

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148483.13 + phase: 0
(63 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
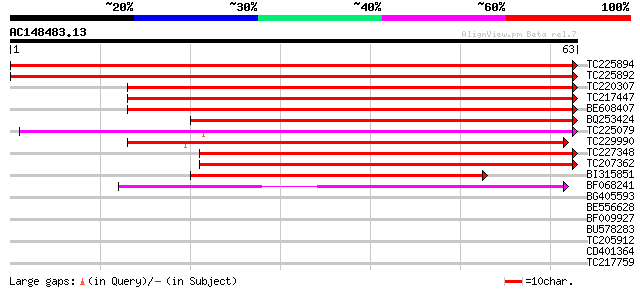
Score E
Sequences producing significant alignments: (bits) Value
TC225894 similar to UP|Q9FIQ0 (Q9FIQ0) Zinc finger protein Glo3-... 127 9e-31
TC225892 similar to UP|ARG3_HUMAN (Q9NP61) ADP-ribosylation fact... 127 9e-31
TC220307 similar to GB|AAM65362.1|21655301|AY103310 At2g37550/F1... 77 1e-15
TC217447 similar to GB|AAM65362.1|21655301|AY103310 At2g37550/F1... 77 2e-15
BE608407 similar to GP|4519792|dbj Asp1 {Arabidopsis thaliana}, ... 72 7e-14
BQ253424 71 1e-13
TC225079 similar to UP|Q8L8M0 (Q8L8M0) ARF GAP-like zinc finger-... 70 2e-13
TC229990 similar to GB|AAP68261.1|31711810|BT008822 At4g21160 {A... 65 9e-12
TC227348 similar to UP|Q9SMX5 (Q9SMX5) GCN4-complementing protei... 61 1e-10
TC207362 similar to UP|Q9FIT8 (Q9FIT8) GCN4-complementing protei... 61 1e-10
BI315851 51 1e-07
BF068241 39 5e-04
BG405593 homologue to GP|10441354|gb| ARF GAP-like zinc finger-c... 37 0.002
BE556628 36 0.003
BF009927 28 0.94
BU578283 weakly similar to GP|16323057|gb| AT5g57660/MRI1_1 {Ara... 27 1.6
TC205912 weakly similar to UP|O24040 (O24040) LTCOR11, partial (... 25 7.9
CD401364 weakly similar to GP|13774348|gb histidine-containing p... 25 7.9
TC217759 similar to UP|Q6ZG80 (Q6ZG80) Zinc finger (C3HC4-type R... 25 7.9
>TC225894 similar to UP|Q9FIQ0 (Q9FIQ0) Zinc finger protein Glo3-like,
partial (32%)
Length = 612
Score = 127 bits (320), Expect = 9e-31
Identities = 59/63 (93%), Positives = 59/63 (93%)
Frame = +1
Query: 1 MASDSFTDKNAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS 60
MASD FTDKN VFRKLK KSENK CFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS
Sbjct: 208 MASDGFTDKNTVFRKLKAKSENKMCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS 387
Query: 61 FVR 63
FVR
Sbjct: 388 FVR 396
>TC225892 similar to UP|ARG3_HUMAN (Q9NP61) ADP-ribosylation factor
GTPase-activating protein 3 (ARF GAP 3), partial (12%)
Length = 1727
Score = 127 bits (320), Expect = 9e-31
Identities = 59/63 (93%), Positives = 59/63 (93%)
Frame = +2
Query: 1 MASDSFTDKNAVFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS 60
MASD FTDKN VFRKLK KSENK CFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS
Sbjct: 209 MASDGFTDKNTVFRKLKAKSENKMCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHIS 388
Query: 61 FVR 63
FVR
Sbjct: 389 FVR 397
>TC220307 similar to GB|AAM65362.1|21655301|AY103310 At2g37550/F13M22.5
{Arabidopsis thaliana;} , partial (28%)
Length = 602
Score = 77.4 bits (189), Expect = 1e-15
Identities = 31/50 (62%), Positives = 41/50 (82%)
Frame = +2
Query: 14 RKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVR 63
R L++++ NK C DC+ KNP WASV+YG+F+C++CS HR LGVHISFVR
Sbjct: 170 RDLQSEAGNKICVDCSQKNPQWASVSYGVFMCLECSGKHRGLGVHISFVR 319
>TC217447 similar to GB|AAM65362.1|21655301|AY103310 At2g37550/F13M22.5
{Arabidopsis thaliana;} , partial (35%)
Length = 563
Score = 77.0 bits (188), Expect = 2e-15
Identities = 31/50 (62%), Positives = 40/50 (80%)
