

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148532.13 + phase: 0
(303 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
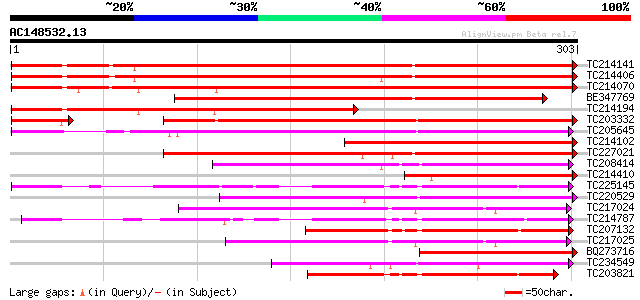
Score E
Sequences producing significant alignments: (bits) Value
TC214141 similar to UP|Q947D4 (Q947D4) Dehydration-responsive pr... 436 e-123
TC214406 weakly similar to UP|Q947D4 (Q947D4) Dehydration-respon... 427 e-120
TC214070 similar to UP|Q8VWQ1 (Q8VWQ1) Dehydration-induced prote... 419 e-117
BE347769 weakly similar to GP|16588826|gb dehydration-responsive... 247 4e-66
TC214194 weakly similar to UP|Q94IC5 (Q94IC5) BURP domain-contai... 239 1e-63
TC203332 weakly similar to UP|Q8H9E9 (Q8H9E9) Resistant specific... 234 5e-62
TC205645 UP|Q8RUD9 (Q8RUD9) Seed coat BURP domain protein 1, com... 213 7e-56
TC214102 similar to UP|Q9MB24 (Q9MB24) S1-3 protein (Fragment), ... 210 6e-55
TC227021 weakly similar to UP|O65009 (O65009) BURP domain contai... 186 1e-47
TC208414 weakly similar to UP|O65009 (O65009) BURP domain contai... 154 4e-38
TC214410 similar to UP|Q8VWQ1 (Q8VWQ1) Dehydration-induced prote... 149 1e-36
TC225145 homologue to UP|O24482 (O24482) Sali3-2, complete 141 3e-34
TC220529 weakly similar to GB|CAE02618.1|38567637|OSA575668 RAFT... 140 9e-34
TC217024 similar to UP|Q40161 (Q40161) Polygalacturonase isoenzy... 135 3e-32
TC214787 UP|Q06765 (Q06765) ADR6 protein (Sali5-4a protein), com... 134 4e-32
TC207132 similar to UP|EA92_VICFA (P21747) Embryonic abundant pr... 132 3e-31
TC217025 similar to UP|Q40161 (Q40161) Polygalacturonase isoenzy... 127 5e-30
BQ273716 weakly similar to SP|Q08298|RD22_ Dehydration-responsiv... 119 1e-27
TC234549 weakly similar to UP|O65009 (O65009) BURP domain contai... 113 9e-26
TC203821 similar to UP|O24482 (O24482) Sali3-2, partial (52%) 109 2e-24
>TC214141 similar to UP|Q947D4 (Q947D4) Dehydration-responsive protein RD22
(Fragment), partial (89%)
Length = 1353
Score = 436 bits (1120), Expect = e-123
Identities = 221/324 (68%), Positives = 258/324 (79%), Gaps = 22/324 (6%)
Frame = +1
Query: 2 LPPQLYWKSMLPNSPMPKAITNLLHPAGYWSEEKGTWVDVGKGGVDVGVRKGYYEGGGTD 61
LPP++YWKS+LP +PMPKAIT++L+ W EEK + V VG GGV+V KG G GT
Sbjct: 142 LPPEVYWKSVLPTTPMPKAITDILYSD--WVEEKSSSVHVGGGGVNVHTGKG--GGSGTT 309
Query: 62 VNVG---------------------VG-RSPFIYNYAASETQLHDKPNVALFFLEKDLHH 99
VNVG VG +SPF Y YAA+ETQLHD PNVALFFLEKDLH
Sbjct: 310 VNVGGKGGGGVNVHAGHKGKPVHVSVGSKSPFDYVYAATETQLHDDPNVALFFLEKDLHS 489
Query: 100 GTKLNLQFSKTTSNAATFLPRQVANSIPFSSNKMEYIINKLNIKKGSKGVQIVKNTISEC 159
GTKL+L F+++TSN ATFL RQVA+SIPFSSNK+++I NK ++K GS+ QI+KNTISEC
Sbjct: 490 GTKLDLHFTRSTSNQATFLSRQVADSIPFSSNKVDFIFNKFSVKPGSEEAQIMKNTISEC 669
Query: 160 EEQGIKGEEKVCVTSLESMVDFTTSKLGKNVEAVSTEVNKESNLQQYTIASGVKKLGEKN 219
EE GIKGEEK C TSLESMVDF+TSKLG NVE VSTEV+KE+ LQ+YT+A GVKKL +
Sbjct: 670 EEGGIKGEEKYCATSLESMVDFSTSKLGNNVEVVSTEVDKETGLQKYTVAPGVKKL-SGD 846
