

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148482.2 + phase: 0
(530 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
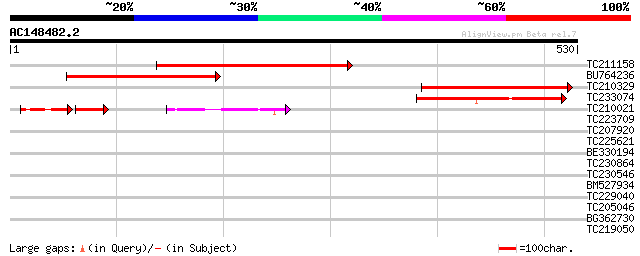
Score E
Sequences producing significant alignments: (bits) Value
TC211158 weakly similar to UP|Q8LG88 (Q8LG88) Sodium-dicarboxyla... 321 6e-88
BU764236 265 3e-71
TC210329 weakly similar to UP|Q8LG88 (Q8LG88) Sodium-dicarboxyla... 239 2e-63
TC233074 similar to UP|Q8LG88 (Q8LG88) Sodium-dicarboxylate cotr... 209 3e-54
TC210021 59 8e-15
TC223709 37 0.017
TC207920 similar to UP|Q6YBV9 (Q6YBV9) Cryptochrome 1, partial (... 30 2.0
TC225621 similar to UP|Q6NQE2 (Q6NQE2) At4g27270, partial (86%) 30 2.0
BE330194 similar to GP|10177043|dbj alcohol dehydrogenase-like p... 29 4.5
TC230864 similar to UP|Q7XJ60 (Q7XJ60) At3g47690, partial (26%) 29 4.5
TC230546 similar to GB|AAM62506.1|21553413|AY085274 VAMP (vesicl... 29 4.5
BM527934 29 5.9
TC229040 weakly similar to GB|AAP68276.1|31711840|BT008837 At3g6... 29 5.9
TC205046 similar to UP|P93166 (P93166) SCOF-1, partial (43%) 29 5.9
BG362730 28 7.7
TC219050 similar to UP|AHM2_ARATH (O64474) Potential cadmium/zin... 28 7.7
>TC211158 weakly similar to UP|Q8LG88 (Q8LG88) Sodium-dicarboxylate
cotransporter-like, partial (32%)
Length = 656
Score = 321 bits (822), Expect = 6e-88
Identities = 154/183 (84%), Positives = 170/183 (92%)
Frame = +2
Query: 138 LNVTLVFCGDPLNPSMLLLGLCATTFFVSMWLHNVATAVMMMPVATGILHRLPPAEEQPE 197
L VTL+FCGDP+NP++LLLGLCAT FFVSMWLHNVATAVMMMPVATGI+ RLP A EQ E
Sbjct: 107 LQVTLLFCGDPVNPALLLLGLCATIFFVSMWLHNVATAVMMMPVATGIVQRLPQAHEQSE 286
Query: 198 LMNKFSRAVILTVVYATPIGGISTLTGTGVNLIIIGMWKSLAPEAKPISFNSWFFFGFPV 257
+NKF RAVILTVVYATPIGGISTLTGTGVNLIIIGMWKSL PEAKPISFN+WFF+GFPV
Sbjct: 287 AVNKFCRAVILTVVYATPIGGISTLTGTGVNLIIIGMWKSLFPEAKPISFNTWFFYGFPV 466
Query: 258 AVLLLLCFWCILCLIYVRKGSSRALSDYLDRALLKRDLEALGPMSFAEKMVLCVFGLLIV 317
AVL+L+C+WCI+CL+YV KGS+RALS YLDRA LKRDLEALGPM+FAEKMVL VFGLLI+
Sbjct: 467 AVLILICYWCIICLLYVPKGSARALSAYLDRAHLKRDLEALGPMAFAEKMVLSVFGLLII 646
Query: 318 LWM 320
LWM
Sbjct: 647 LWM 655
>BU764236
Length = 445
Score = 265 bits (678), Expect = 3e-71
Identities = 127/144 (88%), Positives = 135/144 (93%)
Frame = +3
Query: 54 VKLDAPTTSTKMLGVIAWVFTWWITNAVPLPVTSMCPLFLFPLFGIASADTVAHSYMDDV 113
VKLDAP TS KMLGVIAWVF WW+T AVPLPVTSMCPLFLFP+FGIASAD+VAHSYMDDV
Sbjct: 6 VKLDAPPTSQKMLGVIAWVFAWWVTQAVPLPVTSMCPLFLFPIFGIASADSVAHSYMDDV 185
