

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148532.15 + phase: 1 /partial
(178 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
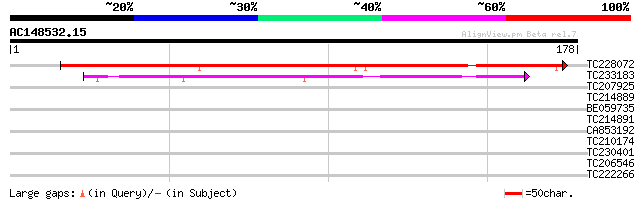
Score E
Sequences producing significant alignments: (bits) Value
TC228072 weakly similar to UP|Q9LW18 (Q9LW18) Emb|CAB87707.1, pa... 174 3e-44
TC233183 similar to UP|Q8IP81 (Q8IP81) CG31856-PA, partial (21%) 44 5e-05
TC207925 similar to UP|O65462 (O65462) Receptor like protein (Fr... 30 0.60
TC214889 homologue to PIR|C86216|C86216 protein T23G18.6 [import... 30 0.79
BE059735 homologue to PIR|C86216|C86 protein T23G18.6 [imported]... 30 0.79
TC214891 homologue to PIR|C86216|C86216 protein T23G18.6 [import... 28 3.0
CA853192 27 3.9
TC210174 27 5.1
TC230401 similar to UP|Q8VZI6 (Q8VZI6) AT4g33380/F17M5_140, part... 26 8.7
TC206546 weakly similar to UP|Q9LFD1 (Q9LFD1) Laccase-like prote... 26 8.7
TC222266 26 8.7
>TC228072 weakly similar to UP|Q9LW18 (Q9LW18) Emb|CAB87707.1, partial (11%)
Length = 1066
Score = 174 bits (440), Expect = 3e-44
Identities = 88/164 (53%), Positives = 118/164 (71%), Gaps = 5/164 (3%)
Frame = +3
Query: 17 FKWGQKRGKGGKKRDIQFYQSFTLRGVDYSLFDNVYVKNDFA--EPRIGKIIKIWETPTL 74
F WG+KRG GGKK+D+QFY+SF+ G +Y++ D V +++ EP IG++IKIWET
Sbjct: 207 FAWGKKRGMGGKKKDVQFYESFSFDGAEYAINDTVCLQSGIGGGEPHIGRLIKIWETRDK 386
Query: 75 ERKIKVQWFFRPIEVSKCLTWIKIYFNELFFAC-GDG-DGLATIHPLESIAGKCNIVCIS 132
RK+KVQWFFRP E+ K L I++ NELF AC GDG G A ++PLE+I GKCN+VCIS
Sbjct: 387 SRKVKVQWFFRPAEICKYLVGIEVKPNELFLACGGDGAKGFANVNPLEAIVGKCNVVCIS 566
Query: 133 KDSRNPQLSDKVTWSAAFVFYRYFDVGQRKIVEEVDDK-SIGIE 175
KD NPQ S + A +V+YR+FDV Q K+V+++D K + GIE
Sbjct: 567 KDVGNPQPSGEA--KADYVYYRFFDVVQLKVVDQIDVKVAAGIE 692
>TC233183 similar to UP|Q8IP81 (Q8IP81) CG31856-PA, partial (21%)
Length = 767
Score = 43.5 bits (101), Expect = 5e-05
Identities = 39/145 (26%), Positives = 65/145 (43%), Gaps = 5/145 (3%)
Frame = +2
Query: 24 GKG-GKKRDIQFYQSFTLRGVDYSLFDNVYV--KNDFAEPRIGKIIKIWETPTLERKIKV 80
GKG G+KR + SF G+ Y+L D V + + +P + I I ++ K+
Sbjct: 263 GKGRGRKRH---HDSFEFDGIQYTLEDPVLLVPEEKGQKPYVAIIKDITQSINGNVKVTG 433
Query: 81 QWFFRPIEVSK--CLTWIKIYFNELFFACGDGDGLATIHPLESIAGKCNIVCISKDSRNP 138
QWF+RP E K W ELF++ D P E++ KC + + + + P
Sbjct: 434 QWFYRPEEAEKKGGGNWQSCDTRELFYSFHRDD-----VPAEAVMHKCVVHFVPRHKQLP 598
Query: 139 QLSDKVTWSAAFVFYRYFDVGQRKI 163
+ D F+ + +D +RK+
Sbjct: 599 KRKD----HPGFIVQKVYDTVERKL 661
>TC207925 similar to UP|O65462 (O65462) Receptor like protein (Fragment),
partial (92%)
Length = 1280
Score = 30.0 bits (66), Expect = 0.60
Identities = 25/92 (27%), Positives = 48/92 (52%), Gaps = 5/92 (5%)
Frame = +3
Query: 2 SNSPPAPSPESETL-EFKWGQKRGKGGKKRDIQFYQSFTLRGVDYSLF--DNVYVK--ND 56
SNSPP P +++ E + + G+K D+ S+T+RG + + D V ++ +
Sbjct: 135 SNSPPKPFCWKQSIRETLHNMAKTRPGRK-DVD---SYTIRGTNKIVRAGDCVLMRPSDT 302
Query: 57 FAEPRIGKIIKIWETPTLERKIKVQWFFRPIE 88
P + ++ KI + K++V+W++RP E
Sbjct: 303 SKPPYVARVEKIEQDNRNNVKVRVRWYYRPEE 398
>TC214889 homologue to PIR|C86216|C86216 protein T23G18.6 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(97%)
Length = 1671
Score = 29.6 bits (65), Expect = 0.79
Identities = 17/48 (35%), Positives = 28/48 (57%)
Frame = -2