Query: 220 KAVVCHKENYPYAVFYCHKTDTTKAYSVPLEGADGSRVKAIAVCHTDTSEWNPKHLAFQV 279
KAVVCHK+NYPYAVFYCHKT+TT+AYSVPLEG +G RVKA+AVCHTDTSEWNPKHLAFQV
Sbjct: 847 KAVVCHKQNYPYAVFYCHKTETTRAYSVPLEGTNGVRVKAVAVCHTDTSEWNPKHLAFQV 1026
Query: 280 LKVQPGTVPVCHLLPEDHVVWIRK 303
LKV+PGT+PVCH LPEDHVVW+ K
Sbjct: 1027LKVKPGTIPVCHFLPEDHVVWVPK 1098
>TC214406 weakly similar to UP|Q947D4 (Q947D4) Dehydration-responsive protein
RD22 (Fragment), partial (90%)
Length = 1379
Score = 427 bits (1099), Expect = e-120
Identities = 219/327 (66%), Positives = 259/327 (78%), Gaps = 25/327 (7%)
Frame = +1
Query: 2 LPPQLYWKSMLPNSPMPKAITNLLHPAGYWSEEKGTWVDVGKGGVDVGVRKGYYEGGGTD 61
LPP++YWKS+LP +PMPKAIT++L+P W EEK T V+VG GV+V KG GGGT+
Sbjct: 190 LPPEVYWKSVLPTTPMPKAITDILYPD--WVEEKSTSVNVGGKGVNVHAGKG---GGGTN 354
Query: 62 VNVG----------------------VG-RSPFIYNYAASETQLHDKPNVALFFLEKDLH 98
VNVG VG +SPF Y YA++ETQLHD PNVALFFLEKDLH
Sbjct: 355 VNVGGKGSGGGVNVHAGHKGKPVHVSVGSKSPFNYIYASTETQLHDDPNVALFFLEKDLH 534
Query: 99 HGTKLNLQFSKTTSNAATFLPRQVANSIPFSSNKMEYIINKLNIKKGSKGVQIVKNTISE 158
GTKLNL F+ +++ ATFLPRQVA+SIPFSS+K+E + NK ++K GS+ QI+KNT+SE
Sbjct: 535 PGTKLNLHFTTSSNIQATFLPRQVADSIPFSSSKVEVVFNKFSVKPGSEEAQIMKNTLSE 714
Query: 159 CEEQGIKGEEKVCVTSLESMVDFTTSKLGKNVEAVSTEV--NKESNLQQYTIASGVKKLG 216
CEE GIKGEEK C TSLESM+DF+TSKLGKNVE VSTEV +KE+ LQ+YT+A GV KL
Sbjct: 715 CEEGGIKGEEKYCATSLESMIDFSTSKLGKNVEVVSTEVVEDKETGLQKYTVAPGVNKL- 891
Query: 217 EKNKAVVCHKENYPYAVFYCHKTDTTKAYSVPLEGADGSRVKAIAVCHTDTSEWNPKHLA 276
+KAVVCHK+NYPYAVFYCHKT+TT+AYSVPLEGA+G RVKA+AVCHT TSEWNPKHLA
Sbjct: 892 SGDKAVVCHKQNYPYAVFYCHKTETTRAYSVPLEGANGVRVKAVAVCHTHTSEWNPKHLA 1071
Query: 277 FQVLKVQPGTVPVCHLLPEDHVVWIRK 303
FQVLKV+PGTVPVCH LPEDHVVW+ K
Sbjct: 1072FQVLKVKPGTVPVCHFLPEDHVVWVPK 1152
>TC214070 similar to UP|Q8VWQ1 (Q8VWQ1) Dehydration-induced protein RD22-like
protein, partial (81%)
Length = 1427
Score = 419 bits (1076), Expect = e-117
Identities = 224/347 (64%), Positives = 257/347 (73%), Gaps = 45/347 (12%)
Frame = +3
Query: 2 LPPQLYWKSMLPNSPMPKAITNLLHPAGYWSEEK---------------------GTWVD 40
LPP++YWKS LP +PMPKAIT++LHP +E+K GT V+
Sbjct: 165 LPPEVYWKSKLPTTPMPKAITDILHPD--LAEDKSTSVAVGKGGVNVNAGKTKPGGTSVN 338
Query: 41 VGKGGVDVGVRKGYYEGGGTDVNVGVG--------------------RSPFIYNYAASET 80
VGKGGV+V KG GT VNVG G SPF YNYAASET
Sbjct: 339 VGKGGVNVNTGKG-KPNKGTSVNVGKGGVNVNTGPKKGKPVHVGVGPHSPFDYNYAASET 515
Query: 81 QLHDKPNVALFFLEKDLHHGTKLNLQFSK--TTSNAATFLPRQVANSIPFSSNKMEYIIN 138
Q HD PNVALFFLEKDLH+GTKLNL F++ T+S A+FLPR VA+SIPFSSNK+ ++N
Sbjct: 516 QWHDDPNVALFFLEKDLHYGTKLNLHFTRYFTSSVDASFLPRSVADSIPFSSNKVNEVLN 695
Query: 139 KLNIKKGSKGVQIVKNTISECEEQGIKGEEKVCVTSLESMVDFTTSKLGKN-VEAVSTEV 197
K +IK+GS Q VKNTISECE GIKGEEK CVTSLESMVDF T+KLG N V+AVSTEV
Sbjct: 696 KFSIKEGSDEAQTVKNTISECEVPGIKGEEKRCVTSLESMVDFATTKLGSNDVDAVSTEV 875
Query: 198 NKESN-LQQYTIASGVKKLGEKNKAVVCHKENYPYAVFYCHKTDTTKAYSVPLEGADGSR 256
