

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148532.6 + phase: 0
(243 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
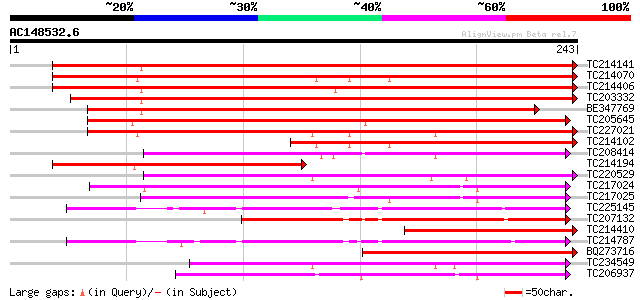
Score E
Sequences producing significant alignments: (bits) Value
TC214141 similar to UP|Q947D4 (Q947D4) Dehydration-responsive pr... 297 3e-81
TC214070 similar to UP|Q8VWQ1 (Q8VWQ1) Dehydration-induced prote... 290 4e-79
TC214406 weakly similar to UP|Q947D4 (Q947D4) Dehydration-respon... 286 8e-78
TC203332 weakly similar to UP|Q8H9E9 (Q8H9E9) Resistant specific... 229 7e-61
BE347769 weakly similar to GP|16588826|gb dehydration-responsive... 195 2e-50
TC205645 UP|Q8RUD9 (Q8RUD9) Seed coat BURP domain protein 1, com... 193 5e-50
TC227021 weakly similar to UP|O65009 (O65009) BURP domain contai... 178 2e-45
TC214102 similar to UP|Q9MB24 (Q9MB24) S1-3 protein (Fragment), ... 163 7e-41
TC208414 weakly similar to UP|O65009 (O65009) BURP domain contai... 146 7e-36
TC214194 weakly similar to UP|Q94IC5 (Q94IC5) BURP domain-contai... 142 1e-34
TC220529 weakly similar to GB|CAE02618.1|38567637|OSA575668 RAFT... 138 2e-33
TC217024 similar to UP|Q40161 (Q40161) Polygalacturonase isoenzy... 134 5e-32
TC217025 similar to UP|Q40161 (Q40161) Polygalacturonase isoenzy... 127 3e-30
TC225145 homologue to UP|O24482 (O24482) Sali3-2, complete 127 5e-30
TC207132 similar to UP|EA92_VICFA (P21747) Embryonic abundant pr... 126 1e-29
TC214410 similar to UP|Q8VWQ1 (Q8VWQ1) Dehydration-induced prote... 119 9e-28
TC214787 UP|Q06765 (Q06765) ADR6 protein (Sali5-4a protein), com... 119 1e-27
BQ273716 weakly similar to SP|Q08298|RD22_ Dehydration-responsiv... 119 2e-27
TC234549 weakly similar to UP|O65009 (O65009) BURP domain contai... 112 2e-25
TC206937 similar to UP|Q40161 (Q40161) Polygalacturonase isoenzy... 108 2e-24
>TC214141 similar to UP|Q947D4 (Q947D4) Dehydration-responsive protein RD22
(Fragment), partial (89%)
Length = 1353
Score = 297 bits (761), Expect = 3e-81
Identities = 142/228 (62%), Positives = 175/228 (76%), Gaps = 3/228 (1%)
Frame = +1
Query: 19 YDASVTQLHDYQNLDLFFFEKDLHNGTKLNMQFTKTG---ATFLPREVANSIPFSSNKVE 75
Y A+ TQLHD N+ LFF EKDLH+GTKL++ FT++ ATFL R+VA+SIPFSSNKV+
Sbjct: 415 YAATETQLHDDPNVALFFLEKDLHSGTKLDLHFTRSTSNQATFLSRQVADSIPFSSNKVD 594
Query: 76 NILNYFSIKQGSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNNVEAVS 135
I N FS+K GS E++ +K I CE I+GEEK C TSLESMVDF+TSKLGNNVE VS
Sbjct: 595 FIFNKFSVKPGSEEAQIMKNTISECEEGGIKGEEKYCATSLESMVDFSTSKLGNNVEVVS 774
Query: 136 TEANKESDKQQYIIAKGVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLPMEGIDGT 195
