

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148526.6 + phase: 0 /pseudo
(420 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
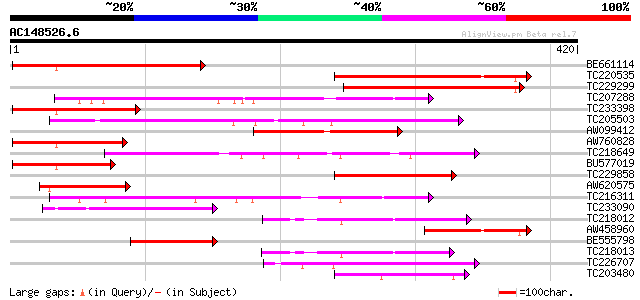
Score E
Sequences producing significant alignments: (bits) Value
BE661114 178 5e-45
TC220535 similar to UP|HXT5_YEAST (P38695) Probable glucose tran... 160 8e-40
TC229299 similar to UP|GT10_HUMAN (O95528) Solute carrier family... 140 1e-33
TC207288 UP|Q7XA51 (Q7XA51) Monosaccharide transporter, complete 130 1e-30
TC233398 125 4e-29
TC205503 UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, complete 112 2e-25
AW099412 110 1e-24
AW760828 107 8e-24
TC218649 similar to UP|Q39416 (Q39416) Integral membrane protein... 104 6e-23
BU577019 96 3e-20
TC229858 weakly similar to UP|Q8VHD6 (Q8VHD6) Glucose transporte... 92 3e-19
AW620575 86 2e-17
TC216311 UP|Q7XA52 (Q7XA52) Monosaccharide transporter, complete 84 2e-16
TC233090 similar to UP|Q9ZVN7 (Q9ZVN7) T7A14.10 protein, partial... 79 4e-15
TC218012 similar to UP|Q6VEF2 (Q6VEF2) Monosaccharide transporte... 72 4e-13
AW458960 70 2e-12
BE555798 homologue to GP|1353516|gb|A sugar transporter {Medicag... 67 2e-11
TC218013 similar to UP|Q6VEF2 (Q6VEF2) Monosaccharide transporte... 64 1e-10
TC226707 similar to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter... 64 1e-10
TC203480 similar to UP|Q8LPQ8 (Q8LPQ8) AT4g35300/F23E12_140, par... 63 3e-10
>BE661114
Length = 634
Score = 178 bits (451), Expect = 5e-45
Identities = 89/146 (60%), Positives = 115/146 (77%), Gaps = 3/146 (2%)
Frame = +3
Query: 3 EGGHPLADKTEFTECWRRTAESPYLMRLALS---GGLLFGYDTGVISGASLYIRDEFEQV 59
EGG P AD + F EC + ++PY++RLA S GGLLFGYDTGVISGA LYI+DEF+ V
Sbjct: 180 EGGVPEADMSAFRECLSLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIKDEFKAV 359
Query: 60 DKKPWLQETIVSMASAGAIIGAAFGGYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWV 119
D+K WLQE IVS A AGAIIGA+ GG++ND+ GRKK I++AD +F G+++MAAA +P +
Sbjct: 360 DRKTWLQEAIVSTAIAGAIIGASVGGWINDRFGRKKGIVIADTLFFIGSVIMAAASSPAI 539
Query: 120 IIIGRLLVGLGVGAASMTAPLYISEA 145
+I+GR+ VG+GV ASM +PLYISEA
Sbjct: 540 LIVGRVFVGIGVXMASMASPLYISEA 617
>TC220535 similar to UP|HXT5_YEAST (P38695) Probable glucose transporter
HXT5, partial (3%)
Length = 1007
Score = 160 bits (406), Expect = 8e-40
Identities = 82/148 (55%), Positives = 107/148 (71%), Gaps = 2/148 (1%)
Frame = +1
Query: 241 VRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLVTSGLNAVGTIVS 300
