

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
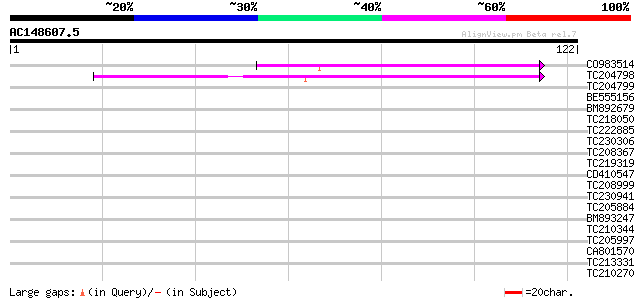
Query= AC148607.5 - phase: 0
(122 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
CO983514 45 4e-06
TC204798 similar to UP|Q7PC86 (Q7PC86) PDR7 ABC transporter, par... 45 6e-06
TC204799 similar to UP|Q7PC86 (Q7PC86) PDR7 ABC transporter, par... 36 0.003
BE555156 31 0.011
BM892679 33 0.014
TC218050 weakly similar to UP|Q7PC82 (Q7PC82) PDR14 ABC transpor... 33 0.014
TC222885 weakly similar to UP|Q8GU89 (Q8GU89) PDR-like ABC trans... 33 0.024
TC230306 weakly similar to UP|Q9VVR2 (Q9VVR2) CG6897-PA (LD27847... 28 0.063
TC208367 similar to GB|AAP37786.1|30725528|BT008427 At1g51560 {A... 30 0.20
TC219319 homologue to UP|Q9FYQ8 (Q9FYQ8) Endosomal protein-like,... 28 0.45
CD410547 28 0.77
TC208999 27 1.0
TC230941 homologue to UP|Q9FYQ8 (Q9FYQ8) Endosomal protein-like,... 27 1.0
TC205884 similar to UP|T9S2_HUMAN (Q99805) Transmembrane 9 super... 27 1.0
BM893247 27 1.3
TC210344 similar to UP|Q9AVE0 (Q9AVE0) SRKb, partial (21%) 27 1.3
TC205997 homologue to UP|Q9LIC2 (Q9LIC2) Multispanning membrane ... 25 1.6
CA801570 26 2.2
TC213331 similar to GB|AAM47365.1|21360499|AY113057 AT4g32140/F1... 26 2.2
TC210270 26 2.9
>CO983514
Length = 575
Score = 45.4 bits (106), Expect = 4e-06
Identities = 20/66 (30%), Positives = 38/66 (57%), Gaps = 4/66 (6%)
Frame = -2
Query: 54 LFIGVAELSMVV----SRLPVFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTY 109
LF G+ S V+ + V Y++R + PWAY+ ++++P F++ V+VI+TY
Sbjct: 571 LFFGINNCSTVLPYVATERTVLYRERFAGMYSPWAYSFAQVLIEVPYIFIQAVVYVIITY 392
Query: 110 YVIGFD 115
++ +D
Sbjct: 391 PMLSYD 374
>TC204798 similar to UP|Q7PC86 (Q7PC86) PDR7 ABC transporter, partial (20%)
Length = 1048
Score = 44.7 bits (104), Expect = 6e-06
Identities = 24/101 (23%), Positives = 50/101 (48%), Gaps = 4/101 (3%)
Frame = +3
Query: 19 AMIAMTIFLRTEMHRDSVAHGDIYVGALFYGCIVILFIGVAELS----MVVSRLPVFYKQ 74
A++ T+F R +R+S A + +GA++ ++F+G+ +V VFY++
Sbjct: 378 ALMIGTVFWRIGKNRESSADLTMIIGAMY---AAVIFVGINNCQTVQPIVAVERTVFYRE 548
Query: 75 RGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIGFD 115
R + P YAL ++P F + + ++ Y ++ F+
Sbjct: 549 RAAGMYAPLPYALAQVFCEVPYVFFQTVYYSLIVYAMVSFE 671
>TC204799 similar to UP|Q7PC86 (Q7PC86) PDR7 ABC transporter, partial (10%)
Length = 555
Score = 35.8 bits (81), Expect = 0.003
Identities = 18/77 (23%), Positives = 37/77 (47%), Gaps = 4/77 (5%)
Frame = +2
Query: 43 VGALFYGCIVILFIGVAELS----MVVSRLPVFYKQRGYLFFPPWAYALPAWILKIPLTF 98
+GA++ ++F+G+ +V VFY++R + P YAL +IP F
Sbjct: 26 IGAMY---AAVIFVGINNCQTVQPIVAVERTVFYRERAAGMYAPLPYALAQVFCEIPYVF 196