TE +KE+ Q+Y +A GVKKLS +K VVCH +YPYAVFYCHK T+ Y +P+EG +G
Sbjct: 775 TEVDKETGLQKYTVAPGVKKLSGDKAVVCHKQNYPYAVFYCHKTETTRAYSVPLEGTNGV 954
Query: 196 KVKTAAICHNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIWFSK 243
+VK A+CH DTS+W+PKHLAF VLKV+PGT+PVCHFL HV+W K
Sbjct: 955 RVKAVAVCHTDTSEWNPKHLAFQVLKVKPGTIPVCHFLPEDHVVWVPK 1098
>TC214070 similar to UP|Q8VWQ1 (Q8VWQ1) Dehydration-induced protein RD22-like
protein, partial (81%)
Length = 1427
Score = 290 bits (742), Expect = 4e-79
Identities = 146/233 (62%), Positives = 176/233 (74%), Gaps = 8/233 (3%)
Frame = +3
Query: 19 YDASVTQLHDYQNLDLFFFEKDLHNGTKLNMQFTK-----TGATFLPREVANSIPFSSNK 73
Y AS TQ HD N+ LFF EKDLH GTKLN+ FT+ A+FLPR VA+SIPFSSNK
Sbjct: 498 YAASETQWHDDPNVALFFLEKDLHYGTKLNLHFTRYFTSSVDASFLPRSVADSIPFSSNK 677
Query: 74 VENILNYFSIKQGSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNN-VE 132
V +LN FSIK+GS E++ +K I CE P I+GEEK CVTSLESMVDF T+KLG+N V+
Sbjct: 678 VNEVLNKFSIKEGSDEAQTVKNTISECEVPGIKGEEKRCVTSLESMVDFATTKLGSNDVD 857
Query: 133 AVSTEANKESDK-QQYIIAKGVKKLSENKI-VVCHLLSYPYAVFYCHKLRATKVYYLPME 190
AVSTE K+ ++ QQY +A GVK+L E+K VVCH +YPYAVFYCHK TK Y +P+E
Sbjct: 858 AVSTEVTKKDNELQQYTMAPGVKRLGEDKASVVCHKENYPYAVFYCHKSETTKAYSVPLE 1037
Query: 191 GIDGTKVKTAAICHNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIWFSK 243
G DG++VK A+CH DTS+W+PKHLAF VLKVQPGTVPVCHFL HV++ K
Sbjct: 1038GADGSRVKAVAVCHTDTSKWNPKHLAFQVLKVQPGTVPVCHFLPQDHVVFVPK 1196
>TC214406 weakly similar to UP|Q947D4 (Q947D4) Dehydration-responsive protein
RD22 (Fragment), partial (90%)
Length = 1379
Score = 286 bits (731), Expect = 8e-78
Identities = 138/230 (60%), Positives = 171/230 (74%), Gaps = 5/230 (2%)
Frame = +1
Query: 19 YDASVTQLHDYQNLDLFFFEKDLHNGTKLNMQFTKTG---ATFLPREVANSIPFSSNKVE 75
Y ++ TQLHD N+ LFF EKDLH GTKLN+ FT + ATFLPR+VA+SIPFSS+KVE
Sbjct: 463 YASTETQLHDDPNVALFFLEKDLHPGTKLNLHFTTSSNIQATFLPRQVADSIPFSSSKVE 642
Query: 76 NILNYFSIKQGSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNNVEAVS 135
+ N FS+K GS E++ +K + CE I+GEEK C TSLESM+DF+TSKLG NVE VS
Sbjct: 643 VVFNKFSVKPGSEEAQIMKNTLSECEEGGIKGEEKYCATSLESMIDFSTSKLGKNVEVVS 822
Query: 136 TEA--NKESDKQQYIIAKGVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLPMEGID 193
TE +KE+ Q+Y +A GV KLS +K VVCH +YPYAVFYCHK T+ Y +P+EG +
Sbjct: 823 TEVVEDKETGLQKYTVAPGVNKLSGDKAVVCHKQNYPYAVFYCHKTETTRAYSVPLEGAN 1002
Query: 194 GTKVKTAAICHNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIWFSK 243
G +VK A+CH TS+W+PKHLAF VLKV+PGTVPVCHFL HV+W K
Sbjct: 1003GVRVKAVAVCHTHTSEWNPKHLAFQVLKVKPGTVPVCHFLPEDHVVWVPK 1152