VRRGLYAG+ +Q+ QQ VGINT+MYYSPTIVQ AG ASN A LSLVT+GLNA G+I+S
Sbjct: 4 VRRGLYAGMGLQIFQQFVGINTVMYYSPTIVQLAGFASNRVALLLSLVTAGLNAFGSILS 183
Query: 301 MVLIDRFGRRKLMLISLIGIFVSLVTLSVTFNQAAHHAPSLSIQDSLSFGGNSTCKAYTT 360
+ ID+ GRRKL+L SL G+ VSLV L+V F++ H+P +S ++ F N+TC Y+T
Sbjct: 184 IYFIDKTGRRKLLLFSLCGVVVSLVVLTVAFHETTTHSPMVSTIETSHF--NNTCPDYST 357
Query: 361 APNHLSWNCMQCL--HEDCAFCANSQNE 386
A N W+CM+CL +C FCA+ N+
Sbjct: 358 AFNPGEWDCMKCLKASPECGFCASRANK 441
>TC229299 similar to UP|GT10_HUMAN (O95528) Solute carrier family 2,
facilitated glucose transporter, member 10 (Glucose
transporter type 10), partial (6%)
Length = 1060
Score = 140 bits (353), Expect = 1e-33
Identities = 70/136 (51%), Positives = 88/136 (64%), Gaps = 2/136 (1%)
Frame = +2
Query: 248 GITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLVTSGLNAVGTIVSMVLIDRF 307
G+ + + QQ VGINT+MYYSPTIVQ AG ASN TA LSL+ SGLNA G+I+S+ ID+
Sbjct: 2 GVGLLIFQQFVGINTVMYYSPTIVQLAGFASNRTALLLSLIISGLNAFGSILSIYFIDKT 181
Query: 308 GRRKLMLISLIGIFVSLVTLSVTFNQAAHHAPSLSIQDSLSFGGNSTCKAYTTAPNHLSW 367
GR+KL LISL G+ SL L+ F ++ H+P +S S F N+TC Y TA N W
Sbjct: 182 GRKKLALISLCGVVFSLALLTAAFRESEIHSPMVSAIQSSQFNNNNTCPDYKTALNSAEW 361
Query: 368 NCMQCL--HEDCAFCA 381
CM CL C +CA
Sbjct: 362 TCMTCLKASPSCGYCA 409
>TC207288 UP|Q7XA51 (Q7XA51) Monosaccharide transporter, complete
Length = 1813
Score = 130 bits (327), Expect = 1e-30
Identities = 101/330 (30%), Positives = 161/330 (48%), Gaps = 49/330 (14%)
Frame = +1
Query: 34 GGLLFGYDTGVISGASL---YIRDEFEQV--DKKPWLQET------------IVSMASAG 76
GG LFGYD GV G ++++ F +V K+ L ET S
Sbjct: 133 GGSLFGYDLGVSGGVPSMDDFLKEFFPKVYRRKQMHLHETDYCKYDDQVLTLFTSSLYFS 312
Query: 77 AIIGAAFGGYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASM 136
A++ F ++ K GRK +I++ + F+AGA++ AAA ++IIGR+L+G G+G +
Sbjct: 313 ALVMTFFASFLTRKKGRKASIIVGALSFLAGAILNAAAKNIAMLIIGRVLLGGGIGFGNQ 492
Query: 137 TAPLYISEASPAKIRGA-------------LVCRLHQWFTHN----RWSISL----LPHQ 175
PLY+SE +PAK RGA L+ L +FT W ISL LP
Sbjct: 493 AVPLYLSEMAPAKNRGAVNQLFQFTTCAGILIANLVNYFTEKIHPYGWRISLGLAGLP-A 669
Query: 176 FGIY-----------QVYRQSKEEEAKQILSKIYRPGEVEEEMKAMHESIEAEKAEDGLI 224
F + + Q + ++AKQ+L +I VE E + + E+ E +A
Sbjct: 670 FAMLVGGICCAETPNSLVEQGRLDKAKQVLQRIRGTENVEAEFEDLKEASEEAQAVKSPF 849
Query: 225 GHSLAQKLKGAWSNDVVRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFA 284
L +K + + + + + QQ+ G N+I++Y+P I Q G +N++ F+
Sbjct: 850 RTLLKRKYR--------PQLIIGALGIPAFQQLTGNNSILFYAPVIFQSLGFGANASLFS 1005
Query: 285 LSLVTSGLNAVGTIVSMVLIDRFGRRKLML 314