Query: 99 VEVAVWVILTYYVIGFD 115
+ + ++ Y ++ F+
Sbjct: 197 FQTVYYSLIVYAMVSFE 247
>BE555156
Length = 384
Score = 31.2 bits (69), Expect(2) = 0.011
Identities = 19/71 (26%), Positives = 33/71 (45%), Gaps = 5/71 (7%)
Frame = +2
Query: 50 CIV-ILFIGVAELS----MVVSRLPVFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVW 104
C+V F+GV +V VFY++R + YA+ I +IP FV+ +
Sbjct: 137 CMVQFFFVGVNNCQTVQPVVAVERTVFYRERAAGMYSALPYAIAQVISEIPYLFVQTIFF 316
Query: 105 VILTYYVIGFD 115
+ Y ++ F+
Sbjct: 317 SFIVYAMVSFE 349
Score = 21.6 bits (44), Expect(2) = 0.011
Identities = 9/29 (31%), Positives = 16/29 (55%)
Frame = +1
Query: 19 AMIAMTIFLRTEMHRDSVAHGDIYVGALF 47
A + T+F R +RD+ + +GAL+
Sbjct: 55 AFLVGTVFWRVGKNRDNTGDLNTIIGALY 141
>BM892679
Length = 422
Score = 33.5 bits (75), Expect = 0.014
Identities = 20/76 (26%), Positives = 36/76 (47%)
Frame = +1
Query: 39 GDIYVGALFYGCIVILFIGVAELSMVVSRLPVFYKQRGYLFFPPWAYALPAWILKIPLTF 98
G +Y+ +F G I L V + V Y+++ + AY+ ++IP
Sbjct: 154 GSMYIAVIFLGLNYCSTI----LPYVATERAVLYREKFAGMYSSTAYSFAQVAIEIPYIL 321
Query: 99 VEVAVWVILTYYVIGF 114
V+ ++V +TY +IGF
Sbjct: 322 VQSILYVAITYPMIGF 369
>TC218050 weakly similar to UP|Q7PC82 (Q7PC82) PDR14 ABC transporter, partial
(28%)
Length = 1436
Score = 33.5 bits (75), Expect = 0.014
Identities = 20/76 (26%), Positives = 36/76 (47%)
Frame = +2
Query: 39 GDIYVGALFYGCIVILFIGVAELSMVVSRLPVFYKQRGYLFFPPWAYALPAWILKIPLTF 98
G +Y+ +F G I L V + V Y+++ + AY+ ++IP
Sbjct: 659 GSMYIAVIFLGLNYCSTI----LPYVATERAVLYREKFAGMYSSTAYSFAQVAIEIPYIL 826
Query: 99 VEVAVWVILTYYVIGF 114
V+ ++V +TY +IGF
Sbjct: 827 VQSILYVAITYPMIGF 874
>TC222885 weakly similar to UP|Q8GU89 (Q8GU89) PDR-like ABC transporter (PDR4
ABC transporter), partial (9%)
Length = 484
Score = 32.7 bits (73), Expect = 0.024
Identities = 21/79 (26%), Positives = 38/79 (47%)
Frame = +2
Query: 37 AHGDIYVGALFYGCIVILFIGVAELSMVVSRLPVFYKQRGYLFFPPWAYALPAWILKIPL 96
A G +Y L G I A+ + V R VFY+++ + AYA ++++P
Sbjct: 179 AMGSMYAAVLLLG---IKNSNSAQPLVAVERT-VFYREKAAGMYSALAYAFAQVVVELPH 346
Query: 97 TFVEVAVWVILTYYVIGFD 115
++ V+ + Y +IGF+
Sbjct: 347 VLLQTVVYSAIVYAMIGFE 403
>TC230306 weakly similar to UP|Q9VVR2 (Q9VVR2) CG6897-PA (LD27847p), partial
(4%)
Length = 1043
Score = 28.1 bits (61), Expect(2) = 0.063
Identities = 11/16 (68%), Positives = 12/16 (74%)
Frame = +3
Query: 31 MHRDSVAHGDIYVGAL 46
MHRDSV HG +YV L
Sbjct: 747 MHRDSVTHGGLYVDTL 794
Score = 21.9 bits (45), Expect(2) = 0.063
Identities = 19/60 (31%), Positives = 29/60 (47%), Gaps = 1/60 (1%)
Frame = +2
Query: 53 ILFIGVAELSMVVSRLPVFYKQRGYLF-FPPWAYALPAWILKIPLTFVEVAVWVILTYYV 111
