

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148483.6 + phase: 0
(125 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
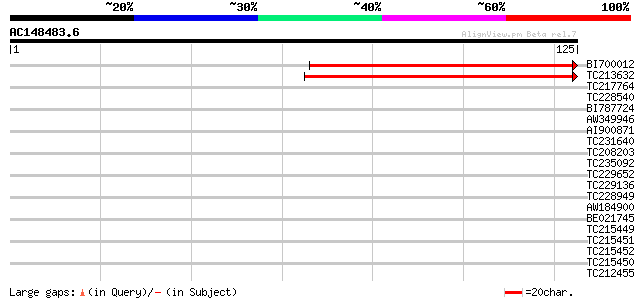
Score E
Sequences producing significant alignments: (bits) Value
BI700012 similar to GP|11320952|gb| peptide deformylase-like pro... 96 3e-21
TC213632 weakly similar to GB|AAG33973.1|11320952|AF250959 pepti... 85 7e-18
TC217764 similar to UP|DEFC_LYCES (Q9FV54) Peptide deformylase, ... 38 0.001
TC228540 29 0.48
BI787724 similar to PIR|T39903|T399 serine-rich protein - fissio... 29 0.48
AW349946 29 0.63
AI900871 similar to PIR|D96519|D96 mysoin-like protein 11013-73... 28 1.1
TC231640 homologue to UP|Q918P0 (Q918P0) Latency associated anti... 28 1.1
TC208203 weakly similar to UP|O04626 (O04626) F2P16.20 protein, ... 28 1.4
TC235092 similar to UP|O91332 (O91332) EBNA-1, partial (5%) 28 1.4
TC229652 28 1.4
TC229136 similar to UP|Q8VYW6 (Q8VYW6) AT4g39640/T19P19_30, part... 28 1.4
TC228949 similar to UP|Q945N1 (Q945N1) AT5g50310/MXI22_1, partia... 28 1.4
AW184900 27 1.8
BE021745 27 1.8
TC215449 similar to UP|P93488 (P93488) NAP1Ps, partial (97%) 27 1.8
TC215451 weakly similar to UP|Q94K07 (Q94K07) Nucleosome assembl... 27 1.8
TC215452 similar to UP|P93488 (P93488) NAP1Ps, partial (29%) 27 1.8
TC215450 similar to UP|P93488 (P93488) NAP1Ps, partial (30%) 27 1.8
TC212455 weakly similar to UP|Q7QIA8 (Q7QIA8) AgCP3984 (Fragment... 27 1.8
>BI700012 similar to GP|11320952|gb| peptide deformylase-like protein
{Arabidopsis thaliana}, partial (34%)
Length = 420
Score = 96.3 bits (238), Expect = 3e-21
Identities = 45/59 (76%), Positives = 54/59 (91%)
Frame = +1
Query: 67 LPYIVKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGIPLRV 125
LP VKAGDPV+HEPA++VD +EIKS+++Q IIDDMI VMRKAPGVG+AAPQIGIPLR+
Sbjct: 199 LPDTVKAGDPVLHEPAQDVDPNEIKSERVQKIIDDMIQVMRKAPGVGLAAPQIGIPLRI 375
>TC213632 weakly similar to GB|AAG33973.1|11320952|AF250959 peptide
deformylase-like protein {Arabidopsis thaliana;} ,
partial (38%)
Length = 464
Score = 85.1 bits (209), Expect = 7e-18
Identities = 41/60 (68%), Positives = 50/60 (83%)
Frame = +1
Query: 66 KLPYIVKAGDPVIHEPAREVDHSEIKSDKIQNIIDDMILVMRKAPGVGVAAPQIGIPLRV 125
KL IVKAG+ V+H A EV+ EIKS+++Q IIDDM+ VMRKAPGVG+AAPQIGIPLR+
Sbjct: 103 KLAKIVKAGEAVLHSRAEEVEAIEIKSERVQKIIDDMVRVMRKAPGVGLAAPQIGIPLRI 282
>TC217764 similar to UP|DEFC_LYCES (Q9FV54) Peptide deformylase, chloroplast
precursor (PDF) (Polypeptide deformylase) , partial
(75%)
Length = 1880
Score = 37.7 bits (86), Expect = 0.001