K+ N LQQYT+A GVK+LGE +VVCHKENYPYAVFYCHK++TTKAYSVPLEGADGSR
Sbjct: 876 TKKDNELQQYTMAPGVKRLGEDKASVVCHKENYPYAVFYCHKSETTKAYSVPLEGADGSR 1055
Query: 257 VKAIAVCHTDTSEWNPKHLAFQVLKVQPGTVPVCHLLPEDHVVWIRK 303
VKA+AVCHTDTS+WNPKHLAFQVLKVQPGTVPVCH LP+DHVV++ K
Sbjct: 1056VKAVAVCHTDTSKWNPKHLAFQVLKVQPGTVPVCHFLPQDHVVFVPK 1196
>BE347769 weakly similar to GP|16588826|gb dehydration-responsive protein
RD22 {Prunus persica}, partial (38%)
Length = 595
Score = 247 bits (631), Expect = 4e-66
Identities = 127/199 (63%), Positives = 156/199 (77%)
Frame = +2
Query: 89 ALFFLEKDLHHGTKLNLQFSKTTSNAATFLPRQVANSIPFSSNKMEYIINKLNIKKGSKG 148
AL+ +E DLH GTKL+L F+++TSN ATFL RQVA+SIPFSSNK+++I NK + K GSK
Sbjct: 2 ALYLVEFDLHSGTKLDLHFTRSTSNQATFLSRQVADSIPFSSNKVDFIFNKNSGKPGSKE 181
Query: 149 VQIVKNTISECEEQGIKGEEKVCVTSLESMVDFTTSKLGKNVEAVSTEVNKESNLQQYTI 208
QI+KNTI ECEE GIKGEEK C TSLESM DF+TSKLG NVE VSTEV+K++ +YT
Sbjct: 182 AQIMKNTIIECEEGGIKGEEKYCATSLESMDDFSTSKLGNNVEVVSTEVDKDTG*HKYTG 361
Query: 209 ASGVKKLGEKNKAVVCHKENYPYAVFYCHKTDTTKAYSVPLEGADGSRVKAIAVCHTDTS 268
A GVKK+ +KAVVCHK NYPYA FYC++TDTT AYSVP EG + R KA++ CHTDTS
Sbjct: 362 APGVKKV-SGDKAVVCHKHNYPYADFYCNRTDTTIAYSVPSEGTNVVRDKAVSRCHTDTS 538
Query: 269 EWNPKHLAFQVLKVQPGTV 287
+ P+H+A VL V P ++
Sbjct: 539 DMYPRHVANHVLTVIPRSI 595
>TC214194 weakly similar to UP|Q94IC5 (Q94IC5) BURP domain-containing protein
(Fragment), partial (49%)
Length = 704
Score = 239 bits (609), Expect = 1e-63
Identities = 132/209 (63%), Positives = 147/209 (70%), Gaps = 24/209 (11%)
Frame = +1
Query: 2 LPPQLYWKSMLPNSPMPKAITNLLHPAGYWSEEKGTWVDVGKGGVDVGVRKGYY-EGGGT 60
LPP YWKS LP +PMPKAIT+LL P W EEK T VDVGKGGV+VGV KG G GT
Sbjct: 82 LPPAFYWKSKLPTTPMPKAITDLLQPD--WKEEKDTSVDVGKGGVNVGVEKGNQGPGDGT 255
Query: 61 DVNVGVG---------------------RSPFIYNYAASETQLHDKPNVALFFLEKDLHH 99
DVNVG G SPF YNYAASETQLHD PNVALFFLEKDLHH
Sbjct: 256 DVNVGGGDGGVNVHTGPKGKPVHVGVGPHSPFDYNYAASETQLHDDPNVALFFLEKDLHH 435
Query: 100 GTKLNLQFS--KTTSNAATFLPRQVANSIPFSSNKMEYIINKLNIKKGSKGVQIVKNTIS 157
GTKLNL F+ T++ ATFLPR V++SIPFSSNK+ ++NK +IK GS + VKNTI+
Sbjct: 436 GTKLNLHFTIYYTSNVDATFLPRSVSDSIPFSSNKVNDVLNKFSIKDGSDEAKTVKNTIN 615
Query: 158 ECEEQGIKGEEKVCVTSLESMVDFTTSKL 186
ECE IKGEEK CVTSLESMVDF T+KL
Sbjct: 616 ECEGPSIKGEEKRCVTSLESMVDFATTKL 702
>TC203332 weakly similar to UP|Q8H9E9 (Q8H9E9) Resistant specific protein-2,
partial (60%)
Length = 1444
Score = 234 bits (596), Expect = 5e-62
Identities = 113/221 (51%), Positives = 144/221 (65%)
Frame = +1
Query: 83 HDKPNVALFFLEKDLHHGTKLNLQFSKTTSNAATFLPRQVANSIPFSSNKMEYIINKLNI 142
HD P F E+ L GTKL+ F K N LPRQ+A IP SS K++ I+ L +
Sbjct: 583 HDIPKADQVFFEEGLRPGTKLDAHFKKR-ENVTPLLPRQIAQHIPLSSAKIKEIVEMLFV 759
Query: 143 KKGSKGVQIVKNTISECEEQGIKGEEKVCVTSLESMVDFTTSKLGKNVEAVSTEVNKESN 202
+ V+I++ TIS CE I GEE+ C TSLESMVDF TSKLGKN +STE KES