++F G+ LSMVVS L + + Y F F W L + K T +++ IL + V
Sbjct: 794 VVFNGLTGLSMVVSLLSLPEQSHLYSFNFIYWTLNLKTEVEKPS*T*LQLGKRRILAFVV 973
>TC208367 similar to GB|AAP37786.1|30725528|BT008427 At1g51560 {Arabidopsis
thaliana;} , partial (34%)
Length = 1062
Score = 29.6 bits (65), Expect = 0.20
Identities = 15/67 (22%), Positives = 27/67 (39%)
Frame = +1
Query: 55 FIGVAELSMVVSRLPVFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIGF 114
F+ + + + VFYK+R ++ Y L ++ P V +TYY++ F
Sbjct: 91 FMSIGGFPSFIEEMKVFYKERLNGYYGVGVYILSNFLSSFPFVAVMSIATGTITYYMVRF 270
Query: 115 DPYIGRY 121
Y
Sbjct: 271 RTEFSHY 291
>TC219319 homologue to UP|Q9FYQ8 (Q9FYQ8) Endosomal protein-like, partial
(26%)
Length = 854
Score = 28.5 bits (62), Expect = 0.45
Identities = 15/40 (37%), Positives = 20/40 (49%), Gaps = 5/40 (12%)
Frame = +2
Query: 80 FPPW-----AYALPAWILKIPLTFVEVAVWVILTYYVIGF 114
+P W A LP L I L F+ ++W+ YYV GF
Sbjct: 74 YPSWLLVLGAGTLPFGTLFIELFFIMSSIWMGRVYYVFGF 193
>CD410547
Length = 352
Score = 27.7 bits (60), Expect = 0.77
Identities = 10/22 (45%), Positives = 13/22 (58%)
Frame = +2
Query: 83 WAYALPAWILKIPLTFVEVAVW 104
WAY + +W L I + FVE W
Sbjct: 257 WAYEIKSWCLLISILFVEFRGW 322
>TC208999
Length = 1019
Score = 27.3 bits (59), Expect = 1.0
Identities = 15/40 (37%), Positives = 20/40 (49%), Gaps = 5/40 (12%)
Frame = +2
Query: 80 FPPW-----AYALPAWILKIPLTFVEVAVWVILTYYVIGF 114
+P W A LP L I L F+ ++W+ YYV GF
Sbjct: 32 YPSWLLVLGAGTLPFGTLFIELFFILSSIWLGRFYYVFGF 151
>TC230941 homologue to UP|Q9FYQ8 (Q9FYQ8) Endosomal protein-like, partial
(22%)
Length = 673
Score = 27.3 bits (59), Expect = 1.0
Identities = 15/39 (38%), Positives = 19/39 (48%), Gaps = 5/39 (12%)
Frame = +1
Query: 81 PPW-----AYALPAWILKIPLTFVEVAVWVILTYYVIGF 114
P W A LP L I L F+ ++W+ YYV GF
Sbjct: 1 PSWLLVLGAGTLPFGTLFIELFFIMSSIWMGRVYYVFGF 117
>TC205884 similar to UP|T9S2_HUMAN (Q99805) Transmembrane 9 superfamily
protein member 2 precursor (p76), partial (6%)
Length = 1205
Score = 27.3 bits (59), Expect = 1.0
Identities = 15/40 (37%), Positives = 20/40 (49%), Gaps = 5/40 (12%)
Frame = +1
Query: 80 FPPW-----AYALPAWILKIPLTFVEVAVWVILTYYVIGF 114
+P W A LP L I L F+ ++W+ YYV GF
Sbjct: 385 YPSWLLVLGAGTLPFGTLFIELFFILSSIWLGRFYYVFGF 504
>BM893247
Length = 405
Score = 26.9 bits (58), Expect = 1.3
Identities = 12/39 (30%), Positives = 21/39 (53%)
Frame = -3
Query: 60 ELSMVVSRLPVFYKQRGYLFFPPWAYALPAWILKIPLTF 98
E+S+V++ P YKQ + +FP + W++ L F
Sbjct: 382 EVSLVLNSTPCHYKQLSHFYFPLDEESFSHWLVDGCLNF 266
>TC210344 similar to UP|Q9AVE0 (Q9AVE0) SRKb, partial (21%)
Length = 870
Score = 26.9 bits (58), Expect = 1.3
