

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148470.10 - phase: 0 /pseudo
(598 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
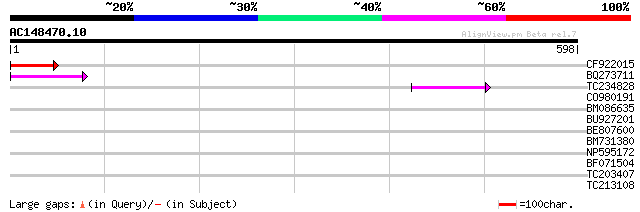
CF922015 82 5e-16
BQ273711 82 7e-16
TC234828 55 1e-07
CO980191 42 0.001
BM086635 33 0.47
BU927201 32 0.61
BE807600 similar to GP|24461867|gb CTV.22 {Poncirus trifoliata},... 30 4.0
BM731380 29 5.2
NP595172 polyprotein [Glycine max] 29 5.2
BF071504 29 6.8
TC203407 weakly similar to UP|Q9ZT16 (Q9ZT16) Arabinogalactan-pr... 28 8.9
TC213108 28 8.9
>CF922015
Length = 172
Score = 82.4 bits (202), Expect = 5e-16
Identities = 35/51 (68%), Positives = 43/51 (83%)
Frame = -3
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEA 51
F RN+PPTFKG YDP+GA+ WL+E+E IFRVM+C + QKV F THMLA+EA
Sbjct: 155 FQRNNPPTFKGGYDPEGAEAWLREIEKIFRVMECQDHQKVLFATHMLADEA 3
>BQ273711
Length = 409
Score = 82.0 bits (201), Expect = 7e-16
Identities = 36/82 (43%), Positives = 45/82 (53%)
Frame = -2
Query: 1 FLRNHPPTFKGRYDPDGAQKWLKEVESIFRVMQCSEVQKVRFGTHMLAEEANDWWVSLLP 60
F +NHPP F G YDP+GA+ WL E E IF M C E KV + T ML EA +WW + P
Sbjct: 246 FRKNHPPKFSGDYDPEGARLWLAETEKIFEAMGCLEEHKVLYATFMLQGEAENWWKFVKP 67
Query: 61 ILEQDDAAMTWAVFRREFLNRY 82
+ W F+ +FL Y
Sbjct: 66 SFVAPGGVIPWNAFKDKFLENY 1
>TC234828
Length = 857
Score = 54.7 bits (130), Expect = 1e-07
Identities = 34/84 (40%), Positives = 43/84 (50%)
Frame = +1
Query: 424 CRWRIQL*LIDCQL*MNFQKFFLMRFLMCHQRGRLNFQLILFPERSRCRWHLTVCRRPS* 483
C W+ L L+ + NFQKFF M F+ CH R + N L R +CR L C R +*
Sbjct: 598 CMWKRTLKLVVYRWCQNFQKFFRMMFVNCHLREKWNSLLTWCLGRIQCRLRLIECLRWN* 777
Query: 484 LN*RNSWRTCLIRNL*DQVFHLGE 507
*R+ +RT NL Q H GE
Sbjct: 778 QR*RHKYRTF*ANNLFVQAHHRGE 849
>CO980191
Length = 802
Score = 41.6 bits (96), Expect = 0.001
Identities = 25/81 (30%), Positives = 39/81 (47%), Gaps = 6/81 (7%)
Frame = +1
Query: 40 VRFGTHM------LAEEANDWWVSLLPILEQDDAAMTWAVFRREFLNRYVPEDVRGQKEI 93
+ F HM + E N + + L + +T VF+ FL +++ ED+ K+
Sbjct: 259 IAFTKHMNLFVSHIYREGNTYEIKLASHGTT*NRVIT*EVFKEIFLEKHLHEDI*NHKKT 438
Query: 94 EFLELKQGDMSVTEYAAEFSK 114
EFLELK G+M V Y F +
Sbjct: 439 EFLELKHGNMIVANYLVNFDE 501
>BM086635
Length = 428
Score = 32.7 bits (73), Expect = 0.47
Identities = 16/29 (55%), Positives = 24/29 (82%)
Frame = +3
Query: 286 VSLIVLL*LLL*ILVLLIVSLLLIVHISW 314
V+L++LL LLL +L+LL++ LLL+VH W
Sbjct: 156 VALLLLLLLLLLLLLLLLLLLLLLVHSVW 242
>BU927201
Length = 444
Score = 32.3 bits (72), Expect = 0.61
Identities = 26/63 (41%), Positives = 34/63 (53%)
Frame = -2
Query: 249 GILAHSARSLRRFGQVARYLH*LVRRL*IRTDLSEVLVSLIVLL*LLL*ILVLLIVSLLL 308
G L H R +RR + A L RRL V ++LL LLL +LVLL++ LL+
Sbjct: 242 GALPHHRRRMRRRRRSAVELR--TRRL----------VERLLLLLLLLLVLVLLVLMLLI 99
Query: 309 IVH 311
IVH
Sbjct: 98 IVH 90
>BE807600 similar to GP|24461867|gb CTV.22 {Poncirus trifoliata}, partial
(4%)
Length = 414
Score = 29.6 bits (65), Expect = 4.0
Identities = 22/57 (38%), Positives = 38/57 (66%)
Frame = -2
