

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148651.5 + phase: 0
(496 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
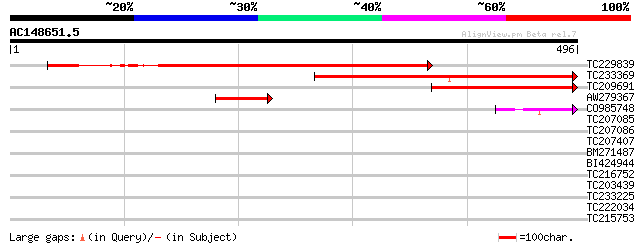
Score E
Sequences producing significant alignments: (bits) Value
TC229839 similar to GB|AAO64872.1|29028986|BT005937 At1g22882 {A... 384 e-107
TC233369 similar to GB|AAO64872.1|29028986|BT005937 At1g22882 {A... 313 1e-85
TC209691 200 1e-51
AW279367 similar to GP|15290067|dbj P0025A05.23 {Oryza sativa (j... 72 4e-13
CO985748 42 5e-04
TC207085 similar to UP|Q9SH20 (Q9SH20) F28K19.1, partial (22%) 32 0.50
TC207086 32 0.65
TC207407 UP|Q6T2Z3 (Q6T2Z3) Cyclin-dependent kinase inhibitor 1;... 30 2.5
BM271487 similar to GP|15810167|gb At1g64110/F22C12_22 {Arabidop... 29 4.2
BI424944 29 5.5
TC216752 similar to UP|Q8LA49 (Q8LA49) Globulin-like protein, pa... 29 5.5
TC203439 weakly similar to UP|Q9LX02 (Q9LX02) ESTs AU082316(E336... 28 7.2
TC233225 28 9.4
TC222034 weakly similar to UP|Q6H5X3 (Q6H5X3) ABA-responsive pro... 28 9.4
TC215753 similar to UP|LSM3_HUMAN (P62310) U6 snRNA-associated S... 28 9.4
>TC229839 similar to GB|AAO64872.1|29028986|BT005937 At1g22882 {Arabidopsis
thaliana;} , partial (28%)
Length = 1063
Score = 384 bits (986), Expect = e-107
Identities = 216/339 (63%), Positives = 244/339 (71%), Gaps = 2/339 (0%)
Frame = +1
Query: 34 SVGLSKWNEVNHGFCEISDTADKYFIKEIDACFPSEALIYSKAGDAEANGLVNESHNGRE 93
SV LS W E H C+ S++A+K +KE +H+
Sbjct: 193 SVRLSNWKEDKHRQCKTSNSANKCLLKE--------------------------THD--- 285
Query: 94 SGAYAVPADINKENTDSANREDHVVENSEYAVKHENDVKKSDILSRAVPLGLNEFKSRAI 153
Y +P KE+ D+ VV D + SD L AVPLGL+EFKSRAI
Sbjct: 286 ---YILPN--YKEDCDTPT----VV-----------DAQMSDHLPWAVPLGLDEFKSRAI 405
Query: 154 SSKVKSGT-GQSRSVIHRLEPGGAEYNYASASKGAKVLGSNKEGKGASNILSRDKDKYLR 212
SSK+KSGT G S SV+HR+EPGGAEYNYASAS GAK+LGSNKE KGASNILSRDKDKYLR
Sbjct: 406 SSKIKSGTSGSSGSVMHRVEPGGAEYNYASASMGAKLLGSNKEAKGASNILSRDKDKYLR 585
Query: 213 NPCSVVGKFVIMELSEETLVDTIEIANFEHHSSNLKDFEIHGSLNFPTNVWDLLGNFTAS 272
NPCS KFVI+ELSEETLVDTIEIANFEHHSSNLK FE+ GSL+FPT+VW LGNFTAS
Sbjct: 586 NPCSAEDKFVIIELSEETLVDTIEIANFEHHSSNLKAFELLGSLSFPTDVWVFLGNFTAS 765
Query: 273 NVRHAQRFVLKEPKWVRYLKLNLQSHYGSEFYCTLSVVEVFGVDAVERMLEDLINTQDNL 332
NVRHAQRFVL++PKWVRYLKLNLQSHYGSEFYCTLSVVEV+GVDAVERMLEDLI+TQDNL
Sbjct: 766 NVRHAQRFVLQQPKWVRYLKLNLQSHYGSEFYCTLSVVEVYGVDAVERMLEDLIHTQDNL 945
Query: 333 LASGEGNADK-TILPHPDPAVIEHVHKKPLEGINSVPAS 370