Sbjct: 760 NPEPENVKILEETISMCEVPAITGEERYCATSLESMVDFVTSKLGKNARVISTEAEKESK 939
Query: 203 LQQYTIASGVKKLGEKNKAVVCHKENYPYAVFYCHKTDTTKAYSVPLEGADGSRVKAIAV 262
Q++++ GVK L E +K +VCH +YPY VF CH+ T A+ +PLEG DG+RVKA AV
Sbjct: 940 SQKFSVKDGVKLLAE-DKVIVCHPMDYPYVVFMCHEISNTTAHFMPLEGEDGTRVKAAAV 1116
Query: 263 CHTDTSEWNPKHLAFQVLKVQPGTVPVCHLLPEDHVVWIRK 303
CH DTSEW+P H+ Q+LK +PG PVCH+ PE H++W K
Sbjct: 1117CHKDTSEWDPNHVFLQMLKTKPGAAPVCHIFPEGHLLWFAK 1239
Score = 43.9 bits (102), Expect = 9e-05
Identities = 18/37 (48%), Positives = 28/37 (75%), Gaps = 4/37 (10%)
Frame = +1
Query: 2 LPPQLYWKSMLPNSPMPKAITNLLH----PAGYWSEE 34
+PP++YW+ MLPN+PMPKAI + L+ P GY +++
Sbjct: 79 IPPEVYWERMLPNTPMPKAIIDFLNLDQLPLGYGAKK 189
>TC205645 UP|Q8RUD9 (Q8RUD9) Seed coat BURP domain protein 1, complete
Length = 1369
Score = 213 bits (543), Expect = 7e-56
Identities = 119/307 (38%), Positives = 166/307 (53%), Gaps = 7/307 (2%)
Frame = +1
Query: 2 LPPQLYWKSMLPNSPMPKAITNLLHPAGYWSEEKGTWVDVGKGGVDVGVRKGYYEGGGTD 61
L P+ YW++MLP +P+PKAIT LL + +E G D
Sbjct: 103 LTPRHYWETMLPRTPLPKAITELLSLES----------------------RSIFEYAGND 216
Query: 62 VNVGVGRSPFIYNYAASETQLHD--KPNV----ALFFLEKDLHHGTKLNLQFSKTTSNAA 115
S I YA D K N+ LFFLE+DL G N++F T
Sbjct: 217 ---DQSESRSILGYAGYNQDEDDVSKHNIQIFNRLFFLEEDLRAGKIFNMKFVNNTKATV 387
Query: 116 TFLPRQVANSIPFSSNKMEYIINKLNIKKGSKGVQIVKNTISECEEQGIKGEEKVCVTSL 175
LPRQ++ IPFS +K + ++ L ++ S +I+ TI C+E +GE K C TSL
Sbjct: 388 PLLPRQISKQIPFSEDKKKQVLAMLGVEANSSNAKIIAETIGLCQEPATEGERKHCATSL 567
Query: 176 ESMVDFTTSKLGKNVEAVSTEVNKESNLQQYTIA-SGVKKLGEKNKAVVCHKENYPYAVF 234
ESMVDF S LGKNV A STE +E+ ++ + +GV+KLG+ +K + CH +YPY VF
Sbjct: 568 ESMVDFVVSALGKNVGAFSTEKERETESGKFVVVKNGVRKLGD-DKVIACHPMSYPYVVF 744
Query: 235 YCHKTDTTKAYSVPLEGADGSRVKAIAVCHTDTSEWNPKHLAFQVLKVQPGTVPVCHLLP 294
CH + Y V L+G DG RVKA+ CH DTS+W+ H AF+VL ++PG VCH+
Sbjct: 745 GCHLVPRSSGYLVRLKGEDGVRVKAVVACHRDTSKWDHNHGAFKVLNLKPGNGTVCHVFT 924
Query: 295 EDHVVWI 301
E +++W+
Sbjct: 925 EGNLLWL 945
>TC214102 similar to UP|Q9MB24 (Q9MB24) S1-3 protein (Fragment), partial
(70%)
Length = 621
Score = 210 bits (535), Expect = 6e-55
Identities = 99/126 (78%), Positives = 113/126 (89%), Gaps = 2/126 (1%)
Frame = +3
Query: 180 DFTTSKLGKN-VEAVSTEVNKESN-LQQYTIASGVKKLGEKNKAVVCHKENYPYAVFYCH 237
DF T+KLG N V+AVSTEV K+ N LQQYT+A GVK+LGE +VVCHKENYPYAVFYCH
Sbjct: 15 DFATTKLGSNDVDAVSTEVTKKDNELQQYTMAPGVKRLGEDKASVVCHKENYPYAVFYCH 194
Query: 238 KTDTTKAYSVPLEGADGSRVKAIAVCHTDTSEWNPKHLAFQVLKVQPGTVPVCHLLPEDH 297
K++TTKAYSVPLEGADGSRVKA+AVCHTDTS+WNPKHLAFQVLKV PGTVP+CH LP+DH
Sbjct: 195 KSETTKAYSVPLEGADGSRVKAVAVCHTDTSKWNPKHLAFQVLKVHPGTVPICHFLPQDH 374
Query: 298 VVWIRK 303
VV++ K
Sbjct: 375 VVFVPK 392
>TC227021 weakly similar to UP|O65009 (O65009) BURP domain containing
protein, partial (47%)
Length = 1147
Score = 186 bits (471), Expect = 1e-47
Identities = 96/227 (42%), Positives = 141/227 (61%), Gaps = 6/227 (2%)
