

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148607.14 - phase: 0
(359 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
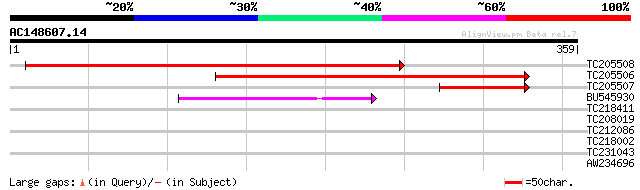
Score E
Sequences producing significant alignments: (bits) Value
TC205508 weakly similar to UP|Q86UA1 (Q86UA1) PRPF39 protein (Fr... 414 e-116
TC205506 similar to UP|Q6J1A1 (Q6J1A1) Fasciclin-like AGP 3, par... 338 2e-93
TC205507 102 3e-22
BU545930 71 7e-13
TC218411 similar to PIR|B96700|B96700 protein F12A21.17 [importe... 30 1.7
TC208019 homologue to UP|PPX2_ARATH (P48528) Serine/threonine pr... 29 2.9
TC212086 similar to UP|Q6RX29 (Q6RX29) RPP13-like protein (Fragm... 28 4.9
TC218002 weakly similar to UP|Q7T2E6 (Q7T2E6) XPA binding protei... 28 6.4
TC231043 28 6.4
AW234696 28 6.4
>TC205508 weakly similar to UP|Q86UA1 (Q86UA1) PRPF39 protein (Fragment),
partial (5%)
Length = 1358
Score = 414 bits (1064), Expect = e-116
Identities = 210/240 (87%), Positives = 222/240 (92%)
Frame = +2
Query: 11 VDQIVKLYERCVIACANYPEYWIRYVLCMEASESMDLANNVLARASQVFVKRQPEIHLFC 70
+ +I KLYERCVIACANYPEYWIRYVLCMEAS SM LANNVLARA+QVFVKRQPEIH+FC
Sbjct: 638 LSKIAKLYERCVIACANYPEYWIRYVLCMEASGSMYLANNVLARATQVFVKRQPEIHIFC 817
Query: 71 ARFKEQAGDIVGARAAYQLVHTEISPGLLEAIIRHANMEHRLGKLEDAFSLYEQAIAIEK 130
ARFKEQ GDI GARAAYQLVHTE SPGLLEAII+HANME+RL K+EDAFSLYEQAIAIEK
Sbjct: 818 ARFKEQTGDIDGARAAYQLVHTETSPGLLEAIIKHANMEYRLEKMEDAFSLYEQAIAIEK 997
Query: 131 GKEHSQTLPMLFAQYSRFVYLASGNSEKAREILVGGLENASLSKPLLEALLHFEAIQPQP 190
GKEHSQTLPMLFAQYSRFVYLASGN+EKAR+ILV GLEN LSKPLLEALLHFEAIQP P
Sbjct: 998 GKEHSQTLPMLFAQYSRFVYLASGNAEKARQILVEGLENVLLSKPLLEALLHFEAIQPLP 1177
Query: 191 KRVDIDFLESLVVKFITPNPENPGVASATEREELSNIFLEFLNLFGDVQSIKRAEDRHAK 250
KRV +DFLES VVKFI PN E+ GVAS TEREELS+IFLEFLNLFGDVQSIKRAEDRHAK
Sbjct: 1178KRVGVDFLESWVVKFIMPNSESAGVASPTEREELSSIFLEFLNLFGDVQSIKRAEDRHAK 1357
>TC205506 similar to UP|Q6J1A1 (Q6J1A1) Fasciclin-like AGP 3, partial (6%)
Length = 1229
Score = 338 bits (868), Expect = 2e-93
Identities = 169/199 (84%), Positives = 177/199 (88%)
Frame = +2
Query: 131 GKEHSQTLPMLFAQYSRFVYLASGNSEKAREILVGGLENASLSKPLLEALLHFEAIQPQP 190
G EHS T MLFAQYSRFVYLASGN+EKAR+ILV GLEN LSKPLLEA+LHFEAIQP P
Sbjct: 2 GNEHSHTSSMLFAQYSRFVYLASGNAEKARQILVEGLENVLLSKPLLEAILHFEAIQPLP 181
Query: 191 KRVDIDFLESLVVKFITPNPENPGVASATEREELSNIFLEFLNLFGDVQSIKRAEDRHAK 250