Query: 114 ITLVLGSFILALAVERYNVHRRLALNVTLVFCGDPLNPSMLLLGLCATTFFVSMWLHNVA 173
ITLVLGSFILALAVERYNVHRRLALNVTL+FCGDP+NP++LLLGLCAT FFVSMWLHNVA
Sbjct: 186 ITLVLGSFILALAVERYNVHRRLALNVTLLFCGDPVNPALLLLGLCATIFFVSMWLHNVA 365
Query: 174 TAVMMMPVATGILHRLPPAEEQPE 197
TAVMMMPVATGI+ RLPP EQ E
Sbjct: 366 TAVMMMPVATGIVQRLPPVHEQSE 437
>TC210329 weakly similar to UP|Q8LG88 (Q8LG88) Sodium-dicarboxylate
cotransporter-like, partial (25%)
Length = 744
Score = 239 bits (611), Expect = 2e-63
Identities = 118/141 (83%), Positives = 131/141 (92%)
Frame = +2
Query: 386 FALADGVQSSGLADVMSKALEFLKDVPYLAIVPAVSLLCSIITEFITSNDATATLLVPLL 445
FA+ADGVQSSGLADV+S+AL+FL+D PYLAIVPAVSL+CSIITEFITSNDATATLLVPLL
Sbjct: 2 FAIADGVQSSGLADVLSRALDFLEDAPYLAIVPAVSLICSIITEFITSNDATATLLVPLL 181
Query: 446 YHIAINMHVHPLLLMIPGGIATEFAFWLPTSTPSNVVGFATGHIEIKDMLKVGVPLKVAG 505
YHIA MHVHPLLLM+PGGIATEFAFWLPTSTPSNVVGFATGHIEIKDMLKVGVPLKV G
Sbjct: 182 YHIARTMHVHPLLLMVPGGIATEFAFWLPTSTPSNVVGFATGHIEIKDMLKVGVPLKVVG 361
Query: 506 VSVLALLMPTLGKLLLSLFSD 526
+ VL+LLMP+LG ++ +D
Sbjct: 362 IVVLSLLMPSLGTIVFGTNND 424
>TC233074 similar to UP|Q8LG88 (Q8LG88) Sodium-dicarboxylate
cotransporter-like, partial (24%)
Length = 445
Score = 209 bits (531), Expect = 3e-54
Identities = 110/142 (77%), Positives = 125/142 (87%), Gaps = 2/142 (1%)
Frame = +1
Query: 381 LLGAGFALADGVQSSGLADVMSKALEFLKDVPYLAIVPAVSLLCSIITEFITSND--ATA 438
LLGAGFALADGVQSSGLADV+S+AL+FL+D PY AI PAVSL+ SIITEFITSND ATA
Sbjct: 13 LLGAGFALADGVQSSGLADVLSRALDFLEDTPYWAIAPAVSLMSSIITEFITSNDATATA 192
Query: 439 TLLVPLLYHIAINMHVHPLLLMIPGGIATEFAFWLPTSTPSNVVGFATGHIEIKDMLKVG 498
TLL+PLLY+IA MHVHPLLLM PG +AT A LPTSTPSNVVGFATGHIEI DMLK+G
Sbjct: 193 TLLIPLLYNIAKTMHVHPLLLMEPGTVAT--AXLLPTSTPSNVVGFATGHIEILDMLKIG 366
Query: 499 VPLKVAGVSVLALLMPTLGKLL 520
VPLKVAG++VL++ MPTLG ++
Sbjct: 367 VPLKVAGIAVLSIFMPTLGAIV 432
>TC210021
Length = 429
Score = 58.9 bits (141), Expect(2) = 8e-15
Identities = 24/31 (77%), Positives = 27/31 (86%)
Frame = +2
Query: 62 STKMLGVIAWVFTWWITNAVPLPVTSMCPLF 92
ST ML VIAWV++WW+T AVPLPVTSMCP F
Sbjct: 110 STSMLAVIAWVYSWWVTAAVPLPVTSMCPKF 202
Score = 54.3 bits (129), Expect = 1e-07
Identities = 43/131 (32%), Positives = 64/131 (48%), Gaps = 15/131 (11%)
Frame = +2
Query: 147 DPLNPSMLLLGLCATTFFVSMWLHNV-ATAVMMMPVATGILHRLPPAEEQPELMNKFSRA 205
DP PS + L + V W+++ TA + +PV + + KF+RA
Sbjct: 77 DPSCPS*YV-SLSTSMLAVIAWVYSWWVTAAVPLPVTS--------------MCPKFNRA 211
Query: 206 VILTVVYATPIGGISTLTGTGVNLIIIGMWKSLAPEAKPIS--------------FNSWF 251