S +T+G V T++SM L+D++GRRK L
Sbjct: 1006-SFITNGALLVATVISMFLVDKYGRRKFFL 1092
>TC233398
Length = 442
Score = 125 bits (314), Expect = 4e-29
Identities = 63/98 (64%), Positives = 77/98 (78%), Gaps = 3/98 (3%)
Frame = +3
Query: 3 EGGHPLADKTEFTECWRRTAESPYLMRLALS---GGLLFGYDTGVISGASLYIRDEFEQV 59
EGG P AD + F EC + ++PY++RLA S GG LFGYDTGVISGA LYIRD+F++V
Sbjct: 147 EGGVPEADISAFRECLSLSWKNPYVLRLAFSAGIGGFLFGYDTGVISGALLYIRDDFKEV 326
Query: 60 DKKPWLQETIVSMASAGAIIGAAFGGYMNDKMGRKKTI 97
D+K WLQE IVSMA AGAIIGA+ GG++ND+ GRKK I
Sbjct: 327 DRKTWLQEAIVSMALAGAIIGASVGGWINDRFGRKKAI 440
>TC205503 UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, complete
Length = 1928
Score = 112 bits (281), Expect = 2e-25
Identities = 86/349 (24%), Positives = 160/349 (45%), Gaps = 42/349 (12%)
Frame = +1
Query: 30 LALSGGLLFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASAGAIIGAAFGGYMND 89
LA +L GYD GV+SGA++YI+ + + D++ E ++ + + ++IG+ G +D
Sbjct: 274 LASMTSILLGYDIGVMSGAAIYIKRDLKVSDEQI---EILLGIINLYSLIGSCLAGRTSD 444
Query: 90 KMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPLYISEASPAK 149
+GR+ TI + +F+ G+ +M P ++ GR + G+G+G A M AP+Y +E SPA
Sbjct: 445 WIGRRYTIGLGGAIFLVGSTLMGFYPHYSFLMCGRFVAGIGIGYALMIAPVYTAEVSPAS 624
Query: 150 IRGALVCRLHQWFTH-------NRWSISLLPHQFGIYQ---------------------- 180
RG L + + + S L + G
Sbjct: 625 SRGFLTSFPEVFINGGILLGYISNYGFSKLTLKVGWRMMLGVGAIPSVVLTEGVLAMPES 804
Query: 181 ---VYRQSKEEEAKQILSKIYRPGEVEEEMKAMHESIEA----EKAEDGLIGHSLAQKLK 233
+ + + EA+++L+K EE + E +A E D ++ + +
Sbjct: 805 PRWLVMRGRLGEARKVLNK--TSDSKEEAQLRLAEIKQAAGIPESCNDDVVQVNKQSNGE 978
Query: 234 GAWS------NDVVRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSL 287
G W +R + A + + QQ G++ ++ YSP I + AGI +++ ++
Sbjct: 979 GVWKELFLYPTPAIRHIVIAALGIHFFQQASGVDAVVLYSPRIFEKAGITNDTHKLLATV 1158
Query: 288 VTSGLNAVGTIVSMVLIDRFGRRKLMLISLIGIFVSLVTLSVTFNQAAH 336
+ V + + +DR GRR L+L S+ G+ +SL+TL+++ H
Sbjct: 1159AVGFVKTVFILAATFTLDRVGRRPLLLSSVGGMVLSLLTLAISLTVIDH 1305
>AW099412
Length = 373
Score = 110 bits (275), Expect = 1e-24
Identities = 61/112 (54%), Positives = 76/112 (67%), Gaps = 1/112 (0%)
Frame = +1
Query: 181 VYRQSKEEEAKQILSKIYRPGEVEEEMKAMHESIEAE-KAEDGLIGHSLAQKLKGAWSND 239
++R+ +EEE K IL KIY P EVE E+ + ES+E E K + S+ + LK
Sbjct: 49 LFRKGREEEGKAILRKIYPPQEVEAEINTLKESVEIEIKEAEASDKVSIVKMLK----TK 216
Query: 240 VVRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLVTSG 291
VRRGLYAG+ +Q+ QQ VGINT+MYYSPTIVQ AG ASN TA LSL+TSG
Sbjct: 217 TVRRGLYAGMGLQIFQQFVGINTVMYYSPTIVQLAGFASNRTALLLSLITSG 372