Query: 285 LVSLIVLL*LLL*ILVLLIVSLLLIVHISWVWLYLI*MEKWLLKLQLRVQ*LLLLFV 341
++ L++LL LLL +L+LL++ LLL + + L+L+ L+ + LR+ *LLLL +
Sbjct: 407 ILLLLLLLLLLLLLLLLLLLLLLLELLPQKLMLHLL-----LVDILLRIL*LLLLLL 252
>BM731380
Length = 433
Score = 29.3 bits (64), Expect = 5.2
Identities = 23/57 (40%), Positives = 36/57 (62%)
Frame = +2
Query: 285 LVSLIVLL*LLL*ILVLLIVSLLLIVHISWVWLYLI*MEKWLLKLQLRVQ*LLLLFV 341
L+ L++LL +L+ L L++V LLL++ + V L L+ + LL L +Q LLLL V
Sbjct: 236 LLLLLLLLLVLVLGLYLVLVLLLLVLLLLLVLLLLLQVLLLLLVFLLLLQVLLLLLV 406
Score = 28.5 bits (62), Expect = 8.9
Identities = 19/55 (34%), Positives = 36/55 (64%)
Frame = +2
Query: 284 VLVSLIVLL*LLL*ILVLLIVSLLLIVHISWVWLYLI*MEKWLLKLQLRVQ*LLL 338
+++ L ++L LLL +L+LL+V LLL+ + + ++L+ ++ LL L L + LL
Sbjct: 266 LVLGLYLVLVLLLLVLLLLLVLLLLLQVLLLLLVFLLLLQVLLLLLVLLLAQFLL 430
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 29.3 bits (64), Expect = 5.2
Identities = 20/39 (51%), Positives = 24/39 (61%)
Frame = +2
Query: 560 I*GQVTIRLR*RMRICRRRPLGHDMVIMNKR*CLSVLLI 598
I*GQ TI+ +RI R+ L H M IMN *C VLL+
Sbjct: 2060 I*GQDTIKFWYSLRIERKLLLEHIMAIMNG**CHLVLLM 2176
>BF071504
Length = 422
Score = 28.9 bits (63), Expect = 6.8
Identities = 27/66 (40%), Positives = 40/66 (59%)
Frame = -3
Query: 284 VLVSLIVLL*LLL*ILVLLIVSLLLIVHISWVWLYLI*MEKWLLKLQLRVQ*LLLLFV*V 343
+L+ L++LL LL* ++L++ LLL+ L+* +K LL L L LLLL + +
Sbjct: 354 LLLMLLLLLLQLL*EQIMLLMLLLLL---------LL*-QKLLLLLLL----LLLLLLLL 217
Query: 344 VHCLCL 349
V CLCL
Sbjct: 216 VQCLCL 199
>TC203407 weakly similar to UP|Q9ZT16 (Q9ZT16) Arabinogalactan-protein
(AtAGP4) (AT5g10430/F12B17_220), partial (50%)
Length = 966
Score = 28.5 bits (62), Expect = 8.9
Identities = 18/47 (38%), Positives = 30/47 (63%), Gaps = 4/47 (8%)
Frame = -2
Query: 285 LVSLIVLL*LLL*ILVLLIVSLLLIVHI----SWVWLYLI*MEKWLL 327
LV L+++L LLL + +LL+V LLL+V + VW L+ + W++
Sbjct: 383 LVVLVLVLVLLLVVELLLVVELLLVVGLLVVGLLVWWVLVQVLIWVV 243
>TC213108
Length = 767
Score = 28.5 bits (62), Expect = 8.9
Identities = 12/31 (38%), Positives = 15/31 (47%)
Frame = +2
Query: 449 FLMCHQRGRLNFQLILFPERSRCRWHLTVCR 479
F+ CH + L P+R C WH T CR
Sbjct: 338 FVPCH*SEFYFM*MFLLPKRLNCIWHQTCCR 430
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.370 0.166 0.609
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,764,320
Number of Sequences: 63676
Number of extensions: 409232
Number of successful extensions: 5542
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 2181
Number of HSP's successfully gapped in prelim test: 198
Number of HSP's that attempted gapping in prelim test: 3074
Number of HSP's gapped (non-prelim): 2593
length of query: 598
length of database: 12,639,632
effective HSP length: 103
effective length of query: 495
effective length of database: 6,081,004
effective search space: 3010096980
effective search space used: 3010096980
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.8 bits)
S2: 62 (28.5 bits)
Medicago: description of AC148470.10