V+LTVVYATPIGGIST + + ++ + + +S F++WF
Sbjct: 212 VVLTVVYATPIGGISTSQFALIQFLEC-KFRPNSTQKLALSILTWLYF*RM*LGFFSTWF 388
Query: 252 FFGFPVAVLLL 262
FFGFPVA+L L
Sbjct: 389 FFGFPVAILFL 421
Score = 39.7 bits (91), Expect(2) = 8e-15
Identities = 21/48 (43%), Positives = 32/48 (65%)
Frame = +1
Query: 11 LLPLQEPIQETPNNNSITIFSILNLQNFYIILGPLLSIIICLLVKLDA 58
LLP+ Q+T +++ T+ S+L Y+I+GP LS++ICL V LDA
Sbjct: 4 LLPIH---QQTQSSSPFTLKSLL-----YVIIGPFLSLVICLFVNLDA 123
>TC223709
Length = 546
Score = 37.4 bits (85), Expect = 0.017
Identities = 17/31 (54%), Positives = 22/31 (70%)
Frame = -2
Query: 11 LLPLQEPIQETPNNNSITIFSILNLQNFYII 41
LLP+ EP++ T N S + SIL LQNFY+I
Sbjct: 98 LLPVHEPVEGTHTNISFSTKSILTLQNFYVI 6
>TC207920 similar to UP|Q6YBV9 (Q6YBV9) Cryptochrome 1, partial (40%)
Length = 954
Score = 30.4 bits (67), Expect = 2.0
Identities = 14/42 (33%), Positives = 26/42 (61%)
Frame = -2
Query: 172 VATAVMMMPVATGILHRLPPAEEQPELMNKFSRAVILTVVYA 213
VAT ++++ V +LH P EE ++ +F + V LT+V++
Sbjct: 629 VATFLIVLLVICFLLHLFPQLEEDSSILAQFQQLVALTLVHS 504
>TC225621 similar to UP|Q6NQE2 (Q6NQE2) At4g27270, partial (86%)
Length = 771
Score = 30.4 bits (67), Expect = 2.0
Identities = 25/71 (35%), Positives = 34/71 (47%), Gaps = 9/71 (12%)
Frame = +3
Query: 275 RKGSSRALSDYLDRALLKRD------LEALGPMSFAEKMVLC-VFGLLIVLWM--TRRIS 325
++ R L L R+LL + L+ L PMSF MV C VF +VLW+ + S
Sbjct: 30 KQNCGRYLKPCLKRSLLSWEHLQRVMLQLLLPMSFLRLMVFCLVFPRDLVLWLLNLKHFS 209
Query: 326 DDLPGWGVLFH 336
L +GV H
Sbjct: 210 MQLEAYGVHSH 242
>BE330194 similar to GP|10177043|dbj alcohol dehydrogenase-like protein
{Arabidopsis thaliana}, partial (16%)
Length = 500
Score = 29.3 bits (64), Expect = 4.5
Identities = 14/38 (36%), Positives = 24/38 (62%), Gaps = 6/38 (15%)
Frame = +3
Query: 148 PLNPSMLLLGLC------ATTFFVSMWLHNVATAVMMM 179
P PS+LLLGLC ++T + ++LH +A +++M
Sbjct: 186 PTTPSLLLLGLCCLAIPQSSTLSLCLFLHFIAKVLVVM 299
>TC230864 similar to UP|Q7XJ60 (Q7XJ60) At3g47690, partial (26%)
Length = 544
Score = 29.3 bits (64), Expect = 4.5
Identities = 12/26 (46%), Positives = 17/26 (65%)
Frame = +2
Query: 245 ISFNSWFFFGFPVAVLLLLCFWCILC 270
I ++ FF FP+ +LLLL W I+C
Sbjct: 242 IQKGTYCFFFFPLGLLLLLLLWFIVC 319
>TC230546 similar to GB|AAM62506.1|21553413|AY085274 VAMP
(vesicle-associated membrane protein)-associated
protein-like {Arabidopsis thaliana;} , partial (60%)
Length = 1019
Score = 29.3 bits (64), Expect = 4.5
Identities = 14/37 (37%), Positives = 22/37 (58%)
Frame = +3