Identities = 29/120 (24%), Positives = 53/120 (44%), Gaps = 11/120 (9%)
Frame = +1
Query: 17 PVLSNGVVSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETALLRKTVNKLPYIVKAGDP 76
P L G V L+ S+ C N L + S + P + T L + GD
Sbjct: 82 PHLHRGSVLLAISNRCFNHSSLLNWILSVNPPRTAPPRAMAKPSSSTAQDL--VASPGDF 255
Query: 77 VIHEPAREVDHSEIK-----------SDKIQNIIDDMILVMRKAPGVGVAAPQIGIPLRV 125
+P + V++ + + D ++ ++ +M VM K G+G++APQ+GI +++
Sbjct: 256 EFAQPLKIVEYPDPRLRARNKRIVAFDDSLKKLVHEMFDVMYKTDGIGLSAPQLGINVQL 435
>TC228540
Length = 1112
Score = 29.3 bits (64), Expect = 0.48
Identities = 17/56 (30%), Positives = 28/56 (49%), Gaps = 2/56 (3%)
Frame = +1
Query: 9 WRLRAFPMPVLSNGVVSLSSSSSCNNKIQLSSTKF--SKFGSTLSSPSSETALLRK 62
WR+ P+P+ + + SSSSS + L ST K GS + ++ +LR+
Sbjct: 223 WRVTLDPIPMFKSTLPPTSSSSSTTPSMTLESTPLMPEKLGSISAHKLGDSLILRR 390
>BI787724 similar to PIR|T39903|T399 serine-rich protein - fission yeast
(Schizosaccharomyces pombe), partial (5%)
Length = 283
Score = 29.3 bits (64), Expect = 0.48
Identities = 15/35 (42%), Positives = 21/35 (59%)
Frame = -3
Query: 24 VSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETA 58
+ LSSS+SC++ LSS + LSS SS T+
Sbjct: 278 IQLSSSTSCSSSSMLSSIRM*SVSDPLSSSSSPTS 174
>AW349946
Length = 567
Score = 28.9 bits (63), Expect = 0.63
Identities = 19/33 (57%), Positives = 21/33 (63%)
Frame = +2
Query: 25 SLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSET 57
S SSSSS N SS+ S F S+LSSPSS T
Sbjct: 389 SNSSSSSSN-----SSSSSSSFSSSLSSPSSAT 472
>AI900871 similar to PIR|D96519|D96 mysoin-like protein 11013-7318
[imported] - Arabidopsis thaliana, partial (6%)
Length = 387
Score = 28.1 bits (61), Expect = 1.1
Identities = 11/27 (40%), Positives = 17/27 (62%)
Frame = +2
Query: 44 SKFGSTLSSPSSETALLRKTVNKLPYI 70
SKFGST+S P+ + + + VN Y+
Sbjct: 116 SKFGSTISEPAERISKIHRPVNSRTYL 196
>TC231640 homologue to UP|Q918P0 (Q918P0) Latency associated antigen, partial
(7%)
Length = 466
Score = 28.1 bits (61), Expect = 1.1
Identities = 18/46 (39%), Positives = 23/46 (49%)
Frame = -3
Query: 12 RAFPMPVLSNGVVSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSET 57
R FP S+ S SSSSS ++ SS+ S F S+ S S T
Sbjct: 143 RTFPSSSSSSSFSSSSSSSSSSSSSSSSSSSSSSFSSSSSWVSDST 6
>TC208203 weakly similar to UP|O04626 (O04626) F2P16.20 protein, partial
(14%)
Length = 733
Score = 27.7 bits (60), Expect = 1.4
Identities = 24/95 (25%), Positives = 39/95 (40%), Gaps = 11/95 (11%)
Frame = -2
Query: 13 AFPMPVLSNGVVSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETALLRKTV-------- 64