Frame = +1
Query: 83 HDKPNVALFFLEKDLHHGTKLNLQFSKTTSNAATFL-PRQVANSIPFSSNKMEYIINKLN 141
H P+V +FF +DL G + + F K + L PR+ A+S+PFS NK+ ++ +
Sbjct: 295 HIDPSVMVFFTIEDLKVGKTMPIHFPKRDPATSPKLWPREEADSLPFSLNKLPNLLKIFS 474
Query: 142 IKKGSKGVQIVKNTISECEEQGIKGEEKVCVTSLESMVDFTTSKLG--KNVEAVSTEVNK 199
+ + S + +++T+ ECE + IKGE K C TSLESM+DFT S LG +++ +ST
Sbjct: 475 VSQNSPKAKAMEDTLRECETKPIKGEVKFCATSLESMLDFTQSILGFTSDLKVLSTSHQT 654
Query: 200 ESNL--QQYTIASGVKKLGEKNKAVVCHKENYPYAVFYCHKTDT-TKAYSVPLEGADGSR 256
+S++ Q YT+ + ++ +K V CH YPY VFYCH ++ K Y VPL G +G R
Sbjct: 655 KSSVTFQNYTMLENIIEI-PASKMVACHTMPYPYTVFYCHSQESENKIYRVPLAGENGDR 831
Query: 257 VKAIAVCHTDTSEWNPKHLAFQVLKVQPGTVPVCHLLPEDHVVWIRK 303
V A+ VCH DTS+W H++FQVLKV+PGT VCH P DH++W+ K
Sbjct: 832 VDAMVVCHMDTSQWGHGHVSFQVLKVKPGTTSVCHFFPADHLIWVPK 972
>TC208414 weakly similar to UP|O65009 (O65009) BURP domain containing
protein, partial (39%)
Length = 957
Score = 154 bits (390), Expect = 4e-38
Identities = 85/198 (42%), Positives = 116/198 (57%), Gaps = 5/198 (2%)
Frame = +2
Query: 109 KTTSNAATFLPRQVANSIPFSSNKMEYIINKLNIKKGSKGVQIVKNTISECEEQGIKGEE 168
KT + F P + FSS + ++ +I K S + +K T+ +CE + ++GE
Sbjct: 215 KTPQHLQNF*PEKKLTRFLFSSKHLPSLLKFFSIPKHSPQAKAMKYTLKQCEFEPMEGET 394
Query: 169 KVCVTSLESMVDFTTSKLGKNVE-AVSTEV---NKESNLQQYTIASGVKKLGEKNKAVVC 224
K C TSLES+ DF G N + V T V N + LQ YTI S VK + N + C
Sbjct: 395 KFCATSLESLFDFAHYLFGSNAQFKVLTTVHLTNSTTLLQNYTI-SEVKVISVPN-VIGC 568
Query: 225 HKENYPYAVFYCHKTDT-TKAYSVPLEGADGSRVKAIAVCHTDTSEWNPKHLAFQVLKVQ 283
H YPYAVFYCH + T Y V +EG +G RV+A A+CH DTS+W+ H++F+VLKVQ
Sbjct: 569 HPMPYPYAVFYCHSQHSDTNLYEVMVEGENGGRVQAAAICHMDTSKWDRDHVSFRVLKVQ 748
Query: 284 PGTVPVCHLLPEDHVVWI 301
PGT PVCH P D++VW+
Sbjct: 749 PGTSPVCHFFPPDNLVWV 802
Score = 30.4 bits (67), Expect = 1.0
Identities = 20/59 (33%), Positives = 30/59 (49%), Gaps = 2/59 (3%)
Frame = +1
Query: 72 IYNYAAS-ETQLHDKPNVALFFLEKDLHHGTKLNLQFS-KTTSNAATFLPRQVANSIPF 128
++ Y S E H + +FF DL G + + FS K +S + FL R+ A+ IPF
Sbjct: 97 VHGYKPSHEDHKHMDLELNVFFTPNDLKVGKIMPIYFSKKNSSTSPKFLTREEADQIPF 273
>TC214410 similar to UP|Q8VWQ1 (Q8VWQ1) Dehydration-induced protein RD22-like
protein, partial (22%)
Length = 575
Score = 149 bits (377), Expect = 1e-36
Identities = 70/96 (72%), Positives = 81/96 (83%), Gaps = 4/96 (4%)
Frame = +1
Query: 212 VKKLGEKNKAVVC----HKENYPYAVFYCHKTDTTKAYSVPLEGADGSRVKAIAVCHTDT 267
+K L +KN +V + PYAVFYCHK++TTKAYSVPLEGADGSRVKA+AVCHTDT
Sbjct: 55 IKILQKKNSRLVLSLVPNSARGPYAVFYCHKSETTKAYSVPLEGADGSRVKAVAVCHTDT 234
Query: 268 SEWNPKHLAFQVLKVQPGTVPVCHLLPEDHVVWIRK 303
S+WNPKHLAFQVLKVQPGTVPVCH LP+DHVV++ K
Sbjct: 235 SKWNPKHLAFQVLKVQPGTVPVCHFLPQDHVVFVPK 342
>TC225145 homologue to UP|O24482 (O24482) Sali3-2, complete
Length = 1382
Score = 141 bits (356), Expect = 3e-34
Identities = 98/301 (32%), Positives = 146/301 (47%), Gaps = 1/301 (0%)
Frame = +3