>TC203332 weakly similar to UP|Q8H9E9 (Q8H9E9) Resistant specific protein-2,
partial (60%)
Length = 1444
Score = 229 bits (585), Expect = 7e-61
Identities = 104/219 (47%), Positives = 140/219 (63%), Gaps = 2/219 (0%)
Frame = +1
Query: 27 HDYQNLDLFFFEKDLHNGTKLNMQFTKTG--ATFLPREVANSIPFSSNKVENILNYFSIK 84
HD D FFE+ L GTKL+ F K LPR++A IP SS K++ I+ +
Sbjct: 583 HDIPKADQVFFEEGLRPGTKLDAHFKKRENVTPLLPRQIAQHIPLSSAKIKEIVEMLFVN 762
Query: 85 QGSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNNVEAVSTEANKESDK 144
+ L++ I CE P I GEE+ C TSLESMVDF TSKLG N +STEA KES
Sbjct: 763 PEPENVKILEETISMCEVPAITGEERYCATSLESMVDFVTSKLGKNARVISTEAEKESKS 942
Query: 145 QQYIIAKGVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLPMEGIDGTKVKTAAICH 204
Q++ + GVK L+E+K++VCH + YPY VF CH++ T +++P+EG DGT+VK AA+CH
Sbjct: 943 QKFSVKDGVKLLAEDKVIVCHPMDYPYVVFMCHEISNTTAHFMPLEGEDGTRVKAAAVCH 1122
Query: 205 NDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIWFSK 243
DTS+W P H+ +LK +PG PVCH GH++WF+K
Sbjct: 1123KDTSEWDPNHVFLQMLKTKPGAAPVCHIFPEGHLLWFAK 1239
>BE347769 weakly similar to GP|16588826|gb dehydration-responsive protein
RD22 {Prunus persica}, partial (38%)
Length = 595
Score = 195 bits (495), Expect = 2e-50
Identities = 104/197 (52%), Positives = 135/197 (67%), Gaps = 3/197 (1%)
Frame = +2
Query: 34 LFFFEKDLHNGTKLNMQFTKTG---ATFLPREVANSIPFSSNKVENILNYFSIKQGSAES 90
L+ E DLH+GTKL++ FT++ ATFL R+VA+SIPFSSNKV+ I N S K GS E+
Sbjct: 5 LYLVEFDLHSGTKLDLHFTRSTSNQATFLSRQVADSIPFSSNKVDFIFNKNSGKPGSKEA 184
Query: 91 ENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNNVEAVSTEANKESDKQQYIIA 150
+ +K I CE I+GEEK C TSLESM DF+TSKLGNNVE VSTE +K++ +Y A
Sbjct: 185 QIMKNTIIECEEGGIKGEEKYCATSLESMDDFSTSKLGNNVEVVSTEVDKDTG*HKYTGA 364
Query: 151 KGVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLPMEGIDGTKVKTAAICHNDTSQW 210
GVKK+S +K VVCH +YPYA FYC++ T Y +P EG + + K + CH DTS
Sbjct: 365 PGVKKVSGDKAVVCHKHNYPYADFYCNRTDTTIAYSVPSEGTNVVRDKAVSRCHTDTSDM 544
Query: 211 SPKHLAFHVLKVQPGTV 227
P+H+A HVL V P ++
Sbjct: 545 YPRHVANHVLTVIPRSI 595
>TC205645 UP|Q8RUD9 (Q8RUD9) Seed coat BURP domain protein 1, complete
Length = 1369
Score = 193 bits (491), Expect = 5e-50
Identities = 90/211 (42%), Positives = 131/211 (61%), Gaps = 4/211 (1%)
Frame = +1
Query: 34 LFFFEKDLHNGTKLNMQF---TKTGATFLPREVANSIPFSSNKVENILNYFSIKQGSAES 90
LFF E+DL G NM+F TK LPR+++ IPFS +K + +L ++ S+ +
Sbjct: 310 LFFLEEDLRAGKIFNMKFVNNTKATVPLLPRQISKQIPFSEDKKKQVLAMLGVEANSSNA 489
Query: 91 ENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNNVEAVSTEANKESDKQQYIIA 150
+ + + IG C+ P EGE K C TSLESMVDF S LG NV A STE +E++ ++++
