

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148405.4 - phase: 0
(609 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
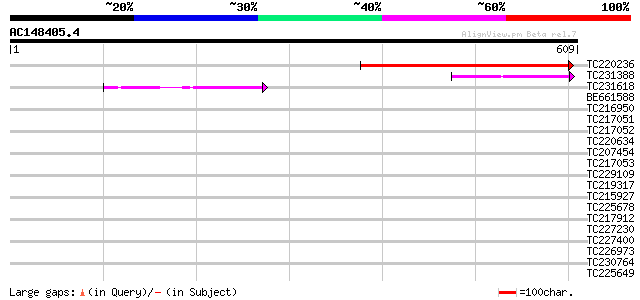
Score E
Sequences producing significant alignments: (bits) Value
TC220236 weakly similar to UP|Q9LSB9 (Q9LSB9) Emb|CAA16573.1 (At... 350 1e-96
TC231388 weakly similar to UP|Q9LSB9 (Q9LSB9) Emb|CAA16573.1 (At... 85 1e-16
TC231618 56 4e-08
BE661588 40 0.002
TC216950 39 0.009
TC217051 33 0.28
TC217052 33 0.48
TC220634 32 0.63
TC207454 32 0.63
TC217053 32 0.63
TC229109 similar to UP|Q6LA42 (Q6LA42) Pseudo-response regulator... 32 0.82
TC219317 similar to GB|AAP13385.1|30023704|BT006277 At3g56360 {A... 31 1.8
TC215927 similar to UP|Q8VWP8 (Q8VWP8) Acyltransferase-like prot... 30 3.1
TC225678 similar to UP|Q6Y236 (Q6Y236) Kinase substrate HASPP28 ... 30 3.1
TC217912 weakly similar to UP|MFP1_ARATH (Q9LW85) MAR binding fi... 30 4.1
TC227230 similar to UP|P93839 (P93839) G/HBF-1, partial (72%) 30 4.1
TC227400 similar to UP|Q6R0G3 (Q6R0G3) MYB transcription factor,... 29 5.3
TC226973 similar to UP|Q6QU76 (Q6QU76) SMC3, partial (36%) 29 5.3
TC230764 similar to UP|Q9FPT0 (Q9FPT0) Ubiquitin-specific protea... 26 6.0
TC225649 similar to UP|Q6UNT3 (Q6UNT3) Hypersensitive-induced re... 29 6.9
>TC220236 weakly similar to UP|Q9LSB9 (Q9LSB9) Emb|CAA16573.1 (At3g15920),
partial (15%)
Length = 1001
Score = 350 bits (897), Expect = 1e-96
Identities = 186/229 (81%), Positives = 202/229 (87%)
Frame = +1
Query: 377 EELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAAREQLEILSKQYGELE 436
EELRR+SLEM KLKSE G N QDLTKESIVQQNDVLL++LDA +EQLEILSKQYGELE
Sbjct: 1 EELRRQSLEMGDKLKSELGRNSYQDLTKESIVQQNDVLLENLDATKEQLEILSKQYGELE 180
Query: 437 AKSKADIKVLVKEVKSLRSSQTELKKELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQL 496
AKSKAD+KVLVKEVKSLR+SQT+LKKELSESIKE E EKLLLHEREK EQAE+A R L
Sbjct: 181 AKSKADVKVLVKEVKSLRNSQTKLKKELSESIKEHSETEKLLLHEREKREQAEVARRILL 360
Query: 497 EKCGLLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQIDILLAEVENLEKDA 556
EKC +L QLQEC+V LPYE E RT + SSSSTDAFN+L TSDDQ+DILLAEVENLEKD
Sbjct: 361 EKCRVLFNQLQECNVSLPYEYEGRTIVNSSSSTDAFNQLTTSDDQMDILLAEVENLEKDY 540
Query: 557 GSAASNVDKTNDIKDGVICGDEMRKIISDLFLDNVRLRKQTNRVTRHML 605
GSAA NVDKTNDIKDGVIC DE+RKII+DLF+DNVRLRKQTNRVTRH L
Sbjct: 541 GSAACNVDKTNDIKDGVICDDEVRKIIADLFIDNVRLRKQTNRVTRHAL 687
>TC231388 weakly similar to UP|Q9LSB9 (Q9LSB9) Emb|CAA16573.1 (At3g15920),
partial (10%)
Length = 684
Score = 84.7 bits (208), Expect = 1e-16
Identities = 50/134 (37%), Positives = 80/134 (59%), Gaps = 2/134 (1%)
Frame = +1
Query: 475 EKLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQSSSSTDAFNK 534
+++L E+++ E + A K L +C +L K+L+ECSV E+ED+ + + S DA +
Sbjct: 4 QRILQKEKQRMENSHNANTKLLHECAILQKRLRECSVNFLVEEEDKLNIDTLPS-DALDL 180
Query: 535 LKTSDDQIDILLAEVENLEKDAGSAASNVDKTNDIKDGVI--CGDEMRKIISDLFLDNVR 592
L TSD++I +LLAE + L +D V++T DI DE+RK+++ +F+DN
Sbjct: 181 LATSDNRIGLLLAEAQLLAQDVEDTVVAVEETRDITTDNTGKTHDELRKMLAHMFVDNAS 360
Query: 593 LRKQTNRVTRHMLS 606
LRKQ N V R L+
Sbjct: 361 LRKQINSVIRCALN 402
>TC231618
Length = 499
Score = 56.2 bits (134), Expect = 4e-08
