASTIDKKF 418
>AI494961 similar to GP|14091078|gb subtilisin-like protein {Glycine max},
partial (15%)
Length = 374
Score = 129 bits (323), Expect = 3e-30
Identities = 66/123 (53%), Positives = 86/123 (69%), Gaps = 2/123 (1%)
Frame = +2
Query: 123 GFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAFDDAISDGVDVITVSLG--PEH 180
G A G+A GG SR+A Y++C N C G AIL AFDDAISDGVDV+++SLG P
Sbjct: 2 GLAAGSATGGSSESRLAVYRVCS---NFGCRGSAILGAFDDAISDGVDVLSLSLGASPGF 172
Query: 181 ASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAPWLVSVAATSIDRQFIDKV 240
D DPIA+G+FHA+E+GIL +AGN GP S+V + APW+++VAA++IDR F V
Sbjct: 173 QPDLTTDPIALGAFHAVERGILVVCSAGNSGPSSSTVVNDAPWILTVAASTIDRDFQSDV 352
Query: 241 ILG 243
+LG
Sbjct: 353 VLG 361
>TC221447 similar to UP|Q75I27 (Q75I27) Putaive subtilisin-like proteinase,
partial (21%)
Length = 491
Score = 126 bits (316), Expect = 2e-29
Identities = 69/158 (43%), Positives = 95/158 (59%)
Frame = +1
Query: 96 SARYSFGHRTHTASTAGGREVEDVSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGD 155
S R GH +HT++TA G V S +GFA GTARG +R+A YK+C G C
Sbjct: 13 SPRDDDGHGSHTSTTAAGSAVVGASLFGFANGTARGMATQARLATYKVCW---LGGCFTS 183
Query: 156 AILAAFDDAISDGVDVITVSLGPEHASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPS 215
I A D AI DGV+++++S+G D+ D IAIG+F A GIL + +AGN GP +
Sbjct: 184 DIAAGIDKAIEDGVNILSMSIGGG-LMDYYKDTIAIGTFAATAHGILVSNSAGNGGPSQA 360
Query: 216 SVCSGAPWLVSVAATSIDRQFIDKVILGNGKTFVGKSI 253
++ + APWL +V A +IDR F + LGNGK + G S+
Sbjct: 361 TLSNVAPWLTTVGAGTIDRDFPAYITLGNGKMYTGVSL 474
>BQ133694 weakly similar to GP|13325079|gb subtilisin-like protease C1
{Glycine max}, partial (17%)
Length = 420
Score = 125 bits (315), Expect = 3e-29
Identities = 66/130 (50%), Positives = 85/130 (64%), Gaps = 2/130 (1%)
Frame = +2
Query: 102 GHRTHTASTAGGREVEDVSFYGFAKGTARGGVPSSRIAAYKICGDIDNGRCSGDAILAAF 161
GH +H ASTA G V S YG GTARGGVP +RIA YK+C C ILAAF
Sbjct: 38 GHGSHCASTAAGNPVRSASLYGLGLGTARGGVPLARIAVYKVCW---TKGCHDADILAAF 208
Query: 162 DDAISDGVDVITVSLGPEHA--SDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCS 219
D+AI DGVD+I++S+GP + + AIG+FHAM++GILT++ A N GP P ++ +
Sbjct: 209 DEAIRDGVDIISISVGPTIVLHLHYFEEVYAIGAFHAMKQGILTSKTADNLGPKPYTMSN 388
Query: 220 GAPWLVSVAA 229
APW +SVAA
Sbjct: 389 YAPWYLSVAA 418
>TC228968 similar to GB|BAB09208.1|9758669|AB012245 subtilisin-like protease
{Arabidopsis thaliana;} , partial (20%)
Length = 1008
Score = 121 bits (304), Expect = 5e-28