Sbjct: 490 KIIAETIGLCQEPATEGERKHCATSLESMVDFVVSALGKNVGAFSTEKERETESGKFVVV 669
Query: 151 K-GVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLPMEGIDGTKVKTAAICHNDTSQ 209
K GV+KL ++K++ CH +SYPY VF CH + + Y + ++G DG +VK CH DTS+
Sbjct: 670 KNGVRKLGDDKVIACHPMSYPYVVFGCHLVPRSSGYLVRLKGEDGVRVKAVVACHRDTSK 849
Query: 210 WSPKHLAFHVLKVQPGTVPVCHFLQHGHVIW 240
W H AF VL ++PG VCH G+++W
Sbjct: 850 WDHNHGAFKVLNLKPGNGTVCHVFTEGNLLW 942
>TC227021 weakly similar to UP|O65009 (O65009) BURP domain containing
protein, partial (47%)
Length = 1147
Score = 178 bits (452), Expect = 2e-45
Identities = 91/219 (41%), Positives = 133/219 (60%), Gaps = 9/219 (4%)
Frame = +1
Query: 34 LFFFEKDLHNGTKLNMQFTK----TGATFLPREVANSIPFSSNKVENILNYFSIKQGSAE 89
+FF +DL G + + F K T PRE A+S+PFS NK+ N+L FS+ Q S +
Sbjct: 316 VFFTIEDLKVGKTMPIHFPKRDPATSPKLWPREEADSLPFSLNKLPNLLKIFSVSQNSPK 495
Query: 90 SENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLG--NNVEAVSTEANKESDK--Q 145
++ ++ + CET I+GE K C TSLESM+DFT S LG ++++ +ST +S Q
Sbjct: 496 AKAMEDTLRECETKPIKGEVKFCATSLESMLDFTQSILGFTSDLKVLSTSHQTKSSVTFQ 675
Query: 146 QYIIAKGVKKLSENKIVVCHLLSYPYAVFYCHKLRA-TKVYYLPMEGIDGTKVKTAAICH 204
Y + + + ++ +K+V CH + YPY VFYCH + K+Y +P+ G +G +V +CH
Sbjct: 676 NYTMLENIIEIPASKMVACHTMPYPYTVFYCHSQESENKIYRVPLAGENGDRVDAMVVCH 855
Query: 205 NDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIWFSK 243
DTSQW H++F VLKV+PGT VCHF H+IW K
Sbjct: 856 MDTSQWGHGHVSFQVLKVKPGTTSVCHFFPADHLIWVPK 972
>TC214102 similar to UP|Q9MB24 (Q9MB24) S1-3 protein (Fragment), partial
(70%)
Length = 621
Score = 163 bits (412), Expect = 7e-41
Identities = 77/126 (61%), Positives = 97/126 (76%), Gaps = 3/126 (2%)
Frame = +3
Query: 121 DFTTSKLGNN-VEAVSTEANKESDK-QQYIIAKGVKKLSENKI-VVCHLLSYPYAVFYCH 177
DF T+KLG+N V+AVSTE K+ ++ QQY +A GVK+L E+K VVCH +YPYAVFYCH
Sbjct: 15 DFATTKLGSNDVDAVSTEVTKKDNELQQYTMAPGVKRLGEDKASVVCHKENYPYAVFYCH 194
Query: 178 KLRATKVYYLPMEGIDGTKVKTAAICHNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGH 237
K TK Y +P+EG DG++VK A+CH DTS+W+PKHLAF VLKV PGTVP+CHFL H
Sbjct: 195 KSETTKAYSVPLEGADGSRVKAVAVCHTDTSKWNPKHLAFQVLKVHPGTVPICHFLPQDH 374
Query: 238 VIWFSK 243
V++ K
Sbjct: 375 VVFVPK 392
>TC208414 weakly similar to UP|O65009 (O65009) BURP domain containing
protein, partial (39%)
Length = 957
Score = 146 bits (369), Expect = 7e-36
Identities = 77/188 (40%), Positives = 114/188 (59%), Gaps = 5/188 (2%)
Frame = +2
Query: 58 FLPREVANSIPFSSNKVENILNYFSIKQGSAESENLKQAIGYCETPTIEGEEKSCVTSLE 117
F P + FSS + ++L +FSI + S +++ +K + CE +EGE K C TSLE