Frame = +3
Query: 14 RKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVR 63
R L+++ NK C DC+ KNP WASV+YG+F+C++CS HR LGVHISFVR
Sbjct: 60 RDLQSQPANKICVDCSQKNPQWASVSYGVFMCLECSGKHRGLGVHISFVR 209
>BE608407 similar to GP|4519792|dbj Asp1 {Arabidopsis thaliana}, partial
(26%)
Length = 421
Score = 71.6 bits (174), Expect = 7e-14
Identities = 29/50 (58%), Positives = 38/50 (76%)
Frame = +2
Query: 14 RKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVR 63
R L+++ NK C DC+ KNP WASV+YG+F+C++CS HR L VHI FVR
Sbjct: 20 RDLQSQPANKICVDCSQKNPQWASVSYGVFMCLECSGKHRGLCVHICFVR 169
>BQ253424
Length = 277
Score = 70.9 bits (172), Expect = 1e-13
Identities = 30/43 (69%), Positives = 33/43 (75%)
Frame = +2
Query: 21 ENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVR 63
ENK C DC AK P WASV GIF+C+ CS +HRSLGVHIS VR
Sbjct: 128 ENKECADCKAKGPRWASVNLGIFICMQCSGIHRSLGVHISKVR 256
>TC225079 similar to UP|Q8L8M0 (Q8L8M0) ARF GAP-like zinc finger-containing
protein ZIGA3, partial (48%)
Length = 1845
Score = 70.1 bits (170), Expect = 2e-13
Identities = 35/73 (47%), Positives = 42/73 (56%), Gaps = 11/73 (15%)
Frame = +3
Query: 2 ASDSFTDKNAVFRKLKTKS-----------ENKSCFDCNAKNPTWASVTYGIFLCIDCSA 50
AS + K V ++L K EN+ C DC AK P WASV GIF+C+ CS
Sbjct: 123 ASSTMNSKANVSKELNAKHKKILEGLLKLPENRECADCKAKGPRWASVNLGIFICMQCSG 302
Query: 51 VHRSLGVHISFVR 63
+HRSLGVHIS VR
Sbjct: 303 IHRSLGVHISKVR 341
>TC229990 similar to GB|AAP68261.1|31711810|BT008822 At4g21160 {Arabidopsis
thaliana;} , partial (22%)
Length = 444
Score = 64.7 bits (156), Expect = 9e-12
Identities = 29/52 (55%), Positives = 36/52 (68%), Gaps = 3/52 (5%)
Frame = +3
Query: 14 RKLKT---KSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFV 62
RKLK +S+N+ C DCNA +P WAS G+F+C+ C VHRSLG HIS V
Sbjct: 207 RKLKDLLLQSDNRLCADCNAPDPKWASANIGVFICLKCCGVHRSLGTHISKV 362
>TC227348 similar to UP|Q9SMX5 (Q9SMX5) GCN4-complementing protein (GCP1),
partial (29%)
Length = 1197
Score = 61.2 bits (147), Expect = 1e-10
Identities = 25/42 (59%), Positives = 31/42 (73%)
Frame = +3
Query: 22 NKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVR 63
N C +C+A P WAS+ GI LCI+CS VHR+LGVH+S VR
Sbjct: 594 NDKCAECSAPEPDWASLNLGILLCIECSGVHRNLGVHVSKVR 719
>TC207362 similar to UP|Q9FIT8 (Q9FIT8) GCN4-complementing protein homolog,
partial (51%)
Length = 1959
Score = 61.2 bits (147), Expect = 1e-10
Identities = 25/42 (59%), Positives = 30/42 (70%)
Frame = +1
Query: 22 NKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFVR 63
N C DC A P WAS+ G+ +CI+CS VHR+LGVHIS VR
Sbjct: 793 NDKCADCGAPEPDWASLNLGVLVCIECSGVHRNLGVHISKVR 918
>BI315851
Length = 385
Score = 51.2 bits (121), Expect = 1e-07
Identities = 19/33 (57%), Positives = 23/33 (69%)
Frame = +1
Query: 21 ENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHR 53
EN+ C DC K P WASV GIF+C+ CS +HR
Sbjct: 286 ENRECADCRNKAPRWASVNLGIFICMQCSGIHR 384
>BF068241
Length = 387
Score = 38.9 bits (89), Expect = 5e-04
Identities = 20/50 (40%), Positives = 26/50 (52%)
Frame = +2
Query: 13 FRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISFV 62