Query: 2 LPPQLYWKSMLPNSPMPKAITNLLHPAGYWSEEKGTWVDVGKGGVDVGVRKGYYEGGGTD 61
LP + YW+++ PN+P+P A+ LL P GV++
Sbjct: 255 LPEEDYWEAVWPNTPIPTALRELLKPL--------------PAGVEID------------ 356
Query: 62 VNVGVGRSPFIYNYAASETQLHDKPNVALFFLEKDLHHGTKLNLQFSKTTSNAATFLPRQ 121
Q+ D FF ++DLH G + +QF+K A +
Sbjct: 357 ---------------ELPKQIDDTQYPKTFFYKEDLHPGKTMKVQFTKRPY-AQPYGVYT 488
Query: 122 VANSIPFSSNKMEYIINKLNIKKGSKGVQIVKNTISECEEQGIKGEEKVCVTSLESMVDF 181
I +S K Y ++ IKK + +GEEK C SL +++ F
Sbjct: 489 WLTDIKDTS-KEGYSFEEICIKK-----------------EAFEGEEKFCAKSLGTVIGF 614
Query: 182 TTSKLGKNVEAVSTE-VNKESNLQQYTIASGVKKLGEKNKAVVCHKENYPYAVFYCHKTD 240
SKLGKN++ +S+ VNK+ +QYT+ GV+ LG+K AV+CH N+ AVFYCHK
Sbjct: 615 AISKLGKNIQVLSSSFVNKQ---EQYTV-EGVQNLGDK--AVMCHGLNFRTAVFYCHKVR 776
Query: 241 TTKAYSVPLEGADGSRVKAIAVCHTDTSEWNPKHLAFQVLKVQPGTVPVCHLLPEDHVVW 300
T A+ VPL DG++ +A+AVCH+DTS N H+ +++ V PGT PVCH L ++W
Sbjct: 777 ETTAFMVPLVAGDGTKTQALAVCHSDTSGMN-HHMLHELMGVDPGTNPVCHFLGSKAILW 953
Query: 301 I 301
+
Sbjct: 954 V 956
>TC220529 weakly similar to GB|CAE02618.1|38567637|OSA575668 RAFTIN1 protein
{Oryza sativa (japonica cultivar-group);} , partial
(15%)
Length = 942
Score = 140 bits (352), Expect = 9e-34
Identities = 69/197 (35%), Positives = 115/197 (58%), Gaps = 6/197 (3%)
Frame = +3
Query: 113 NAATFLPRQVANSIPFSSNKMEYIINKLNIKKGSKGVQIVKNTISECEEQGIKGEEKVCV 172
+ + FLPR+ A SIPFS +++ ++ +I + S +++T+ +CE + I GE K+C
Sbjct: 204 DVSQFLPRKEAESIPFSISQLPSVLQLFSISEDSPEANAMRDTLEQCEAEPITGETKICA 383
Query: 173 TSLESMVDFTTSKLGK----NVEAVSTEVNKESNLQQYTIASGVKKLGEKNKAVVCHKEN 228
TSLESM++F + +G N+ + LQ++TI + + K V CH
Sbjct: 384 TSLESMLEFIGTIIGSETKHNILTTTLPTASGVPLQKFTILEVSEDINAA-KWVACHPLP 560
Query: 229 YPYAVFYCHKTDT-TKAYSVPLEGADG-SRVKAIAVCHTDTSEWNPKHLAFQVLKVQPGT 286
YPYA++YCH T +K + V L +G +++A+ +CH DTS+W+P H+ F+ L ++PG
Sbjct: 561 YPYAIYYCHFIATGSKVFKVSLGSENGDDKIEALGICHLDTSDWSPNHIIFRQLGIKPGK 740
Query: 287 VPVCHLLPEDHVVWIRK 303
VCH P H++W+ K
Sbjct: 741 DAVCHFFPIKHLMWVPK 791
>TC217024 similar to UP|Q40161 (Q40161) Polygalacturonase isoenzyme 1 beta
subunit precursor , partial (33%)
Length = 958
Score = 135 bits (339), Expect = 3e-32
Identities = 74/215 (34%), Positives = 116/215 (53%), Gaps = 5/215 (2%)
Frame = +1
Query: 91 FFLEKDLHHGTKLNLQFSKTTSNAATFLPRQVANSIPFSSNKMEYIINKLNIKKGSKGVQ 150
FF E L GT + + + +FLPR + +PFSS K++ + + + S +
Sbjct: 85 FFRETMLKEGTVMPMPDIRDKMPKRSFLPRAILTKLPFSSAKVDELKRVFKVSENSSMDK 264
Query: 151 IVKNTISECEEQGIKGEEKVCVTSLESMVDFTTSKLGKNVEAVSTEVNKESNLQQYTIAS 210
++ +++ ECE GE K CV S+E M+DF+TS LG+NV +TE K SN + +
Sbjct: 265 MIMDSLGECERAPSVGETKRCVASVEDMIDFSTSVLGRNVAVWTTENVKGSN--KNVMVG 438
Query: 211 GVKKL--GEKNKAVVCHKENYPYAVFYCHKTDTTKAYSVPLEGADGSRVK---AIAVCHT 265
VK + G+ K+V CH+ +PY ++YCH + Y L + S+ K +A+CH
Sbjct: 439 RVKGMNGGKVTKSVSCHQSLFPYLLYYCHSLPKVRVYEANLLDPE-SKAKINHGVAICHL 615
Query: 266 DTSEWNPKHLAFQVLKVQPGTVPVCHLLPEDHVVW 300