Sbjct: 239 F*PEKKLTRFLFSSKHLPSLLKFFSIPKHSPQAKAMKYTLKQCEFEPMEGETKFCATSLE 418
Query: 118 SMVDFTTSKLGNNVE--AVSTE--ANKESDKQQYIIAKGVKKLSENKIVVCHLLSYPYAV 173
S+ DF G+N + ++T N + Q Y I++ VK +S ++ CH + YPYAV
Sbjct: 419 SLFDFAHYLFGSNAQFKVLTTVHLTNSTTLLQNYTISE-VKVISVPNVIGCHPMPYPYAV 595
Query: 174 FYCHKLRA-TKVYYLPMEGIDGTKVKTAAICHNDTSQWSPKHLAFHVLKVQPGTVPVCHF 232
FYCH + T +Y + +EG +G +V+ AAICH DTS+W H++F VLKVQPGT PVCHF
Sbjct: 596 FYCHSQHSDTNLYEVMVEGENGGRVQAAAICHMDTSKWDRDHVSFRVLKVQPGTSPVCHF 775
Query: 233 LQHGHVIW 240
+++W
Sbjct: 776 FPPDNLVW 799
>TC214194 weakly similar to UP|Q94IC5 (Q94IC5) BURP domain-containing protein
(Fragment), partial (49%)
Length = 704
Score = 142 bits (358), Expect = 1e-34
Identities = 73/114 (64%), Positives = 86/114 (75%), Gaps = 5/114 (4%)
Frame = +1
Query: 19 YDASVTQLHDYQNLDLFFFEKDLHNGTKLNMQFT-----KTGATFLPREVANSIPFSSNK 73
Y AS TQLHD N+ LFF EKDLH+GTKLN+ FT ATFLPR V++SIPFSSNK
Sbjct: 361 YAASETQLHDDPNVALFFLEKDLHHGTKLNLHFTIYYTSNVDATFLPRSVSDSIPFSSNK 540
Query: 74 VENILNYFSIKQGSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKL 127
V ++LN FSIK GS E++ +K I CE P+I+GEEK CVTSLESMVDF T+KL
Sbjct: 541 VNDVLNKFSIKDGSDEAKTVKNTINECEGPSIKGEEKRCVTSLESMVDFATTKL 702
>TC220529 weakly similar to GB|CAE02618.1|38567637|OSA575668 RAFTIN1 protein
{Oryza sativa (japonica cultivar-group);} , partial
(15%)
Length = 942
Score = 138 bits (348), Expect = 2e-33
Identities = 71/192 (36%), Positives = 110/192 (56%), Gaps = 6/192 (3%)
Frame = +3
Query: 58 FLPREVANSIPFSSNKVENILNYFSIKQGSAESENLKQAIGYCETPTIEGEEKSCVTSLE 117
FLPR+ A SIPFS +++ ++L FSI + S E+ ++ + CE I GE K C TSLE
Sbjct: 216 FLPRKEAESIPFSISQLPSVLQLFSISEDSPEANAMRDTLEQCEAEPITGETKICATSLE 395
Query: 118 SMVDFTTSKLG----NNVEAVSTEANKESDKQQYIIAKGVKKLSENKIVVCHLLSYPYAV 173
SM++F + +G +N+ + Q++ I + + ++ K V CH L YPYA+
Sbjct: 396 SMLEFIGTIIGSETKHNILTTTLPTASGVPLQKFTILEVSEDINAAKWVACHPLPYPYAI 575
Query: 174 FYCHKL-RATKVYYLPMEGIDG-TKVKTAAICHNDTSQWSPKHLAFHVLKVQPGTVPVCH 231
+YCH + +KV+ + + +G K++ ICH DTS WSP H+ F L ++PG VCH
Sbjct: 576 YYCHFIATGSKVFKVSLGSENGDDKIEALGICHLDTSDWSPNHIIFRQLGIKPGKDAVCH 755
Query: 232 FLQHGHVIWFSK 243
F H++W K
Sbjct: 756 FFPIKHLMWVPK 791
>TC217024 similar to UP|Q40161 (Q40161) Polygalacturonase isoenzyme 1 beta
subunit precursor , partial (33%)
Length = 958
Score = 134 bits (336), Expect = 5e-32
Identities = 78/213 (36%), Positives = 111/213 (51%), Gaps = 7/213 (3%)
Frame = +1
Query: 35 FFFEKDLHNGTKLNMQFTKTGA---TFLPREVANSIPFSSNKVENILNYFSIKQGSAESE 91
FF E L GT + M + +FLPR + +PFSS KV+ + F + + S+ +
Sbjct: 85 FFRETMLKEGTVMPMPDIRDKMPKRSFLPRAILTKLPFSSAKVDELKRVFKVSENSSMDK 264
