

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148607.15 - phase: 0
(174 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
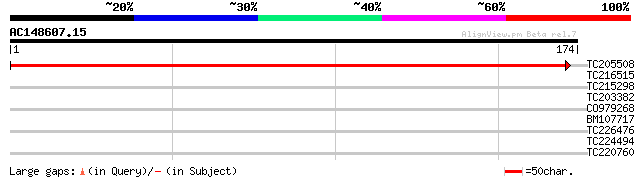
TC205508 weakly similar to UP|Q86UA1 (Q86UA1) PRPF39 protein (Fr... 276 3e-75
TC216515 weakly similar to UP|Q9LHY8 (Q9LHY8) ESTs D15336(C0474)... 29 0.98
TC215298 similar to UP|Q40090 (Q40090) SPF1 protein, partial (21%) 29 1.3
TC203382 similar to GB|AAL09765.1|15982836|AY057525 At1g27000/T7... 27 3.7
CO979268 27 4.9
BM107717 similar to SP|P37225|MAON_ NAD-dependent malic enzyme 5... 27 4.9
TC226476 homologue to UP|Q8L693 (Q8L693) 1-deoxy-D-xylulose 5-ph... 27 6.4
TC224494 26 8.3
TC220760 similar to UP|Q756S8 (Q756S8) AER176Wp, partial (9%) 26 8.3
>TC205508 weakly similar to UP|Q86UA1 (Q86UA1) PRPF39 protein (Fragment),
partial (5%)
Length = 1358
Score = 276 bits (707), Expect = 3e-75
Identities = 138/172 (80%), Positives = 146/172 (84%)
Frame = +2
Query: 1 MQQDWARLAMIYTRILENPNQQLDRYFNSFKELASNRPLSELRTADEAAAVAGVVSEGID 60
MQQDWA LA+IYTRILENPNQQLDRYF+SFKELA NRPLSELRTADEAAAVAGV SE
Sbjct: 197 MQQDWACLAVIYTRILENPNQQLDRYFSSFKELAGNRPLSELRTADEAAAVAGVASEATG 376
Query: 61 QGVEGEVHPDGADNSPKPASAGLTEAEELEKYIAIREEMYKKAKEFDSKIIGFETTIRRP 120
Q EGEVHPDGA+ SPK SAGLTEAEELEKYIAIREEMYKKAKEFDSKIIG ET IRRP
Sbjct: 377 QATEGEVHPDGAERSPKSVSAGLTEAEELEKYIAIREEMYKKAKEFDSKIIGCETAIRRP 556
Query: 121 YFHVRPLNIGELENWHNYLDFIEREGDLSKVTKLLSYKGTFCSIFSCYRLTY 172
YFHVRPLN+GELENWHNYLDFIEREGDLSK+ KL C+ + Y + Y
Sbjct: 557 YFHVRPLNVGELENWHNYLDFIEREGDLSKIAKLYERCVIACANYPEYWIRY 712
>TC216515 weakly similar to UP|Q9LHY8 (Q9LHY8) ESTs D15336(C0474), partial
(58%)
Length = 1320
Score = 29.3 bits (64), Expect = 0.98
Identities = 26/84 (30%), Positives = 37/84 (43%), Gaps = 10/84 (11%)
Frame = +3
Query: 36 NRPLSELRTADEAAAVAGV-----VSEGIDQGVEGEVHPDGADNSPKPASAGLTEAEE-- 88
NR + E+R + A VA VS I++G+ SP L+ A++
Sbjct: 456 NREMEEVRVVENAINVAPSENSENVSSNIEEGIA----------SPTNRRIYLSRAQDAV 605
Query: 89 ---LEKYIAIREEMYKKAKEFDSK 109
L K AIR++ KAK FD K
Sbjct: 606 TNMLAKGSAIRQDAVNKAKAFDEK 677
>TC215298 similar to UP|Q40090 (Q40090) SPF1 protein, partial (21%)
Length = 701
Score = 28.9 bits (63), Expect = 1.3
Identities = 28/99 (28%), Positives = 39/99 (39%), Gaps = 25/99 (25%)
Frame = -2
Query: 33 LASNRPLSELRTADEAAAVAGVVSEGIDQGVEGEV-------------------HPDGAD 73
+ R L ELR E+ + AG+ GI++ EGE+ P +D
Sbjct: 361 VGDGRMLEELRRTGESRSSAGLKPGGIEK*EEGEMGGGERGREGGGVDLNLGTPEPVLSD 182
Query: 74 NSPKPASAGLT----EAEELEKYIAIR--EEMYKKAKEF 106
P P GL EA EK + ++ E K KEF
Sbjct: 181 KPPPPPCGGLLLSVGEARRSEKEVVMKGCVEKVKLVKEF 65