Identities = 87/265 (32%), Positives = 133/265 (49%), Gaps = 16/265 (6%)
Frame = +3
Query: 113 GREVEDVSFYG-FAKGTARGGVPSSRIAAYKICGDI------DNGRCSGDAILAAFDDAI 165
GR V + S G FAKGTA GG P +R+A YK C I + C+ +L A DDAI
Sbjct: 12 GRVVPNASAIGGFAKGTALGGAPLARLAIYKACWPIKGKSKHEGNICTNIDMLKAIDDAI 191
Query: 166 SDGVDVITVSLGPEHASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAPWLV 225
DGVDV+++S+G + D IA G+ HA+ K I+ +AGN GP+P ++ + APW++
Sbjct: 192 GDGVDVLSISIGFSAPISYEEDVIARGALHAVRKNIVVVCSAGNSGPLPQTLSNPAPWII 371
Query: 226 SVAATSIDRQFIDKVILGNGKTFVGKSINITPSNGKKIPIAVRN--AQACLAGGNASPEM 283
+VAA+++DR F + L NG G+SI P+ + L N+ +
Sbjct: 372 TVAASTVDRSFHAPIKL-NGTIIEGRSITPLHMGNSFYPLVLARDVEHPGLPSNNSGFCL 548
Query: 284 CDCIDENMVKGKLVLC----GSRNGKDLAYAN-GAIGSI--HNVTKSQLGASFVTPRPSL 336
+ + N +GK+VLC G R K L G +G I +N + S P+
Sbjct: 549 DNTLQPNKARGKIVLCMRGQGERLKKGLEVQRAGGVGFILGNNKLNGKDVPSDPHFIPAT 728
Query: 337 NLKTNDFVHIQSYTNSSKYPVADFL 361
+ + + + Y +S+ P+A L
Sbjct: 729 GVSYENSLKLIQYVHSTPNPMAQIL 803
>TC234978 similar to UP|P93204 (P93204) SBT1 protein (Subtilisin-like
protease) , partial (15%)
Length = 422
Score = 115 bits (287), Expect = 4e-26
Identities = 58/122 (47%), Positives = 75/122 (60%), Gaps = 16/122 (13%)
Frame = +3
Query: 39 DLVIGVIDIRIWPESESFNDKGLGPIPKKWRGVCAGGGNFS---CNNKIIGARFYGDG-- 93
D++IGV+D +WPES SF+D G+ IP +WRG C G +FS CN K+IGAR + G
Sbjct: 36 DVIIGVLDTGVWPESPSFDDAGMPEIPARWRGECETGPDFSPKMCNRKLIGARSFSKGFH 215
Query: 94 -----------DVSARYSFGHRTHTASTAGGREVEDVSFYGFAKGTARGGVPSSRIAAYK 142
SAR GH THT+STA G V + S G+A GTARG P++R+AAYK
Sbjct: 216 MASGIGVREKEPASARDRDGHGTHTSSTAAGSHVTNASLLGYASGTARGMAPTARVAAYK 395
Query: 143 IC 144
+C
Sbjct: 396 VC 401
>BE660772
Length = 826
Score = 106 bits (265), Expect = 2e-23
Identities = 74/209 (35%), Positives = 113/209 (53%), Gaps = 11/209 (5%)
Frame = +2
Query: 136 SRIAAYKICGDIDNGRCSGDAILAAFDDAISDGVDVITVSLGPE-HASDFLNDPIAIGSF 194
+RIAAYKIC + C ILAA D+A+SDGV VI++S+G +A + D IA+G+F
Sbjct: 5 ARIAAYKICWKLG---CFDSDILAAMDEAVSDGVHVISLSVGSSGYAPQYYRDSIAVGAF 175
Query: 195 HAMEKGILTTQAAGNFGPIPSSVCSGAPWLVSVAATSIDRQFIDKVILGNGKTFVGKSIN 254
A + +L + +AGN GP PS+ + APW+++V A+++DR+F VILG+G+ F G S+
Sbjct: 176 GAAKHNVLVSCSAGNSGPGPSTAVNIAPWILTVGASTVDREFPADVILGDGRVFGGVSLY 355
Query: 255 ITPS-NGKKIPIAVR---NAQACLAGGNASPEMCDCIDENMVKGKLVLCGSRNGK----- 305
S K+P+ ++ C G ++ + V+GK+ +C R G
Sbjct: 356 YGESLPDFKLPLVYAKDCGSRYCYIGS---------LESSKVQGKIXVC-DRGGNARVEK 505
Query: 306 -DLAYANGAIGSIHNVTKSQLGASFVTPR 333
G +G I T SQ SF+ R