Query: 92 NLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNNVEAVSTEANKESDKQQYI-IA 150
+ ++G CE GE K CV S+E M+DF+TS LG NV +TE K S+K +
Sbjct: 265 MIMDSLGECERAPSVGETKRCVASVEDMIDFSTSVLGRNVAVWTTENVKGSNKNVMVGRV 444
Query: 151 KGVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLPMEGIDGTKVKT---AAICHNDT 207
KG+ K V CH +PY ++YCH L +VY + + +K K AICH DT
Sbjct: 445 KGMNGGKVTKSVSCHQSLFPYLLYYCHSLPKVRVYEANLLDPE-SKAKINHGVAICHLDT 621
Query: 208 SQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIW 240
+ WSP H AF L PG + VCH++ + W
Sbjct: 622 TAWSPTHGAFLALGSGPGRIEVCHWIFENDLTW 720
>TC217025 similar to UP|Q40161 (Q40161) Polygalacturonase isoenzyme 1 beta
subunit precursor , partial (32%)
Length = 933
Score = 127 bits (320), Expect = 3e-30
Identities = 69/190 (36%), Positives = 103/190 (53%), Gaps = 6/190 (3%)
Frame = +2
Query: 57 TFLPREVANSIPFSSNKVENILNYFSIKQGSAESENLKQAIGYCETPTIEGEEKSCVTSL 116
+FLPR + +PFSS+KV + F + S+ + + ++G CE GE K CV S+
Sbjct: 41 SFLPRSILTKLPFSSSKVHELKRLFKVSDNSSMEKMIMDSLGECERVPSMGETKRCVGSI 220
Query: 117 ESMVDFTTSKLGNNVEAVSTEANKESDKQQYIIAKGVKKLSENKI---VVCHLLSYPYAV 173
E M+DF+TS LG NV +TE S+K ++ VK ++ K+ V CH +PY +
Sbjct: 221 EDMIDFSTSVLGRNVAVWTTENVNGSNKN--VMVGRVKGMNGGKVTQSVSCHQSLFPYML 394
Query: 174 FYCHKLRATKVYYLPMEGIDGTKVKT---AAICHNDTSQWSPKHLAFHVLKVQPGTVPVC 230
+YCH + +VY + + +K K AICH DT+ WSP H AF L PG + VC
Sbjct: 395 YYCHSVPKVRVYQADLLDPE-SKAKINHGVAICHLDTTAWSPTHGAFMALGSGPGRIEVC 571
Query: 231 HFLQHGHVIW 240
H++ + W
Sbjct: 572 HWIFENDLTW 601
>TC225145 homologue to UP|O24482 (O24482) Sali3-2, complete
Length = 1382
Score = 127 bits (319), Expect = 5e-30
Identities = 75/217 (34%), Positives = 119/217 (54%), Gaps = 1/217 (0%)
Frame = +3
Query: 25 QLHDYQNLDLFFFEKDLHNGTKLNMQFTKTGATFLPREVANSIPFSSNKVENILNYFS-I 83
Q+ D Q FF+++DLH G + +QFTK P++ + + + + I
Sbjct: 369 QIDDTQYPKTFFYKEDLHPGKTMKVQFTKR-------------PYA--QPYGVYTWLTDI 503
Query: 84 KQGSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNNVEAVSTEANKESD 143
K S E + ++ + EGEEK C SL +++ F SKLG N++ +S+ +
Sbjct: 504 KDTSKEGYSFEEIC--IKKEAFEGEEKFCAKSLGTVIGFAISKLGKNIQVLSSSF---VN 668
Query: 144 KQQYIIAKGVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLPMEGIDGTKVKTAAIC 203
KQ+ +GV+ L + K V+CH L++ AVFYCHK+R T + +P+ DGTK + A+C
Sbjct: 669 KQEQYTVEGVQNLGD-KAVMCHGLNFRTAVFYCHKVRETTAFMVPLVAGDGTKTQALAVC 845
Query: 204 HNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIW 240
H+DTS + H+ ++ V PGT PVCHFL ++W
Sbjct: 846 HSDTSGMN-HHMLHELMGVDPGTNPVCHFLGSKAILW 953
>TC207132 similar to UP|EA92_VICFA (P21747) Embryonic abundant protein USP92