>AW760828
Length = 423
Score = 107 bits (268), Expect = 8e-24
Identities = 55/88 (62%), Positives = 67/88 (75%), Gaps = 3/88 (3%)
Frame = +1
Query: 3 EGGHPLADKTEFTECWRRTAESPYLMRLALS---GGLLFGYDTGVISGASLYIRDEFEQV 59
EGG D + F EC + ++PY++RLA S GGLLFGYDTGVISGA LYIRD+F+ V
Sbjct: 160 EGGGVEVDASAFRECLSLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGAILYIRDDFKAV 339
Query: 60 DKKPWLQETIVSMASAGAIIGAAFGGYM 87
D+K WLQE IVSMA AGAI+GAA GG++
Sbjct: 340 DRKTWLQEAIVSMALAGAIVGAAVGGWI 423
>TC218649 similar to UP|Q39416 (Q39416) Integral membrane protein, partial
(93%)
Length = 1695
Score = 104 bits (260), Expect = 6e-23
Identities = 87/309 (28%), Positives = 158/309 (50%), Gaps = 31/309 (10%)
Frame = +2
Query: 71 SMASAGAIIGAAFGGYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLG 130
S+++ GA++GA G + + +GRK ++++A + + G L ++ A + +GRLL G G
Sbjct: 305 SLSNVGAMVGAIASGQIAEYIGRKGSLMIASIPNIIGWLAISFAKDSSFLYMGRLLEGFG 484
Query: 131 VGAASMTAPLYISEASPAKIRGALVCRLHQWFTHNRWSIS---LLPHQFGIYQVYRQSK- 186
VG S T P+YI+E SP +RG LV + N+ S++ +L + GI+ +R
Sbjct: 485 VGIISYTVPVYIAEISPPNLRGGLV-------SVNQLSVTIGIMLAYLLGIFVEWRILAI 643
Query: 187 ---------------EEEAKQILSKIYRPGEVEEEMKAMHE-----SIEAEKAEDGLIGH 226
E+ + L+K+ E E ++ + S+E + + +
Sbjct: 644 IGILPCTILIPALFFIPESPRWLAKMGMTEEFETSLQVLRGFDTDISVEVNEIKRAVAST 823
Query: 227 SLAQKLKGAWSNDVVRR----GLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTA 282
+ ++ A D+ +R L GI + ++QQ+ GIN +++YS TI + AGI+S+ A
Sbjct: 824 NTRITVRFA---DLKQRRYWLPLMIGIGLLILQQLSGINGVLFYSSTIFRNAGISSSDAA 994
Query: 283 FALSLVTSGLNAV---GTIVSMVLIDRFGRRKLMLISLIGIFVSLVTLSVTFNQAAHHAP 339
T G+ AV T +++ L D+ GRR L+++S G+ SL+ +++TF A +
Sbjct: 995 ------TFGVGAVQVLATSLTLWLADKSGRRLLLIVSATGMSFSLLVVAITFYIKASISE 1156
Query: 340 SLSIQDSLS 348
+ S+ LS
Sbjct: 1157TSSLYGILS 1183
>BU577019
Length = 409
Score = 95.9 bits (237), Expect = 3e-20
Identities = 50/79 (63%), Positives = 59/79 (74%), Gaps = 3/79 (3%)
Frame = +3
Query: 3 EGGHPLADKTEFTECWRRTAESPYLMRLALS---GGLLFGYDTGVISGASLYIRDEFEQV 59
EGG D + F EC + ++PY++RLA S GGLLFGYDTGVISGA LYIRD+F++V
Sbjct: 171 EGGGVEVDVSAFRECLSLSWKNPYVLRLAFSAGIGGLLFGYDTGVISGALLYIRDDFKEV 350
Query: 60 DKKPWLQETIVSMASAGAI 78
D K WLQE IVSMA AGAI
Sbjct: 351 DSKTWLQEAIVSMALAGAI 407
>TC229858 weakly similar to UP|Q8VHD6 (Q8VHD6) Glucose transporter 10 (Mus
musculus 13 days embryo heart cDNA, RIKEN full-length
enriched library, clone:D330001H08 product:solute
carrier family 2 (facilitated glucose transporter),