KRVDIDFLES VVKFI PN E+PGVASATEREELS+IFLEFLNLFGDVQSIKRAEDRHAK
Sbjct: 182 KRVDIDFLESWVVKFIMPNSESPGVASATEREELSSIFLEFLNLFGDVQSIKRAEDRHAK 361
Query: 251 LFLPNRGLSELKKRHAEDFLASDKTKVSRAYSAQSPAQSVAGAYPNGPNQWPNYGVQPQT 310
LFLP+R +SELKKRHAEDFLASDKTK R+YSAQSPAQS GAYPN NQW NYGVQPQT
Sbjct: 362 LFLPHRSMSELKKRHAEDFLASDKTKAPRSYSAQSPAQSGMGAYPNAQNQWSNYGVQPQT 541
Query: 311 WPATTQAQGQQWPAGYTQQ 329
WP TQAQGQQW AGYTQQ
Sbjct: 542 WPPVTQAQGQQWTAGYTQQ 598
>TC205507
Length = 623
Score = 102 bits (254), Expect = 3e-22
Identities = 46/57 (80%), Positives = 47/57 (81%)
Frame = +1
Query: 273 DKTKVSRAYSAQSPAQSVAGAYPNGPNQWPNYGVQPQTWPATTQAQGQQWPAGYTQQ 329
DKTKV R+ SAQSPAQS GAYPN NQW NYGVQPQTWP TQAQGQQW AGYTQQ
Sbjct: 1 DKTKVPRSXSAQSPAQSAMGAYPNAQNQWTNYGVQPQTWPPVTQAQGQQWTAGYTQQ 171
>BU545930
Length = 453
Score = 71.2 bits (173), Expect = 7e-13
Identities = 36/125 (28%), Positives = 69/125 (54%)
Frame = -2
Query: 108 MEHRLGKLEDAFSLYEQAIAIEKGKEHSQTLPMLFAQYSRFVYLASGNSEKAREILVGGL 167
ME RLG E AFS+Y++A+ + ++ LP + +SR Y ++ + + A ++L+ G+
Sbjct: 452 MEKRLGNTESAFSIYKEALKMASAEKMLHALPXXYVHFSRLKYXSTNSVDAAGDVLIDGV 273
Query: 168 ENASLSKPLLEALLHFEAIQPQPKRVDIDFLESLVVKFITPNPENPGVASATEREELSNI 227
+K LLE L+ F + K + + ++S++ I+P E S + E++SN+
Sbjct: 272 RTLPQNKLLLEELIKFXMMHGGTKHMAV--IDSIIADTISPRSEGSQGFSTEDAEDISNL 99
Query: 228 FLEFL 232
+LE +
Sbjct: 98 YLEVI 84
>TC218411 similar to PIR|B96700|B96700 protein F12A21.17 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(59%)
Length = 1850
Score = 30.0 bits (66), Expect = 1.7
Identities = 34/123 (27%), Positives = 51/123 (40%), Gaps = 8/123 (6%)
Frame = +1
Query: 60 VKRQPEIHLFCARFKEQAGDIVGARAAYQLV-----HTEISPGLLEAIIRHA-NMEHRLG 113
++ P KE+AGDI GA A + L I + A + + R G
Sbjct: 856 IQHMPATVATLVSLKERAGDIDGAAAVLDAAIKWWSNAMTEDNKLNTITQEAASFKLRHG 1035
Query: 114 KLEDAFSLYEQAIAIEKGKEHSQTLPMLFAQYSRFVYLASGNSEKAREILVG--GLENAS 171
K EDA LYEQ + K + + L L +R + + EK + L G G++ S
Sbjct: 1036KEEDAAQLYEQLV---KSQGSIEALVGLVTTVARMDVVKAELYEKQLKALPGLKGIDVDS 1206
Query: 172 LSK 174
L +
Sbjct: 1207LER 1215
>TC208019 homologue to UP|PPX2_ARATH (P48528) Serine/threonine protein
phosphatase PP-X isozyme 2 , partial (43%)
Length = 791
Score = 29.3 bits (64), Expect = 2.9
Identities = 12/31 (38%), Positives = 15/31 (47%)
Frame = -1
Query: 322 WPAGYTQQQVNWFFPSTIFLVCCFPSVQIPV 352
WP Y Q + FFP I+L FP P+
Sbjct: 788 WPKLYNQNSTSGFFPQIIYLESYFPIFPNPI 696