Query: 100 FNELFFACGDGDGLATIHPLESIAGKCNIVCISKDSRNPQLSDKVTWS 147
FN +F GLAT+H LE ++ + I+K S+NP+ + +TW+
Sbjct: 869 FNSIFNVNEGSLGLATVHKLEGLSSE---KIIAKASKNPR--NTLTWT 741
>BE059735 homologue to PIR|C86216|C86 protein T23G18.6 [imported] -
Arabidopsis thaliana, partial (37%)
Length = 440
Score = 29.6 bits (65), Expect = 0.79
Identities = 17/48 (35%), Positives = 28/48 (57%)
Frame = -2
Query: 100 FNELFFACGDGDGLATIHPLESIAGKCNIVCISKDSRNPQLSDKVTWS 147
FN +F GLAT+H LE ++ + I+K S+NP+ + +TW+
Sbjct: 256 FNSIFNVNEGSLGLATVHKLEGLSSE---KIIAKASKNPR--NTLTWT 128
>TC214891 homologue to PIR|C86216|C86216 protein T23G18.6 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(66%)
Length = 1257
Score = 27.7 bits (60), Expect = 3.0
Identities = 16/48 (33%), Positives = 27/48 (55%)
Frame = -2
Query: 100 FNELFFACGDGDGLATIHPLESIAGKCNIVCISKDSRNPQLSDKVTWS 147
FN +F GLAT+H LE ++ + I+K +NP+ + +TW+
Sbjct: 416 FNSIFNVNKGSLGLATVHKLEGLSSE---KIIAKACKNPR--NTLTWT 288
>CA853192
Length = 549
Score = 27.3 bits (59), Expect = 3.9
Identities = 14/41 (34%), Positives = 25/41 (60%), Gaps = 1/41 (2%)
Frame = -2
Query: 52 YVKNDFAEPRIGKI-IKIWETPTLERKIKVQWFFRPIEVSK 91
+V+N+F EP+IGKI I + P++ Q +F +++ K
Sbjct: 314 FVRNNFGEPKIGKIYINVMAKPSIS-----QVYFLKLDILK 207
>TC210174
Length = 861
Score = 26.9 bits (58), Expect = 5.1
Identities = 18/51 (35%), Positives = 24/51 (46%)
Frame = -3
Query: 44 DYSLFDNVYVKNDFAEPRIGKIIKIWETPTLERKIKVQWFFRPIEVSKCLT 94
+Y L + YVK F GK + + + KIK+ WFF P S C T
Sbjct: 835 NYQLQLSQYVKFFF----FGK*AHVSQVRLRKTKIKISWFF*PGPKSSCNT 695
>TC230401 similar to UP|Q8VZI6 (Q8VZI6) AT4g33380/F17M5_140, partial (13%)
Length = 537
Score = 26.2 bits (56), Expect = 8.7
Identities = 9/24 (37%), Positives = 17/24 (70%)
Frame = +2
Query: 74 LERKIKVQWFFRPIEVSKCLTWIK 97
+E I+++W F+P+E+ K W+K
Sbjct: 398 MEESIQIRWEFKPVELIK---WVK 460
>TC206546 weakly similar to UP|Q9LFD1 (Q9LFD1) Laccase-like protein, partial
(32%)
Length = 1281
Score = 26.2 bits (56), Expect = 8.7
Identities = 19/66 (28%), Positives = 27/66 (40%)
Frame = +2
Query: 24 GKGGKKRDIQFYQSFTLRGVDYSLFDNVYVKNDFAEPRIGKIIKIWETPTLERKIKVQWF 83
GKG + FY + D+ D + DF +P+I K TP K+K F
Sbjct: 530 GKGYSMLEASFYNVSGVYTTDFP--DKPPIIFDFTDPKIALDTKYLFTPPKSTKVKKLKF 703
Query: 84 FRPIEV 89
+EV
Sbjct: 704 NSTVEV 721
>TC222266
Length = 995
Score = 26.2 bits (56), Expect = 8.7
Identities = 11/39 (28%), Positives = 19/39 (48%)
Frame = +1
Query: 17 FKWGQKRGKGGKKRDIQFYQSFTLRGVDYSLFDNVYVKN 55
F+ RGK K+ FY++ G D+S+ ++ N
Sbjct: 103 FREKTPRGKWSKQETELFYEAVREMGTDFSMIQEIFFPN 219
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.140 0.446
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,611,481
Number of Sequences: 63676
Number of extensions: 147164
Number of successful extensions: 717
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 707
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 714
length of query: 178
length of database: 12,639,632
effective HSP length: 91
effective length of query: 87
effective length of database: 6,845,116
effective search space: 595525092
effective search space used: 595525092
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 56 (26.2 bits)
Medicago: description of AC148532.15