DT+ W+P H AF L PG + VCH + E+ + W
Sbjct: 616 DTTAWSPTHGAFLALGSGPGRIEVCHWIFENDLTW 720
>TC214787 UP|Q06765 (Q06765) ADR6 protein (Sali5-4a protein), complete
Length = 1150
Score = 134 bits (338), Expect = 4e-32
Identities = 93/300 (31%), Positives = 147/300 (49%), Gaps = 5/300 (1%)
Frame = +3
Query: 7 YWKSMLPNSPMPKAITNLLHPAGYWSEEKGTWVDVGKGGVDVGVRKGYYEGGGTDVNVGV 66
+W ++ PN+P+P ++ +LL P +V +
Sbjct: 141 FWHAVWPNTPIPSSLRDLLKPG--------------------------------PASVEI 224
Query: 67 GRSPFIYNYAASETQLHDKPNVALFFLEKDLHHGTKLNLQFSKTTSN----AATFLPRQV 122
P Q+ + FF ++DLH G + +QFSK T+L +++
Sbjct: 225 DDHPM---------QIEETQYPKTFFYKEDLHPGKTMKVQFSKPPFQQPWGVGTWL-KEI 374
Query: 123 ANSIPFSSNKMEYIINKLNIKKGSKGVQIVKNTISECEEQGIKGEEKVCVTSLESMVDFT 182
++ K Y +L IKK + I+GEEK C SL +++ F
Sbjct: 375 KDT-----TKEGYSFEELCIKK-----------------EAIEGEEKFCAKSLGTVIGFA 488
Query: 183 TSKLGKNVEAVSTE-VNKESNLQQYTIASGVKKLGEKNKAVVCHKENYPYAVFYCHKTDT 241
SKLGKN++ +S+ VNK+ QYT+ GV+ LG+K AV+CH+ N+ AVFYCH+
Sbjct: 489 ISKLGKNIQVLSSSFVNKQD---QYTV-EGVQNLGDK--AVMCHRLNFRTAVFYCHEVRE 650
Query: 242 TKAYSVPLEGADGSRVKAIAVCHTDTSEWNPKHLAFQVLKVQPGTVPVCHLLPEDHVVWI 301
T A+ VPL DG++ +A+A+CH++TS N + L Q++ V PGT PVCH L ++W+
Sbjct: 651 TTAFMVPLVAGDGTKTQALAICHSNTSGMNHQML-HQLMGVDPGTNPVCHFLGSKAILWV 827
>TC207132 similar to UP|EA92_VICFA (P21747) Embryonic abundant protein USP92
precursor, partial (51%)
Length = 847
Score = 132 bits (331), Expect = 3e-31
Identities = 64/143 (44%), Positives = 94/143 (64%)
Frame = +3
Query: 159 CEEQGIKGEEKVCVTSLESMVDFTTSKLGKNVEAVSTEVNKESNLQQYTIASGVKKLGEK 218
C + KGEEK C TSL+SM+ F SKLGKN++A+S+ ++ + QY + V K+GEK
Sbjct: 423 CGKAAAKGEEKFCATSLQSMMGFAISKLGKNIKAISSSFAQDHD--QYVVEE-VNKIGEK 593
Query: 219 NKAVVCHKENYPYAVFYCHKTDTTKAYSVPLEGADGSRVKAIAVCHTDTSEWNPKHLAFQ 278
AV+CH+ N+ VFYCH+ + T Y VPL +DG++ KA+ +CH DT +P + ++
Sbjct: 594 --AVMCHRLNFENVVFYCHQINATTTYMVPLVASDGTKAKALTICHHDTRGMDP-IVVYE 764
Query: 279 VLKVQPGTVPVCHLLPEDHVVWI 301
VLKV+ GTVPVCH + + W+
Sbjct: 765 VLKVKTGTVPVCHFVGNKAIAWV 833
>TC217025 similar to UP|Q40161 (Q40161) Polygalacturonase isoenzyme 1 beta
subunit precursor , partial (32%)
Length = 933
Score = 127 bits (320), Expect = 5e-30
Identities = 66/190 (34%), Positives = 105/190 (54%), Gaps = 5/190 (2%)
Frame = +2
Query: 116 TFLPRQVANSIPFSSNKMEYIINKLNIKKGSKGVQIVKNTISECEEQGIKGEEKVCVTSL 175
+FLPR + +PFSS+K+ + + S +++ +++ ECE GE K CV S+
Sbjct: 41 SFLPRSILTKLPFSSSKVHELKRLFKVSDNSSMEKMIMDSLGECERVPSMGETKRCVGSI 220
Query: 176 ESMVDFTTSKLGKNVEAVSTEVNKESNLQQYTIASGVKKL--GEKNKAVVCHKENYPYAV 233
E M+DF+TS LG+NV +TE SN + + VK + G+ ++V CH+ +PY +
Sbjct: 221 EDMIDFSTSVLGRNVAVWTTENVNGSN--KNVMVGRVKGMNGGKVTQSVSCHQSLFPYML 394
Query: 234 FYCHKTDTTKAYSVPLEGADGSRVK---AIAVCHTDTSEWNPKHLAFQVLKVQPGTVPVC 290
+YCH + Y L + S+ K +A+CH DT+ W+P H AF L PG + VC
Sbjct: 395 YYCHSVPKVRVYQADLLDPE-SKAKINHGVAICHLDTTAWSPTHGAFMALGSGPGRIEVC 571