F+ +K N +C C S G+F+C+ C VHRSLG HIS V
Sbjct: 134 FKLVKE*LPNGTCKTC------LKSANIGVFICLKCCGVHRSLGTHISKV 265
>BG405593 homologue to GP|10441354|gb| ARF GAP-like zinc finger-containing
protein ZiGA4 {Arabidopsis thaliana}, partial (8%)
Length = 385
Score = 37.0 bits (84), Expect = 0.002
Identities = 12/41 (29%), Positives = 22/41 (53%)
Frame = +1
Query: 12 VFRKLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDCSAVH 52
+ R L N+ C +CN+ P + ++ F+C+ CS +H
Sbjct: 262 IIRGLMKLPPNRRCINCNSLGPQYVCTSFWTFICMTCSGIH 384
>BE556628
Length = 112
Score = 36.2 bits (82), Expect = 0.003
Identities = 13/26 (50%), Positives = 18/26 (69%)
Frame = +2
Query: 22 NKSCFDCNAKNPTWASVTYGIFLCID 47
N C +C+A +P W+S+ GI LCID
Sbjct: 35 NDKCAECSAPDPYWSSLNLGILLCID 112
>BF009927
Length = 228
Score = 28.1 bits (61), Expect = 0.94
Identities = 11/17 (64%), Positives = 14/17 (81%)
Frame = +1
Query: 1 MASDSFTDKNAVFRKLK 17
M S+ FT+KN +FRKLK
Sbjct: 178 MTSNGFTNKNTIFRKLK 228
>BU578283 weakly similar to GP|16323057|gb| AT5g57660/MRI1_1 {Arabidopsis
thaliana}, partial (11%)
Length = 409
Score = 27.3 bits (59), Expect = 1.6
Identities = 15/39 (38%), Positives = 18/39 (45%), Gaps = 1/39 (2%)
Frame = +1
Query: 15 KLKTKSENKSCFDCNAKNPTWASVTYGIFLCIDC-SAVH 52
+LKTK K C C A P +LC+ C S VH
Sbjct: 202 ELKTKEMEKVCEFCTALRPLVYCKADAAYLCLSCDSKVH 318
>TC205912 weakly similar to UP|O24040 (O24040) LTCOR11, partial (61%)
Length = 957
Score = 25.0 bits (53), Expect = 7.9
Identities = 13/31 (41%), Positives = 17/31 (53%)
Frame = -3
Query: 17 KTKSENKSCFDCNAKNPTWASVTYGIFLCID 47
K SEN SC N K P W ++ + I LC +
Sbjct: 280 KRNSENISC---NMKLPKWPNLWHFIGLCFE 197
>CD401364 weakly similar to GP|13774348|gb histidine-containing
phosphotransfer protein {Catharanthus roseus}, partial
(74%)
Length = 608
Score = 25.0 bits (53), Expect = 7.9
Identities = 15/42 (35%), Positives = 23/42 (54%)
Frame = -2
Query: 20 SENKSCFDCNAKNPTWASVTYGIFLCIDCSAVHRSLGVHISF 61
++NKSC C NPT S+ Y LC+ V + + + +SF
Sbjct: 208 TQNKSCPICKQNNPT--SILY-FSLCV*FCPVDQVVKLDLSF 92
>TC217759 similar to UP|Q6ZG80 (Q6ZG80) Zinc finger (C3HC4-type RING
finger) protein-like, partial (72%)
Length = 1362
Score = 25.0 bits (53), Expect = 7.9
Identities = 13/36 (36%), Positives = 21/36 (58%), Gaps = 2/36 (5%)
Frame = +1
Query: 18 TKSENKSCFDCNAKNPTWASVTY--GIFLCIDCSAV 51
T+ ++KS FDC+ ++ +TY G FL + S V
Sbjct: 1 TRMKSKSGFDCSCRSLYLVLITYY*GFFLLLKWSLV 108
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.324 0.132 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,423,133
Number of Sequences: 63676
Number of extensions: 43266
Number of successful extensions: 325
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 325
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 325
length of query: 63
length of database: 12,639,632
effective HSP length: 39
effective length of query: 24
effective length of database: 10,156,268
effective search space: 243750432
effective search space used: 243750432
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 52 (24.6 bits)
Medicago: description of AC148483.13