A P ++ G+ S S+++S K PS++T L ++
Sbjct: 414 AIPCSMIDTGIGSRSTATSAGKARARQLAKVCFISD--ERPSAKTTLQGYSLPLTDRYSS 241
Query: 65 ---NKLPYIVKAGDPVIHEPAREVDHSEIKSDKIQ 96
+ LPYI D V+HE + V H K DK++
Sbjct: 240 *KLSSLPYIYAKEDDVVHE--KRVFHIVAKGDKVK 142
>TC235092 similar to UP|O91332 (O91332) EBNA-1, partial (5%)
Length = 641
Score = 27.7 bits (60), Expect = 1.4
Identities = 19/45 (42%), Positives = 26/45 (57%)
Frame = +2
Query: 15 PMPVLSNGVVSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETAL 59
P+PV S+ SL+S++S SS+ SK + SSPSS AL
Sbjct: 71 PLPVTSSTTSSLTSAASPPPAPPPSSSPRSKDPTRNSSPSSSRAL 205
>TC229652
Length = 1485
Score = 27.7 bits (60), Expect = 1.4
Identities = 25/82 (30%), Positives = 40/82 (48%), Gaps = 3/82 (3%)
Frame = -3
Query: 7 LVWRLRAFPMPVLSNGVVSLSSSSSCNNKIQLSSTKFSKFGSTLSS-PSSETALLRKTVN 65
LV R FP+ ++S+ V+ SS S NK+ L + F + + PS T LLRK +
Sbjct: 883 LVNRSAPFPLKLVSSYVM*NSSKHSFTNKLCLGKDITAPFKALPAP*PSIPTELLRKEIG 704
Query: 66 KLPYIVKAG--DPVIHEPAREV 85
I ++ P ++ AR +
Sbjct: 703 FNTMISESR*IPPYLYRAARRI 638
>TC229136 similar to UP|Q8VYW6 (Q8VYW6) AT4g39640/T19P19_30, partial (62%)
Length = 1627
Score = 27.7 bits (60), Expect = 1.4
Identities = 16/43 (37%), Positives = 22/43 (50%)
Frame = +2
Query: 17 PVLSNGVVSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETAL 59
P+ +G LS N I ++ST S FGS + SPS+ L
Sbjct: 512 PINDHGTSHLSIIDPERNAISMTSTVNSYFGSKILSPSTGIVL 640
>TC228949 similar to UP|Q945N1 (Q945N1) AT5g50310/MXI22_1, partial (27%)
Length = 1134
Score = 27.7 bits (60), Expect = 1.4
Identities = 15/34 (44%), Positives = 22/34 (64%)
Frame = -3
Query: 25 SLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETA 58
S SSSSS ++ ++LS + S S+ SS SSE +
Sbjct: 259 SSSSSSSSSSSVRLSPSDSSSSSSSFSSSSSEAS 158
>AW184900
Length = 422
Score = 27.3 bits (59), Expect = 1.8
Identities = 15/42 (35%), Positives = 25/42 (58%)
Frame = +1
Query: 9 WRLRAFPMPVLSNGVVSLSSSSSCNNKIQLSSTKFSKFGSTL 50
++L A P VLSN ++SL+S+++ N + + GSTL
Sbjct: 160 FQLDATPATVLSNSLMSLASNNNTQNNNNFWALQDRSQGSTL 285
>BE021745
Length = 419
Score = 27.3 bits (59), Expect = 1.8
Identities = 18/64 (28%), Positives = 32/64 (49%)
Frame = +3
Query: 7 LVWRLRAFPMPVLSNGVVSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETALLRKTVNK 66
L+W + M + + SSS+ N +++S+ S+ + L+ PSS A + N
Sbjct: 186 LIWNCESGAM---LSSLTQSPSSSAPNCSSRMASSSRSRTKAWLTEPSSSLACICLPPNH 356
Query: 67 LPYI 70
LP+I
Sbjct: 357 LPFI 368
>TC215449 similar to UP|P93488 (P93488) NAP1Ps, partial (97%)
Length = 1495
Score = 27.3 bits (59), Expect = 1.8
Identities = 21/60 (35%), Positives = 33/60 (55%)