Sbjct: 506 GSAVKLTGGLGMIMGNT*SQR*KSFLQMR 592
>BM143804 similar to GP|29028287|gb| subtilase {Casuarina glauca}, partial
(11%)
Length = 408
Score = 94.4 bits (233), Expect = 8e-20
Identities = 54/144 (37%), Positives = 80/144 (55%), Gaps = 1/144 (0%)
Frame = +2
Query: 162 DDAISDGVDVITVSLGPEHASDFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGA 221
D AISDGVDV+++S G + + DP+AI +F AMEKGI + +AGN GP + +G
Sbjct: 2 DSAISDGVDVLSLSFGFDDVPLY-EDPVAIATFSAMEKGIFVSTSAGNEGPFLGRLHNGI 178
Query: 222 PWLVSVAATSIDRQFIDKVILGNGKTFVGKSINITPSNGKKIPIAVRNAQACLAGGNASP 281
PW+++VAA ++DR+F + LGNG G S+ + +PI
Sbjct: 179 PWVITVAAGTLDREFHGTLTLGNGVQITGMSLYHGNFSSSNVPIVFMG------------ 322
Query: 282 EMCDCIDE-NMVKGKLVLCGSRNG 304
+CD + E VK K+V+C +NG
Sbjct: 323 -LCDNVKELAKVKSKIVVCEDKNG 391
>BE801875 similar to PIR|T07172|T07 subtilisin-like proteinase (EC 3.4.21.-)
2 - tomato, partial (15%)
Length = 405
Score = 89.7 bits (221), Expect = 2e-18
Identities = 44/104 (42%), Positives = 66/104 (63%)
Frame = +2
Query: 150 GRCSGDAILAAFDDAISDGVDVITVSLGPEHASDFLNDPIAIGSFHAMEKGILTTQAAGN 209
G C IL+A D A+ DGVDV+++SLG S + D +++ SF AMEKG+ + + GN
Sbjct: 5 GGCFSSDILSAVDRAVDDGVDVLSISLGGG-VSSYYRDSLSVASFGAMEKGVFVSCSGGN 181
Query: 210 FGPIPSSVCSGAPWLVSVAATSIDRQFIDKVILGNGKTFVGKSI 253
GP P S+ + +PW+ +V A+++DR F V LGNG+ G S+
Sbjct: 182 AGPDPVSLTNVSPWITTVGASTMDRDFPADVSLGNGRKITGTSL 313
>TC226349 similar to PIR|JC7519|JC7519 subtilisin-like serine proteinase -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(56%)
Length = 1773
Score = 82.0 bits (201), Expect = 4e-16
Identities = 66/202 (32%), Positives = 95/202 (46%), Gaps = 18/202 (8%)
Frame = +1
Query: 183 DFLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAPWLVSVAATSIDRQFIDKVIL 242
D+ D +AIG+F AMEKGIL + +AGN GP P S+ + APW+ +V A ++DR F V L
Sbjct: 1 DYYRDSVAIGAFSAMEKGILVSCSAGNSGPGPYSLSNVAPWITTVGAGTLDRDFPAYVAL 180
Query: 243 GNGKTFVGKSI---NITPSNGKKIPIA------VRNAQACLAGGNASPEMCDCIDENMVK 293
GNG F G S+ N P + + A N C+ G SPE V
Sbjct: 181 GNGLNFSGVSLYRGNALPDSSLPLVYAGNVSNGAMNGNLCIT-GTLSPE--------KVA 333
Query: 294 GKLVLCG----SRNGK-DLAYANGAIGSIHNVTKSQ----LGASFVTPRPSLNLKTNDFV 344
GK+VLC +R K + + GA+G + + T + + + + P ++ K D
Sbjct: 334 GKIVLCDRGLTARVQKGSVVKSAGALGMVLSNTAANGEELVADAHLLPATAVGQKAGD-- 507
Query: 345 HIQSYTNSSKYPVADFLFSWPK 366
I+ Y S P F K
Sbjct: 508 AIKKYLVSDAKPTVKIFFEGTK 573
>TC232007 similar to PIR|JC7519|JC7519 subtilisin-like serine proteinase -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(20%)