>TC203382 similar to GB|AAL09765.1|15982836|AY057525 At1g27000/T7N9_6
{Arabidopsis thaliana;} , partial (50%)
Length = 1335
Score = 27.3 bits (59), Expect = 3.7
Identities = 14/44 (31%), Positives = 23/44 (51%)
Frame = +2
Query: 22 QLDRYFNSFKELASNRPLSELRTADEAAAVAGVVSEGIDQGVEG 65
Q+ R N ++LASNRP++ L E + ++ +V G G
Sbjct: 224 QVRRLANEVRQLASNRPITVLNGGSEQSNLSSLVVPAAALGALG 355
>CO979268
Length = 450
Score = 26.9 bits (58), Expect = 4.9
Identities = 13/38 (34%), Positives = 19/38 (49%), Gaps = 4/38 (10%)
Frame = -2
Query: 61 QGVEGEVHPDGADN----SPKPASAGLTEAEELEKYIA 94
QG EGE +P G N +P P ++ E K++A
Sbjct: 296 QGCEGEGNPQGTTNEKRANPNPEMGSTSQGERKRKWLA 183
>BM107717 similar to SP|P37225|MAON_ NAD-dependent malic enzyme 59 kDa
isoform mitochondrial precursor (EC 1.1.1.39) (NAD-ME).
[Potato], partial (13%)
Length = 408
Score = 26.9 bits (58), Expect = 4.9
Identities = 10/18 (55%), Positives = 13/18 (71%)
Frame = +3
Query: 21 QQLDRYFNSFKELASNRP 38
QQ DR+ NSF+ L +N P
Sbjct: 345 QQYDRFMNSFRSLXNNTP 398
>TC226476 homologue to UP|Q8L693 (Q8L693) 1-deoxy-D-xylulose 5-phosphate
synthase 1 precursor, partial (85%)
Length = 2000
Score = 26.6 bits (57), Expect = 6.4
Identities = 16/56 (28%), Positives = 26/56 (45%)
Frame = +3
Query: 13 TRILENPNQQLDRYFNSFKELASNRPLSELRTADEAAAVAGVVSEGIDQGVEGEVH 68
T L+ P + ++ L SNRPL ELR V++G+ + + G +H
Sbjct: 723 TATLDGPIPPVGALSSALSRLQSNRPLRELRE----------VAKGVTKRIGGPMH 860
>TC224494
Length = 568
Score = 26.2 bits (56), Expect = 8.3
Identities = 13/34 (38%), Positives = 20/34 (58%)
Frame = -3
Query: 57 EGIDQGVEGEVHPDGADNSPKPASAGLTEAEELE 90
+G+ G E E P SP+PAS+ ++ +ELE
Sbjct: 563 DGL*LGPEHEPRPACTSMSPQPASSPVSPEQELE 462
>TC220760 similar to UP|Q756S8 (Q756S8) AER176Wp, partial (9%)
Length = 721
Score = 26.2 bits (56), Expect = 8.3
Identities = 15/36 (41%), Positives = 19/36 (52%)
Frame = -1
Query: 41 ELRTADEAAAVAGVVSEGIDQGVEGEVHPDGADNSP 76
E+ E AG V G +GVEGEV GA++ P
Sbjct: 115 EVPRGGEGLGGAGEVGNG--EGVEGEVGEGGAEDEP 14
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.316 0.135 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,791,815
Number of Sequences: 63676
Number of extensions: 74183
Number of successful extensions: 408
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 407
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 408
length of query: 174
length of database: 12,639,632
effective HSP length: 91
effective length of query: 83
effective length of database: 6,845,116
effective search space: 568144628
effective search space used: 568144628
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 55 (25.8 bits)
Medicago: description of AC148607.15