member 10, full insert sequence), partial (4%)
Length = 715
Score = 92.4 bits (228), Expect = 3e-19
Identities = 47/91 (51%), Positives = 59/91 (64%)
Frame = +3
Query: 241 VRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLVTSGLNAVGTIVS 300
+R G + QQ GINT+MYYSPTIVQ AG +N A LSL+ +G+NA GTI+
Sbjct: 6 IRLAFLVGAGLLAFQQFTGINTVMYYSPTIVQMAGFHANELALLLSLIVAGMNAAGTILG 185
Query: 301 MVLIDRFGRRKLMLISLIGIFVSLVTLSVTF 331
+ LID GR+KL L SL G+ VSLV L+ F
Sbjct: 186 IYLIDHAGRKKLALSSLGGVIVSLVILAFAF 278
>AW620575
Length = 377
Score = 86.3 bits (212), Expect = 2e-17
Identities = 44/70 (62%), Positives = 55/70 (77%), Gaps = 3/70 (4%)
Frame = +2
Query: 23 ESPYLM---RLALSGGLLFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASAGAII 79
++PY+M +A GGLLFGYDTGVISGA LYI+D+F +V +LQETIVSMA GAI+
Sbjct: 167 QNPYIMGFTAVASIGGLLFGYDTGVISGALLYIKDDFPEVRHSNFLQETIVSMAVTGAIV 346
Query: 80 GAAFGGYMND 89
GAA GG++ND
Sbjct: 347 GAAAGGWIND 376
>TC216311 UP|Q7XA52 (Q7XA52) Monosaccharide transporter, complete
Length = 1925
Score = 83.6 bits (205), Expect = 2e-16
Identities = 79/340 (23%), Positives = 140/340 (40%), Gaps = 55/340 (16%)
Frame = +1
Query: 30 LALSGGLLFGYDTGVISGASL---YIRDEFEQVDKKPWLQETI--------------VSM 72
+A GGL+FGYD G+ G + ++ F V +K +T+ S
Sbjct: 181 VAAMGGLIFGYDIGISGGVTSMDPFLLKFFPSVFRKKNSDKTVNQYCQYDSQTLTMFTSS 360
Query: 73 ASAGAIIGAAFGGYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVG 132
A++ + + + GRK ++L ++F+ GAL+ A W++I+GR+L+G G+G
Sbjct: 361 LYLAALLSSLVASTVTRRFGRKLSMLFGGLLFLVGALINGFAQHVWMLIVGRILLGFGIG 540
Query: 133 AASM---TAPLYISEASPAKIRGALVCRLHQWFTHNR--------WSISLLPHQFGI--- 178
A+ T P+ + G L F R W S++ + I
Sbjct: 541 FANQVCATLPI*NGFIQI*RSIGTLAFNCQSQFGIPRGQCVELLLWLKSMVAWGWKIEVW 720
Query: 179 ---------------------YQVYRQSKEEEAKQILSKIYRPGEVEEEMKAMHESIEAE 217
+ + E+AK L +I V+EE + + E+
Sbjct: 721 EGAMVPALIITVGSLVLPDTPNSMIERGDREKAKAQLQRIRGIDNVDEEFNDLVAASES- 897
Query: 218 KAEDGLIGHSLAQKLKGAWSNDVVRR---GLYAGITVQVVQQIVGINTIMYYSPTIVQFA 274
+ +++ W N + R+ L + + QQ+ GIN IM+Y+P +
Sbjct: 898 -----------SSQVEHPWRNLLQRKYRPHLTMAVLIPFFQQLTGINVIMFYAPVLFSSI 1044
Query: 275 GIASNSTAFALSLVTSGLNAVGTIVSMVLIDRFGRRKLML 314
G + A +++T +N V T VS+ +D++GRR L L
Sbjct: 1045GF-KDDAALMSAVITGVVNVVATCVSIYGVDKWGRRALFL 1161
>TC233090 similar to UP|Q9ZVN7 (Q9ZVN7) T7A14.10 protein, partial (22%)
Length = 594
Score = 79.0 bits (193), Expect = 4e-15
Identities = 45/130 (34%), Positives = 76/130 (57%)
Frame = +3