LA G+GNADK T+ PHP+P E H+ GINS PAS
Sbjct: 946 LAPGDGNADKMTVSPHPNPPESEDAHQNTFGGINSYPAS 1062
>TC233369 similar to GB|AAO64872.1|29028986|BT005937 At1g22882 {Arabidopsis
thaliana;} , partial (16%)
Length = 945
Score = 313 bits (802), Expect = 1e-85
Identities = 158/233 (67%), Positives = 190/233 (80%), Gaps = 3/233 (1%)
Frame = +1
Query: 267 GNFTASNVRHAQRFVLKEPKWVRYLKLNLQSHYGSEFYCTLSVVEVFGVDAVERMLEDLI 326
GNFTASNV+ AQRFVL+E KW+RY+KLNLQSHYGSEFYCTLS+VEV+GVDA+ERMLEDLI
Sbjct: 1 GNFTASNVKQAQRFVLEEQKWMRYIKLNLQSHYGSEFYCTLSIVEVYGVDAIERMLEDLI 180
Query: 327 NTQDNLLASGEGNADKTIL-PHPDPAVIEHVHKKPLEGINSVPASDISSSKHETANIK-- 383
QD ASGEGN +K + P + A ++V + GINS PAS+ISS E +K
Sbjct: 181 YAQDKPFASGEGNGEKRVASPLSNAAKADNVRPNTITGINSDPASEISSENQEAIIVKRN 360
Query: 384 VPDPVEEIRQQVGRMPGDTVLKILMQKVRTLDVNLFVLERYMEDLNSRYVNIFKDYSKDT 443
VPDPVEEIRQQVGRMPGDTVLKILMQKVR LD+NL VLE+YMEDLNSRY+NIFK+YSKD
Sbjct: 361 VPDPVEEIRQQVGRMPGDTVLKILMQKVRYLDLNLSVLEQYMEDLNSRYINIFKEYSKDM 540
Query: 444 GEKDIVLQKIKEDIKNLIDHQDVIAKDASDLISWKSQVSSQLNHLIQDNAVLR 496
GEKD++L+KIKE+I ++ QDV+ K+ SDL SW+S S QL+H+++DNAVLR
Sbjct: 541 GEKDLLLEKIKEEISRFLERQDVMMKEFSDLDSWRSHFSVQLDHVLRDNAVLR 699
>TC209691
Length = 654
Score = 200 bits (508), Expect = 1e-51
Identities = 98/127 (77%), Positives = 109/127 (85%)
Frame = +3
Query: 370 SDISSSKHETANIKVPDPVEEIRQQVGRMPGDTVLKILMQKVRTLDVNLFVLERYMEDLN 429
S ISS+ HE N VPDPVEEIRQQVGRMPGDTVLKILMQKVRTLD+NLFVLERYMEDLN
Sbjct: 12 SRISSANHEKLNSNVPDPVEEIRQQVGRMPGDTVLKILMQKVRTLDLNLFVLERYMEDLN 191
Query: 430 SRYVNIFKDYSKDTGEKDIVLQKIKEDIKNLIDHQDVIAKDASDLISWKSQVSSQLNHLI 489
+RYVNIFK+YSKD G KDI++Q IKEDI+NL+D QD I KD SDL SWKS +S Q HL+
Sbjct: 192 TRYVNIFKEYSKDIGGKDILIQNIKEDIRNLVDQQDAITKDGSDLKSWKSHISMQFGHLL 371
Query: 490 QDNAVLR 496
+DNAVLR
Sbjct: 372 RDNAVLR 392
>AW279367 similar to GP|15290067|dbj P0025A05.23 {Oryza sativa (japonica
cultivar-group)}, partial (9%)
Length = 151
Score = 72.4 bits (176), Expect = 4e-13
Identities = 37/50 (74%), Positives = 42/50 (84%)
Frame = +2
Query: 181 ASASKGAKVLGSNKEGKGASNILSRDKDKYLRNPCSVVGKFVIMELSEET 230
ASASKGAKV SN+E KGASNILS DKDKYLRNPCS K VI+E+SE++
Sbjct: 2 ASASKGAKVSCSNQEAKGASNILSGDKDKYLRNPCSAEEKSVIIEISEKS 151
>CO985748
Length = 787
Score = 42.4 bits (98), Expect = 5e-04
Identities = 27/73 (36%), Positives = 40/73 (53%), Gaps = 2/73 (2%)
Frame = -3
Query: 426 EDLNSRYVNIFKDYSKDTGEKDIVLQKIKEDIKNLID--HQDVIAKDASDLISWKSQVSS 483