precursor, partial (51%)
Length = 847
Score = 126 bits (316), Expect = 1e-29
Identities = 63/141 (44%), Positives = 90/141 (63%)
Frame = +3
Query: 100 CETPTIEGEEKSCVTSLESMVDFTTSKLGNNVEAVSTEANKESDKQQYIIAKGVKKLSEN 159
C +GEEK C TSL+SM+ F SKLG N++A+S+ ++ D QY++ + V K+ E
Sbjct: 423 CGKAAAKGEEKFCATSLQSMMGFAISKLGKNIKAISSSFAQDHD--QYVVEE-VNKIGE- 590
Query: 160 KIVVCHLLSYPYAVFYCHKLRATKVYYLPMEGIDGTKVKTAAICHNDTSQWSPKHLAFHV 219
K V+CH L++ VFYCH++ AT Y +P+ DGTK K ICH+DT P + + V
Sbjct: 591 KAVMCHRLNFENVVFYCHQINATTTYMVPLVASDGTKAKALTICHHDTRGMDP-IVVYEV 767
Query: 220 LKVQPGTVPVCHFLQHGHVIW 240
LKV+ GTVPVCHF+ + + W
Sbjct: 768 LKVKTGTVPVCHFVGNKAIAW 830
>TC214410 similar to UP|Q8VWQ1 (Q8VWQ1) Dehydration-induced protein RD22-like
protein, partial (22%)
Length = 575
Score = 119 bits (299), Expect = 9e-28
Identities = 50/74 (67%), Positives = 59/74 (79%)
Frame = +1
Query: 170 PYAVFYCHKLRATKVYYLPMEGIDGTKVKTAAICHNDTSQWSPKHLAFHVLKVQPGTVPV 229
PYAVFYCHK TK Y +P+EG DG++VK A+CH DTS+W+PKHLAF VLKVQPGTVPV
Sbjct: 121 PYAVFYCHKSETTKAYSVPLEGADGSRVKAVAVCHTDTSKWNPKHLAFQVLKVQPGTVPV 300
Query: 230 CHFLQHGHVIWFSK 243
CHFL HV++ K
Sbjct: 301 CHFLPQDHVVFVPK 342
>TC214787 UP|Q06765 (Q06765) ADR6 protein (Sali5-4a protein), complete
Length = 1150
Score = 119 bits (298), Expect = 1e-27
Identities = 76/217 (35%), Positives = 118/217 (54%), Gaps = 1/217 (0%)
Frame = +3
Query: 25 QLHDYQNLDLFFFEKDLHNGTKLNMQFTKTGATFLPREVANSIPFSSN-KVENILNYFSI 83
Q+ + Q FF+++DLH G + +QF+K PF V L I
Sbjct: 240 QIEETQYPKTFFYKEDLHPGKTMKVQFSKP-------------PFQQPWGVGTWLK--EI 374
Query: 84 KQGSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNNVEAVSTEANKESD 143
K + E + ++ + IEGEEK C SL +++ F SKLG N++ +S+ + D
Sbjct: 375 KDTTKEGYSFEELC--IKKEAIEGEEKFCAKSLGTVIGFAISKLGKNIQVLSSSFVNKQD 548
Query: 144 KQQYIIAKGVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLPMEGIDGTKVKTAAIC 203
QY + +GV+ L + K V+CH L++ AVFYCH++R T + +P+ DGTK + AIC
Sbjct: 549 --QYTV-EGVQNLGD-KAVMCHRLNFRTAVFYCHEVRETTAFMVPLVAGDGTKTQALAIC 716
Query: 204 HNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIW 240
H++TS + + L ++ V PGT PVCHFL ++W
Sbjct: 717 HSNTSGMNHQML-HQLMGVDPGTNPVCHFLGSKAILW 824
>BQ273716 weakly similar to SP|Q08298|RD22_ Dehydration-responsive protein
RD22 precursor. [Mouse-ear cress] {Arabidopsis
thaliana}, partial (13%)
Length = 419
Score = 119 bits (297), Expect = 2e-27
Identities = 44/92 (47%), Positives = 65/92 (69%)
Frame = +1
Query: 152 GVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLPMEGIDGTKVKTAAICHNDTSQWS 211
G+ ++ K++VCH + YPY VF CH++ T +++P+EG DGT+VK AA+CH DTS+W