Query: 25 PYLMRLALSGGLLFGYDTGVISGASLYIRDEFEQVDKKPWLQETIVSMASAGAIIGAAFG 84
P+++ ++S +FGY GV++G + I E + +++ +VS+ AGA IG+
Sbjct: 6 PHVLVASMSN-FIFGYHIGVMNGPIVSIARELG-FEGNSFIEGLVVSIFIAGAFIGSISS 179
Query: 85 GYMNDKMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPLYISE 144
+ D++G + T + + + GA++ A A + II GR LVGLG+G ++ P+YISE
Sbjct: 180 ASLLDRLGSRLTFQINSIPLILGAIISAQAHSLNEIIGGRFLVGLGIGVNTVLVPIYISE 359
Query: 145 ASPAKIRGAL 154
+P K RGAL
Sbjct: 360 VAPTKYRGAL 389
>TC218012 similar to UP|Q6VEF2 (Q6VEF2) Monosaccharide transporter 4, partial
(54%)
Length = 1142
Score = 72.4 bits (176), Expect = 4e-13
Identities = 46/159 (28%), Positives = 87/159 (53%), Gaps = 4/159 (2%)
Frame = +3
Query: 188 EEAKQILSKIYRPGEVEEEMKAMHESIEAEKAEDGLIGHSLAQKLKGAWSNDVVRRG--- 244
EE K +L KI +E E + E +EA + +A+++K + N + RR
Sbjct: 39 EEGKTVLKKIRGTDNIELEFQ---ELVEASR---------VAKEVKHPFRNLLKRRNRPQ 182
Query: 245 LYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLVTSGLNAVGTIVSMVLI 304
L I +Q+ QQ GIN IM+Y+P + G ++++ ++ +++T +N + T+VS+ +
Sbjct: 183 LVISIALQIFQQFTGINAIMFYAPVLFNTLGFKNDASLYS-AVITGAVNVLSTVVSIYSV 359
Query: 305 DRFGRRKLMLISLIGIFVSLVTLSVTFN-QAAHHAPSLS 342
D+ GRR L+L + + +F+S V +++ + H+ LS
Sbjct: 360 DKLGRRMLLLEAGVQMFLSQVVIAIILGIKVTDHSDDLS 476
>AW458960
Length = 285
Score = 69.7 bits (169), Expect = 2e-12
Identities = 35/81 (43%), Positives = 51/81 (62%), Gaps = 2/81 (2%)
Frame = +2
Query: 308 GRRKLMLISLIGIFVSLVTLSVTFNQAAHHAPSLSIQDSLSFGGNSTCKAYTTAPNHLSW 367
GR+KL+L SL G+ SLV L+V F+Q+ H+P +S ++ F N+TC Y +A N W
Sbjct: 5 GRKKLVLFSLCGVVFSLVVLTVVFHQSTTHSPMVSALETSHF--NNTCPDYHSAVNPGGW 178
Query: 368 NCMQCLHED--CAFCANSQNE 386
+CM+CL C FCA+ N+
Sbjct: 179 DCMKCLKASPGCGFCASGANK 241
>BE555798 homologue to GP|1353516|gb|A sugar transporter {Medicago
truncatula}, partial (18%)
Length = 312
Score = 67.0 bits (162), Expect = 2e-11
Identities = 30/65 (46%), Positives = 46/65 (70%)
Frame = +2
Query: 90 KMGRKKTILMADVVFVAGALVMAAAPAPWVIIIGRLLVGLGVGAASMTAPLYISEASPAK 149
K GRK ++L ++F+ GAL+ A W++I+GR+L+G G+G A+ + PLY+SE +P K
Sbjct: 8 KFGRKLSMLF*GLLFLVGALINGFAQHVWMLIVGRILLGFGIGFANQSVPLYLSEMAPYK 187
Query: 150 IRGAL 154
RGAL
Sbjct: 188 YRGAL 202
>TC218013 similar to UP|Q6VEF2 (Q6VEF2) Monosaccharide transporter 4, partial
(33%)
Length = 568
Score = 63.9 bits (154), Expect = 1e-10
Identities = 40/146 (27%), Positives = 81/146 (55%), Gaps = 3/146 (2%)
Frame = +2
Query: 187 EEEAKQILSKIYRPGEVEEEMKAMHESIEAEKAEDGLIGHSLAQKLKGAWSNDVVRRG-- 244
E + KQ L + +E E + E +EA + +A+++K + N + RR