Frame = -2
Query: 23 VVSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETALLRKTVNKLPYIVKAGDPVIHEPA 82
+V LSSSSS ++ SS+ S S++SS SS ++ ++ N P + PV H+ A
Sbjct: 1149 LVLLSSSSSSSSSSSSSSSSSSSSSSSISSSSSSSS---RSPNSSP*M---ASPVNHDTA 988
>TC215451 weakly similar to UP|Q94K07 (Q94K07) Nucleosome assembly protein,
partial (20%)
Length = 777
Score = 27.3 bits (59), Expect = 1.8
Identities = 21/60 (35%), Positives = 33/60 (55%)
Frame = -1
Query: 23 VVSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETALLRKTVNKLPYIVKAGDPVIHEPA 82
+V LSSSSS ++ SS+ S S++SS SS ++ ++ N P + PV H+ A
Sbjct: 195 LVLLSSSSSSSSSSSSSSSSSSSSSSSISSSSSSSS---RSPNSSP*M---ASPVNHDTA 34
>TC215452 similar to UP|P93488 (P93488) NAP1Ps, partial (29%)
Length = 609
Score = 27.3 bits (59), Expect = 1.8
Identities = 21/60 (35%), Positives = 33/60 (55%)
Frame = -3
Query: 23 VVSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETALLRKTVNKLPYIVKAGDPVIHEPA 82
+V LSSSSS ++ SS+ S S++SS SS ++ ++ N P + PV H+ A
Sbjct: 244 LVLLSSSSSSSSSSSSSSSSSSSSSSSISSSSSSSS---RSPNSSP*M---ASPVNHDTA 83
>TC215450 similar to UP|P93488 (P93488) NAP1Ps, partial (30%)
Length = 672
Score = 27.3 bits (59), Expect = 1.8
Identities = 21/60 (35%), Positives = 33/60 (55%)
Frame = -3
Query: 23 VVSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETALLRKTVNKLPYIVKAGDPVIHEPA 82
+V LSSSSS ++ SS+ S S++SS SS ++ ++ N P + PV H+ A
Sbjct: 316 LVLLSSSSSSSSSSSSSSSSSSSSSSSISSSSSSSS---RSPNSSP*M---ASPVNHDTA 155
>TC212455 weakly similar to UP|Q7QIA8 (Q7QIA8) AgCP3984 (Fragment), partial
(11%)
Length = 944
Score = 27.3 bits (59), Expect = 1.8
Identities = 16/41 (39%), Positives = 24/41 (58%)
Frame = -1
Query: 18 VLSNGVVSLSSSSSCNNKIQLSSTKFSKFGSTLSSPSSETA 58
+L + ++ L SSSSC+ +SS+K S S L S S T+
Sbjct: 914 LLLSSLILLRSSSSCSGNCNVSSSKSSGSTSLLVSLLSSTS 792
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.316 0.132 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,435,508
Number of Sequences: 63676
Number of extensions: 71016
Number of successful extensions: 581
Number of sequences better than 10.0: 72
Number of HSP's better than 10.0 without gapping: 560
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 581
length of query: 125
length of database: 12,639,632
effective HSP length: 86
effective length of query: 39
effective length of database: 7,163,496
effective search space: 279376344
effective search space used: 279376344
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 53 (25.0 bits)
Medicago: description of AC148483.6