Query: 291 HLLPEDHVVW 300
H + E+ + W
Sbjct: 572 HWIFENDLTW 601
>BQ273716 weakly similar to SP|Q08298|RD22_ Dehydration-responsive protein
RD22 precursor. [Mouse-ear cress] {Arabidopsis
thaliana}, partial (13%)
Length = 419
Score = 119 bits (299), Expect = 1e-27
Identities = 48/84 (57%), Positives = 61/84 (72%)
Frame = +1
Query: 220 KAVVCHKENYPYAVFYCHKTDTTKAYSVPLEGADGSRVKAIAVCHTDTSEWNPKHLAFQV 279
K +VCH +YPY VF CH+ T A+ +PLEG DG+RVKA AVCH DTSEW+P H+ Q+
Sbjct: 67 KVIVCHPMDYPYVVFMCHEISNTTAHFMPLEGEDGTRVKAAAVCHKDTSEWDPNHVFLQM 246
Query: 280 LKVQPGTVPVCHLLPEDHVVWIRK 303
LK +PG PVCH+ PE H++W K
Sbjct: 247 LKTKPGAAPVCHIFPEGHLLWFAK 318
>TC234549 weakly similar to UP|O65009 (O65009) BURP domain containing
protein, partial (14%)
Length = 791
Score = 113 bits (283), Expect = 9e-26
Identities = 57/169 (33%), Positives = 96/169 (56%), Gaps = 8/169 (4%)
Frame = +1
Query: 141 NIKKGSKGVQIVKNTISECEEQGIKGEEKVCVTSLESMVDFTTSKLGKNVE--AVSTEVN 198
+I + S +++T+ +C + I GE K+C TSLESM++F +G + ++T +
Sbjct: 37 SISEDSPXANAMRDTLDQCXXEPITGETKICATSLESMLEFVGKIIGLETKHNIITTTLP 216
Query: 199 KESN--LQQYTIASGVKKLGEKNKAVVCHKENYPYAVFYCHKTDT-TKAYSVPL---EGA 252
S LQ++TI + + +K V CH YPYA++YCH T +K + V L
Sbjct: 217 TASGVPLQKFTILEVSEDINA-SKWVACHPLPYPYAIYYCHFIATGSKVFKVSLGSENNG 393
Query: 253 DGSRVKAIAVCHTDTSEWNPKHLAFQVLKVQPGTVPVCHLLPEDHVVWI 301
D +++A+ +CH DTS+W+P H+ F+ L ++PG VCH H++W+
Sbjct: 394 DDDKIEALGICHLDTSDWSPNHIIFRQLGIKPGKDSVCHFFTIKHLMWV 540
>TC203821 similar to UP|O24482 (O24482) Sali3-2, partial (52%)
Length = 952
Score = 109 bits (272), Expect = 2e-24
Identities = 59/135 (43%), Positives = 86/135 (63%), Gaps = 1/135 (0%)
Frame = +2
Query: 160 EEQGIKGEEKVCVTSLESMVDFTTSKLGKNVEAVSTE-VNKESNLQQYTIASGVKKLGEK 218
E+ +GEEK C SL +++ F SKLGKN++ +S+ VNK+ +QYT+ GV+ LG+K
Sbjct: 500 EQLWFEGEEKFCAKSLGTVIGFAISKLGKNIQVLSSSFVNKQ---EQYTV-EGVQNLGDK 667
Query: 219 NKAVVCHKENYPYAVFYCHKTDTTKAYSVPLEGADGSRVKAIAVCHTDTSEWNPKHLAFQ 278
AV+CH N+ AVFYCHK T A+ VPL DG++ A+A H++TS N H+ +
Sbjct: 668 --AVMCHGLNFRTAVFYCHKVHETTAFIVPLVAGDGTKTHALACSHSNTSGMN-HHMLHE 838
Query: 279 VLKVQPGTVPVCHLL 293
++ P T P+CH L
Sbjct: 839 LIVFNPKTNPICHFL 883
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.314 0.132 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,493,558
Number of Sequences: 63676
Number of extensions: 196712
Number of successful extensions: 1092
Number of sequences better than 10.0: 93
Number of HSP's better than 10.0 without gapping: 948
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1035
length of query: 303
length of database: 12,639,632
effective HSP length: 97
effective length of query: 206
effective length of database: 6,463,060
effective search space: 1331390360
effective search space used: 1331390360
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 59 (27.3 bits)
Medicago: description of AC148532.13