Identities = 16/54 (29%), Positives = 29/54 (53%)
Frame = +3
Query: 23 MTIFLRTEMHRDSVAHGDIYVGALFYGCIVILFIGVAELSMVVSRLPVFYKQRG 76
M I+L++ RD + +++G ++Y C L V +S+ ++RL FY G
Sbjct: 210 MVIWLQSMQWRDYIP*SLMFLGLVYY-CWR*LQGNVTRVSITLTRLQAFYHMHG 368
>TC205997 homologue to UP|Q9LIC2 (Q9LIC2) Multispanning membrane
protein-like, partial (36%)
Length = 1407
Score = 25.0 bits (53), Expect(2) = 1.6
Identities = 10/28 (35%), Positives = 16/28 (56%)
Frame = +3
Query: 87 LPAWILKIPLTFVEVAVWVILTYYVIGF 114
LP + I L F+ ++W+ YY+ GF
Sbjct: 291 LPFGAVFIELFFILTSIWLNQFYYIFGF 374
Score = 20.0 bits (40), Expect(2) = 1.6
Identities = 6/18 (33%), Positives = 10/18 (55%)
Frame = +1
Query: 76 GYLFFPPWAYALPAWILK 93
G ++ PW+ W+LK
Sbjct: 133 GLVYQYPWSLLAVTWVLK 186
>CA801570
Length = 428
Score = 26.2 bits (56), Expect = 2.2
Identities = 20/69 (28%), Positives = 30/69 (42%), Gaps = 7/69 (10%)
Frame = +3
Query: 54 LFIGVAELSMVVSRLPVFYKQRGYLFFPPW-------AYALPAWILKIPLTFVEVAVWVI 106
LF+G + V S+LPV YK R L W +Y + + L + ++
Sbjct: 78 LFVGCYHIRGV*SKLPVVYKARRSL*NCTWWSMSQHISYTVVDSVSHFRLLLITKV--IV 251
Query: 107 LTYYVIGFD 115
L Y+I FD
Sbjct: 252 LNSYIINFD 278
>TC213331 similar to GB|AAM47365.1|21360499|AY113057 AT4g32140/F10N7_50
{Arabidopsis thaliana;} , partial (19%)
Length = 520
Score = 26.2 bits (56), Expect = 2.2
Identities = 17/53 (32%), Positives = 27/53 (50%), Gaps = 2/53 (3%)
Frame = +1
Query: 17 IMAMIAMTIFLRTEMHRDSVAHGDIYVGALFYGCIVILFIG--VAELSMVVSR 67
+++ + M++ + M D V HG Y GCI + F G +A LS +SR
Sbjct: 139 LVSTLGMSLTIPVAMIADMVIHGRKYSAMYILGCIQV-FAGFTLANLSGKISR 294
>TC210270
Length = 610
Score = 25.8 bits (55), Expect = 2.9
Identities = 16/50 (32%), Positives = 29/50 (58%)
Frame = +1
Query: 68 LPVFYKQRGYLFFPPWAYALPAWILKIPLTFVEVAVWVILTYYVIGFDPY 117
LP+F KQ +L P+AYA+ ++L + L+ + +++ Y IG P+
Sbjct: 85 LPLFKKQSFHLSLYPYAYAM--YMLILSLSI*PLI*PLLVYYNQIGATPF 228
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.336 0.149 0.486
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,093,532
Number of Sequences: 63676
Number of extensions: 84454
Number of successful extensions: 761
Number of sequences better than 10.0: 59
Number of HSP's better than 10.0 without gapping: 757
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 760
length of query: 122
length of database: 12,639,632
effective HSP length: 98
effective length of query: 24
effective length of database: 6,399,384
effective search space: 153585216
effective search space used: 153585216
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 51 (24.3 bits)
Medicago: description of AC148607.5