E L Y N + Y +D + +I +D L D + + K+ SDL SW+S S
Sbjct: 581 EGLEVCYENEIRVYLED------IYFEISDDANLLADCGANNPVMKEFSDLDSWRSHFSM 420
Query: 484 QLNHLIQDNAVLR 496
QL+H+++DNAVLR
Sbjct: 419 QLDHVLRDNAVLR 381
>TC207085 similar to UP|Q9SH20 (Q9SH20) F28K19.1, partial (22%)
Length = 1626
Score = 32.3 bits (72), Expect = 0.50
Identities = 15/39 (38%), Positives = 21/39 (53%)
Frame = +3
Query: 362 EGINSVPASDISSSKHETANIKVPDPVEEIRQQVGRMPG 400
+G VPA +S+ KH ++ +PVE R VG PG
Sbjct: 1032 KGYAYVPADCLSNEKHSNEDVYASEPVEHDR*SVGSQPG 1148
>TC207086
Length = 634
Score = 32.0 bits (71), Expect = 0.65
Identities = 15/39 (38%), Positives = 21/39 (53%)
Frame = +2
Query: 362 EGINSVPASDISSSKHETANIKVPDPVEEIRQQVGRMPG 400
+G VPA +S+ KH ++ +PVE R VG PG
Sbjct: 20 KGYAYVPADCLSNEKHSDEDVYASEPVEHDR*SVGSQPG 136
>TC207407 UP|Q6T2Z3 (Q6T2Z3) Cyclin-dependent kinase inhibitor 1;1, complete
Length = 1236
Score = 30.0 bits (66), Expect = 2.5
Identities = 19/48 (39%), Positives = 26/48 (53%)
Frame = +2
Query: 266 LGNFTASNVRHAQRFVLKEPKWVRYLKLNLQSHYGSEFYCTLSVVEVF 313
+G+ A NVR R ++E K L+ N H+G YC LS+V VF
Sbjct: 872 VGSVEALNVRVPVRIAIEEEK----LRSN*TCHFGLFIYC*LSLV*VF 1003
>BM271487 similar to GP|15810167|gb At1g64110/F22C12_22 {Arabidopsis
thaliana}, partial (15%)
Length = 411
Score = 29.3 bits (64), Expect = 4.2
Identities = 21/68 (30%), Positives = 33/68 (47%), Gaps = 3/68 (4%)
Frame = +2
Query: 428 LNSRYVNIFKDYSKDTGEKDIVLQKIKEDIKNLIDHQDVIA--KDASDLISWKSQVSSQL 485
L SR ++ DY +++ E I +L + I +D S L+SWKSQ+ L
Sbjct: 83 LGSRVIDSGNDY-----------EEVDEKINSLFPYNIEIRPPEDESHLVSWKSQLEEDL 229
Query: 486 NHL-IQDN 492
+ +QDN
Sbjct: 230 KMIQVQDN 253
>BI424944
Length = 350
Score = 28.9 bits (63), Expect = 5.5
Identities = 15/42 (35%), Positives = 21/42 (49%)
Frame = +2
Query: 2 KTIFLLLWCRCISHSLILVSLLYLYVDGSEELSVGLSKWNEV 43
K F L++C ISHSL LVS ++ G +WN +
Sbjct: 113 KLTFTLIFCIVISHSLGLVSFTLCIASETKRNKKGDLRWNGI 238
>TC216752 similar to UP|Q8LA49 (Q8LA49) Globulin-like protein, partial (53%)
Length = 748
Score = 28.9 bits (63), Expect = 5.5
Identities = 23/80 (28%), Positives = 35/80 (43%), Gaps = 5/80 (6%)
Frame = -3
Query: 212 RNPCSVVGKFVIMELSEETLVDTIEIANFEHHSS---NLKDFEIHGSLNFPT--NVWDLL 266
R C V GK+ I + + +I I+N HH N + HG L PT N +L
Sbjct: 470 RTSCQV-GKYRIRGSDDREPLQSIRISNLGHHKKPWHNEQITSFHGCLQNPTAVNTNNLN 294
Query: 267 GNFTASNVRHAQRFVLKEPK 286
T N+ + Q + +E +
Sbjct: 293 PATTPDNISNLQSRITRESR 234