Sbjct: 2 ERKEKQCLKRFGALYNIELEFQ---ELLEASR---------VAKEVKHPFRNLLKRRNRP 145
Query: 245 -LYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLVTSGLNAVGTIVSMVL 303
L + +Q+ QQ GIN IM+Y+P + G ++++ ++ +++T +N + T+VS+
Sbjct: 146 QLVISVALQIFQQFTGINAIMFYAPVLFNTLGFKNDASLYS-AVITGAVNVLSTVVSIYS 322
Query: 304 IDRFGRRKLMLISLIGIFVSLVTLSV 329
+D+ GRR L+L + + +F+S V +++
Sbjct: 323 VDKVGRRILLLEAGVQMFLSQVVIAI 400
>TC226707 similar to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, partial
(62%)
Length = 2338
Score = 63.9 bits (154), Expect = 1e-10
Identities = 48/171 (28%), Positives = 79/171 (46%), Gaps = 11/171 (6%)
Frame = +3
Query: 189 EAKQILSKIYRPGEVEEEMKAMHESIE-----AEKAEDGLIGHSLAQKLKGAWSN----- 238
EAK++L KI E EEE + I+ + +D ++ S G W
Sbjct: 168 EAKRVLYKI---SESEEEARLRLADIKDTAGIPQDCDDDVVLVSKQTHGHGVWRELFLHP 338
Query: 239 -DVVRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFAGIASNSTAFALSLVTSGLNAVGT 297
VR A + + Q GI+ ++ YSP I + AGI S++ ++ + V
Sbjct: 339 TPAVRHIFIASLGIHFFAQATGIDAVVLYSPRIFEKAGIKSDNYRLLATVAVGFVKTVSI 518
Query: 298 IVSMVLIDRFGRRKLMLISLIGIFVSLVTLSVTFNQAAHHAPSLSIQDSLS 348
+V+ +DR GRR L+L S+ G+ +SL+TL ++ H +L+ LS
Sbjct: 519 LVATFFLDRAGRRVLLLCSVSGLILSLLTLGLSLTVVDHSQTTLNWAVGLS 671
>TC203480 similar to UP|Q8LPQ8 (Q8LPQ8) AT4g35300/F23E12_140, partial (38%)
Length = 1040
Score = 62.8 bits (151), Expect = 3e-10
Identities = 41/113 (36%), Positives = 62/113 (54%), Gaps = 13/113 (11%)
Frame = +2
Query: 241 VRRGLYAGITVQVVQQIVGINTIMYYSPTIVQFA---------GIASNSTAFALSLVTSG 291
V+ L G+ +Q++QQ GIN ++YY+P I++ A GI S S +F +S T+
Sbjct: 197 VKHALVVGVGIQILQQFSGINGVLYYTPQILEEAGVEVLLSDIGIGSESASFLISAFTTF 376
Query: 292 LNAVGTIVSMVLIDRFGRRKLMLISLIGIFVSLVTL----SVTFNQAAHHAPS 340
L V+M L+D GRR+L+L ++ + VSL+ L V F AH A S
Sbjct: 377 LMLPCIGVAMKLMDVSGRRQLLLTTIPVLIVSLIILVIGSLVNFGNVAHAAIS 535
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.134 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,168,061
Number of Sequences: 63676
Number of extensions: 264603
Number of successful extensions: 1488
Number of sequences better than 10.0: 82
Number of HSP's better than 10.0 without gapping: 1455
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1470
length of query: 420
length of database: 12,639,632
effective HSP length: 100
effective length of query: 320
effective length of database: 6,272,032
effective search space: 2007050240
effective search space used: 2007050240
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC148526.6