Length = 669
Score = 81.3 bits (199), Expect = 7e-16
Identities = 37/91 (40%), Positives = 60/91 (65%), Gaps = 5/91 (5%)
Frame = +3
Query: 3 GVISVFPSEEFHIQTTRSWDFLGLRQS--IKRDQIIETDLVIGVIDIRIWPESESFNDKG 60
G+++V P + + TTR+ FLGL +S + + +D+++GV+D +WPES+SF+D G
Sbjct: 309 GILAVLPETRYELFTTRTPLFLGLDKSADLFPESSSGSDVIVGVLDTGVWPESKSFDDTG 488
Query: 61 LGPIPKKWRGVCAGGGNF---SCNNKIIGAR 88
LGP+P W+G C G NF +CN K+I ++
Sbjct: 489 LGPVPSTWKGACQTGTNFTASNCNRKVIRSK 581
>BE806680
Length = 383
Score = 80.1 bits (196), Expect = 2e-15
Identities = 44/104 (42%), Positives = 61/104 (58%)
Frame = +1
Query: 150 GRCSGDAILAAFDDAISDGVDVITVSLGPEHASDFLNDPIAIGSFHAMEKGILTTQAAGN 209
G C I A D AI DGV+V+++S+G ++ D IAIGSF AM GIL + + N
Sbjct: 19 GGCFTSDIAAGIDKAIEDGVNVLSMSIGGS-LMEYYRDIIAIGSFTAMSHGILVSTSTVN 195
Query: 210 FGPIPSSVCSGAPWLVSVAATSIDRQFIDKVILGNGKTFVGKSI 253
GP S+ + APW+ +V A +IDR + LG GKT+ G S+
Sbjct: 196 GGPSQGSLSNVAPWITTVGAGTIDRDCPAYITLGTGKTYTGASL 327
>TC205345 similar to UP|Q9SZV5 (Q9SZV5) Proteinase-like protein (AT4g30020
protein) (AT4g30020/F6G3_50), partial (62%)
Length = 1729
Score = 79.0 bits (193), Expect = 4e-15
Identities = 54/163 (33%), Positives = 82/163 (50%), Gaps = 17/163 (10%)
Frame = +1
Query: 184 FLNDPIAIGSFHAMEKGILTTQAAGNFGPIPSSVCSGAPWLVSVAATSIDRQFIDKVILG 243
FLN P A++ G+ QAAGN GP+P ++ S +PW+ SVAA DR++ + +ILG
Sbjct: 25 FLN-PFDATLLGAVKAGVFVAQAAGNHGPLPKTLVSYSPWIASVAAAIDDRRYKNHLILG 201
Query: 244 NGKTFVGKSINITPSNGKKIPIAVRNAQACLAGGNA---SPEMC---DCIDENMVKGKLV 297
NGKT G I ++PS + A L + SP C + +++N++KG ++
Sbjct: 202 NGKTLAG--IGLSPSTHLNETYTLVAANDVLLDSSLMKYSPTDCQRPELLNKNLIKGNIL 375
Query: 298 LCG-----------SRNGKDLAYANGAIGSIHNVTKSQLGASF 329
LCG + + A A GA+G + V LG F
Sbjct: 376 LCGYSFNFVVGTASIKKVSETAKALGAVGFVLCVENISLGTKF 504
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.137 0.417
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,327,398
Number of Sequences: 63676
Number of extensions: 240762
Number of successful extensions: 1144
Number of sequences better than 10.0: 64
Number of HSP's better than 10.0 without gapping: 1079
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1083
length of query: 395
length of database: 12,639,632
effective HSP length: 99
effective length of query: 296
effective length of database: 6,335,708
effective search space: 1875369568
effective search space used: 1875369568
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC148652.13