Query: 21 TPNNNSITIFSILNLQNFYIILGPLLSIIICLLVKLD 57
T N NSI +F +L L + + + I++C+LV LD
Sbjct: 48 TCNRNSIPLFFLLPLSHAPTLFASHVVIVVCMLVLLD 158
>BM527934
Length = 425
Score = 28.9 bits (63), Expect = 5.9
Identities = 15/34 (44%), Positives = 19/34 (55%)
Frame = +3
Query: 61 TSTKMLGVIAWVFTWWITNAVPLPVTSMCPLFLF 94
T +L +W FTW +TN VP T+MC L F
Sbjct: 60 TGHSLLTCFSW-FTWLLTNCVP---TTMCALHAF 149
>TC229040 weakly similar to GB|AAP68276.1|31711840|BT008837 At3g61310
{Arabidopsis thaliana;} , partial (42%)
Length = 893
Score = 28.9 bits (63), Expect = 5.9
Identities = 11/24 (45%), Positives = 13/24 (53%)
Frame = -2
Query: 250 WFFFGFPVAVLLLLCFWCILCLIY 273
W F FP LLL +WC+ C Y
Sbjct: 754 WHFIKFPRVSWLLLWWWCLPCCSY 683
>TC205046 similar to UP|P93166 (P93166) SCOF-1, partial (43%)
Length = 1110
Score = 28.9 bits (63), Expect = 5.9
Identities = 14/40 (35%), Positives = 25/40 (62%)
Frame = +3
Query: 256 PVAVLLLLCFWCILCLIYVRKGSSRALSDYLDRALLKRDL 295
PV ++ +LCF+ L L+Y+ GS SD++ L +++L
Sbjct: 891 PVNLIFILCFFFCLFLVYI*VGSGLG-SDFVSAPLYRQNL 1007
>BG362730
Length = 508
Score = 28.5 bits (62), Expect = 7.7
Identities = 14/37 (37%), Positives = 19/37 (50%), Gaps = 3/37 (8%)
Frame = +1
Query: 245 ISFNSWFFFGFPVAV---LLLLCFWCILCLIYVRKGS 278
+ F FF F V + LLLL +W +LC+ Y S
Sbjct: 52 VVFKCLFFIAFDVLMVQFLLLLMYWSVLCMNYAHFAS 162
>TC219050 similar to UP|AHM2_ARATH (O64474) Potential
cadmium/zinc-transporting ATPase 2 , partial (4%)
Length = 589
Score = 28.5 bits (62), Expect = 7.7
Identities = 14/67 (20%), Positives = 31/67 (45%)
Frame = +2
Query: 392 VQSSGLADVMSKALEFLKDVPYLAIVPAVSLLCSIITEFITSNDATATLLVPLLYHIAIN 451
+ ++ +A K L++V L +VP + ++ ++ T ++ + H ++
Sbjct: 272 IATNVVAQKRQKNCSHLRNVVVLMVVPLIIIITTVNINMNTITMTNTINIIRSVPHKQVS 451
Query: 452 MHVHPLL 458
HVHP L
Sbjct: 452 QHVHPAL 472
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.327 0.142 0.440
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,763,255
Number of Sequences: 63676
Number of extensions: 426943
Number of successful extensions: 3570
Number of sequences better than 10.0: 33
Number of HSP's better than 10.0 without gapping: 3524
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3564
length of query: 530
length of database: 12,639,632
effective HSP length: 102
effective length of query: 428
effective length of database: 6,144,680
effective search space: 2629923040
effective search space used: 2629923040
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC148482.2