>TC203439 weakly similar to UP|Q9LX02 (Q9LX02) ESTs AU082316(E3368), partial
(27%)
Length = 1497
Score = 28.5 bits (62), Expect = 7.2
Identities = 16/64 (25%), Positives = 33/64 (51%)
Frame = +2
Query: 146 NEFKSRAISSKVKSGTGQSRSVIHRLEPGGAEYNYASASKGAKVLGSNKEGKGASNILSR 205
NE + +++S +V + S HR+ G +Y + + KGA+ G +++ K + I
Sbjct: 227 NELRLQSLSDRV----AELESRFHRVLGGDRKYTWTAEIKGAEKNGFDRKYKWVAEIAEE 394
Query: 206 DKDK 209
++ K
Sbjct: 395 EEKK 406
>TC233225
Length = 888
Score = 28.1 bits (61), Expect = 9.4
Identities = 13/40 (32%), Positives = 23/40 (57%)
Frame = +1
Query: 223 IMELSEETLVDTIEIANFEHHSSNLKDFEIHGSLNFPTNV 262
+ +++ E +V+ + I S NLK+ +I GS N P +V
Sbjct: 634 LYQVNFEDMVEILVILRLITSSPNLKELQISGSSNIPVSV 753
>TC222034 weakly similar to UP|Q6H5X3 (Q6H5X3) ABA-responsive protein-like,
partial (47%)
Length = 557
Score = 28.1 bits (61), Expect = 9.4
Identities = 20/87 (22%), Positives = 42/87 (47%), Gaps = 9/87 (10%)
Frame = +1
Query: 135 DILSRAVPLGLNEFKSRAISSK--VKSGTGQSRSVIHRLE------PGGAEYNYASASKG 186
++L +++ + + + AI+ K GTG+SRS HR+ P +E S G
Sbjct: 34 NLLHKSISVDAYTYTNTAIAEKEIATKGTGKSRSFTHRIHDHVKMGPNLSEILKGKLSLG 213
Query: 187 AKVLGSNKEGKGASNILS-RDKDKYLR 212
A+++ G ++ ++K++ L+
Sbjct: 214 ARIIQEGGRGSIFKSVFGMQEKEQLLK 294
>TC215753 similar to UP|LSM3_HUMAN (P62310) U6 snRNA-associated Sm-like
protein LSm3 (MDS017), partial (60%)
Length = 630
Score = 28.1 bits (61), Expect = 9.4
Identities = 20/73 (27%), Positives = 36/73 (48%), Gaps = 7/73 (9%)
Frame = +1
Query: 103 INKENTDSANREDHVVENSEYAVKH--ENDVKKSDILSRA-----VPLGLNEFKSRAISS 155
I N+D+ DH+V S +AV + E D+ S + +R V L ++ + S +IS+
Sbjct: 28 IQFHNSDTCY*NDHIVHQSSHAVLNGGETDITPSPLTNRKGTVRFVVLTISSYVSSSIST 207
Query: 156 KVKSGTGQSRSVI 168
V + R ++
Sbjct: 208 VVTISSTSPRIIL 246
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.314 0.133 0.377
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,138,178
Number of Sequences: 63676
Number of extensions: 262687
Number of successful extensions: 1099
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 1085
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1095
length of query: 496
length of database: 12,639,632
effective HSP length: 101
effective length of query: 395
effective length of database: 6,208,356
effective search space: 2452300620
effective search space used: 2452300620
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 61 (28.1 bits)
Medicago: description of AC148651.5