Sbjct: 43 GLVLVTLKKVIVCHPMDYPYVVFMCHEISNTTAHFMPLEGEDGTRVKAAAVCHKDTSEWD 222
Query: 212 PKHLAFHVLKVQPGTVPVCHFLQHGHVIWFSK 243
P H+ +LK +PG PVCH GH++WF+K
Sbjct: 223 PNHVFLQMLKTKPGAAPVCHIFPEGHLLWFAK 318
>TC234549 weakly similar to UP|O65009 (O65009) BURP domain containing
protein, partial (14%)
Length = 791
Score = 112 bits (280), Expect = 2e-25
Identities = 58/171 (33%), Positives = 90/171 (51%), Gaps = 8/171 (4%)
Frame = +1
Query: 78 LNYFSIKQGSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLG----NNVEA 133
L FSI + S + ++ + C I GE K C TSLESM++F +G +N+
Sbjct: 25 LQLFSISEDSPXANAMRDTLDQCXXEPITGETKICATSLESMLEFVGKIIGLETKHNIIT 204
Query: 134 VSTEANKESDKQQYIIAKGVKKLSENKIVVCHLLSYPYAVFYCHKLRA-TKVYYLPM--- 189
+ Q++ I + + ++ +K V CH L YPYA++YCH + +KV+ + +
Sbjct: 205 TTLPTASGVPLQKFTILEVSEDINASKWVACHPLPYPYAIYYCHFIATGSKVFKVSLGSE 384
Query: 190 EGIDGTKVKTAAICHNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIW 240
D K++ ICH DTS WSP H+ F L ++PG VCHF H++W
Sbjct: 385 NNGDDDKIEALGICHLDTSDWSPNHIIFRQLGIKPGKDSVCHFFTIKHLMW 537
>TC206937 similar to UP|Q40161 (Q40161) Polygalacturonase isoenzyme 1 beta
subunit precursor , partial (23%)
Length = 537
Score = 108 bits (271), Expect = 2e-24
Identities = 61/174 (35%), Positives = 87/174 (49%), Gaps = 5/174 (2%)
Frame = +2
Query: 72 NKVENILNYFSIKQGSAESENLKQAIGYCETPTIEGEEKSCVTSLESMVDFTTSKLGNNV 131
+K++ + F + + +K ++ CE GE K CV SLE M+DF TS LG NV
Sbjct: 5 SKIDELKQVFKASDNGSMEKMMKDSLEECERAPSSGETKRCVGSLEDMIDFATSVLGRNV 184
Query: 132 EAVSTEANKESDKQQYII--AKGVKKLSENKIVVCHLLSYPYAVFYCHKLRATKVYYLPM 189
AV T N K+ ++ +G+ + V CH +PY ++YCH + +VY +
Sbjct: 185 -AVRTTQNVNGSKKSVVVGPVRGINGGKVTQSVSCHQSLFPYLLYYCHAVPKVRVYEADL 361
Query: 190 EGIDGTKVKT---AAICHNDTSQWSPKHLAFHVLKVQPGTVPVCHFLQHGHVIW 240
TK K AICH DTS WSP H AF L PG + VCH++ + W
Sbjct: 362 LD-PKTKAKINRGVAICHLDTSDWSPTHGAFLSLGSVPGRIEVCHWIFENDMAW 520
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.133 0.402
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,726,541
Number of Sequences: 63676
Number of extensions: 161934
Number of successful extensions: 985
Number of sequences better than 10.0: 80
Number of HSP's better than 10.0 without gapping: 934
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 937
length of query: 243
length of database: 12,639,632
effective HSP length: 95
effective length of query: 148
effective length of database: 6,590,412
effective search space: 975380976
effective search space used: 975380976
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Medicago: description of AC148532.6