>TC212086 similar to UP|Q6RX29 (Q6RX29) RPP13-like protein (Fragment),
partial (5%)
Length = 571
Score = 28.5 bits (62), Expect = 4.9
Identities = 28/105 (26%), Positives = 48/105 (45%), Gaps = 18/105 (17%)
Frame = +1
Query: 210 PENPGVASATEREELSNIFLEFLNLFGDVQSIKRAEDRHAKL-----FLPNRGLSELKKR 264
P+N G+A TE E++++ +L+ L VQ KR D K L + +SE K
Sbjct: 34 PQNIGIADTTELEDVADFYLDELVDRSLVQVAKRRSDGGVKTCRIHELLRDLCISESKSD 213
Query: 265 HAEDFLASDKTK-------VSRAYSA------QSPAQSVAGAYPN 296
+ D++ +R+YS+ +S +S+ GA P+
Sbjct: 214 KYDKDTEIDESNGEADIRFDARSYSSRLNSMIRSDTESILGASPD 348
>TC218002 weakly similar to UP|Q7T2E6 (Q7T2E6) XPA binding protein 2, partial
(19%)
Length = 1500
Score = 28.1 bits (61), Expect = 6.4
Identities = 24/80 (30%), Positives = 36/80 (45%), Gaps = 3/80 (3%)
Frame = +1
Query: 100 EAIIRHANMEHRLGKLEDAFSLYEQAIAIEKGKEHSQTLPMLFAQYSRFVYLASGNS-EK 158
+ II +A EDAF +YE+ + I K + + S+FV N E+
Sbjct: 22 QIIINYAYFLEEHKYFEDAFKVYERGVKIFK---YPHVKDIWVTYLSKFVRRYGKNKLER 192
Query: 159 AREILVGGLENASLS--KPL 176
ARE+ +E+A KPL
Sbjct: 193 ARELFENAVESAPADQVKPL 252
>TC231043
Length = 891
Score = 28.1 bits (61), Expect = 6.4
Identities = 18/50 (36%), Positives = 27/50 (54%)
Frame = +1
Query: 155 NSEKAREILVGGLENASLSKPLLEALLHFEAIQPQPKRVDIDFLESLVVK 204
NS RE+ + +E AS KPL+ A H E+ P P + D + + VV+
Sbjct: 88 NSIMGRELCIAEVEAAS-GKPLVIATSHLESPCPAPPKWDQMYSKERVVQ 234
>AW234696
Length = 411
Score = 28.1 bits (61), Expect = 6.4
Identities = 20/67 (29%), Positives = 33/67 (48%)
Frame = +2
Query: 261 LKKRHAEDFLASDKTKVSRAYSAQSPAQSVAGAYPNGPNQWPNYGVQPQTWPATTQAQGQ 320
L+ + F + +++ + SA +P+ S G+ GPN P YG T P+T +A
Sbjct: 26 LRNKTTPSFHPTAISQLDSSKSATTPSASRYGS--QGPNDPPFYGWPTATSPSTAEAHTS 199
Query: 321 QWPAGYT 327
P+G T
Sbjct: 200 --PSGKT 214
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.136 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,836,791
Number of Sequences: 63676
Number of extensions: 215822
Number of successful extensions: 1134
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 1131
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1134
length of query: 359
length of database: 12,639,632
effective HSP length: 98
effective length of query: 261
effective length of database: 6,399,384
effective search space: 1670239224
effective search space used: 1670239224
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC148607.14