Identities = 49/177 (27%), Positives = 80/177 (44%)
Frame = +3
Query: 101 SLTAGSSSVASDYGSDTPYDASEVGTPRIGQDDNSEVGTDDLTLDEDTTNPIEKLVKYGI 160
S+ A SS A G++ + SE+GTP+ G+D+ S+ D+ TL+ NP E V
Sbjct: 30 SVKAHGSSAAIVPGNN---EVSELGTPQHGKDNCSDQIMDNSTLEHGLINPTETNVHCAT 200
Query: 161 SNIDEGLFMGQTILEQLEGLPRHKVNARHGNNVTGKDRSNGNSYDSSLLGNNTMELFSQP 220
S+ N + +N+T D+ G + D+ + + E
Sbjct: 201 SS----------------------ENFVYKDNIT--DKVTGYTADAIAIHLDGTEFTPAV 308
Query: 221 GHAKVLGHIRKLSNESVGSDGSSIRGSDISNFGIPNSSGDGSVDLPGSASVSRETDI 277
+ H+++LS ES+ SD SS+R ++ SN D S +L GS SR +D+
Sbjct: 309 RDYNLNAHVKRLSTESIRSDLSSLRNTETSNLTTTTLVQDASHNLAGSHEASRNSDL 479
>BE661588
Length = 697
Score = 40.4 bits (93), Expect = 0.002
Identities = 55/211 (26%), Positives = 93/211 (44%), Gaps = 12/211 (5%)
Frame = +1
Query: 300 KLNRILSTMQRRLVTAKTD--MEDLIVRLNQEIAAKDFLTTKVKDLEVELETSKQKNKEN 357
K +RI + +R LV + + ED+ Q AA+ +K+L+ + ++ K N
Sbjct: 10 KAHRIHTVERRTLVEQELEKAQEDIPEYKKQAEAAEQEKGQVLKELD-STKRLIEELKLN 186
Query: 358 LQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKESIVQQNDVLLQD 417
L++A ER Q + D E + + EME + ES +V E + + D
Sbjct: 187 LERAQTEER----QARQDSELAKLRVEEMEQGIADES--SVAAKAQLEVAKARYTAAVSD 348
Query: 418 LDAAREQLEILSKQYGELEAKSKADIKVLVKEV---KSLRSSQTELKKEL---SESIKEK 471
L A +E+L L K+Y L IK + V K + S +L EL ES++
Sbjct: 349 LIAVKEELAALHKEYASLVTDRDVAIKKAEEAVAASKEVEKSVEDLTVELIAAKESLETT 528
Query: 472 Y----EAEKLLLHEREKSEQAEIAWRKQLEK 498
+ EAE+ + +Q + W K+L++
Sbjct: 529 HAAHLEAEEQRIGTVMARDQDSLNWEKELKQ 621
>TC216950
Length = 873
Score = 38.5 bits (88), Expect = 0.009
Identities = 37/162 (22%), Positives = 73/162 (44%), Gaps = 12/162 (7%)
Frame = +2
Query: 341 KDLEVELETSKQKNKENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQ 400
+ LE EL+ +++ + +++AI R+R E L+ + +++E++ + E G
Sbjct: 203 RQLEAELKLIEEETAKRVEEAI---RKRVE------ESLKSEEVQVEIERRLEEGRK--- 346
Query: 401 DLTKESIVQQNDVLLQDLDAAREQLEILSKQYGELEAKSKADIKVLVKEVK--------- 451
+ ND + L+ +E I +KQ E K K D++ +++E +
Sbjct: 347 --------RLNDEVAAQLEKEKEAALIEAKQKEEQARKEKEDLERMLEENRRKVEEAQRR 502
Query: 452 ---SLRSSQTELKKELSESIKEKYEAEKLLLHEREKSEQAEI 490
+ + E KEL E ++K EA + HE E+ +I
Sbjct: 503 EALEQQRREEERYKELEEMQRQKEEAMRKKKHEEEQERLNQI 628
>TC217051
Length = 582
Score = 33.5 bits (75), Expect = 0.28
Identities = 27/102 (26%), Positives = 49/102 (47%), Gaps = 3/102 (2%)
Frame = +2
Query: 101 SLTAGSSSVASDYGSDTPYDASEVGTPR-IGQDDNSEVGTDDLTLDEDTTNPIEKL--VK 157
S+ A S V S YG PY + + +PR +G + ++ D L LD++ T ++ +K
Sbjct: 215 SMVANSHRVGSSYGGAAPYRSRDGLSPRPVGASEEIQLRIDPLDLDDEITGLHRQVRRLK 394
Query: 158 YGISNIDEGLFMGQTILEQLEGLPRHKVNARHGNNVTGKDRS 199
+ I + + LE+L+ + K A NN+ ++S
Sbjct: 395 HVAEEIGTEVKYQKNFLEELQ-MTMIKAQAGVKNNLRRLNKS 517
>TC217052
Length = 793
Score = 32.7 bits (73), Expect = 0.48
Identities = 26/102 (25%), Positives = 49/102 (47%), Gaps = 3/102 (2%)
Frame = +1
Query: 101 SLTAGSSSVASDYGSDTPYDASE-VGTPRIGQDDNSEVGTDDLTLDEDTTNPIEKL--VK 157
S+ A S + S YG PY + + + T +G + ++ D L LD++ T ++ +K
Sbjct: 205 SMAANSHRLGSSYGGAAPYRSRDGLSTRPVGASEEIQLRIDPLDLDDEITGLHRQVRRLK 384
Query: 158 YGISNIDEGLFMGQTILEQLEGLPRHKVNARHGNNVTGKDRS 199
+ I + +T LE+L+ + K A NN+ ++S
Sbjct: 385 HVAEEIGTEVKYQKTFLEELQ-MTMIKAQAGVKNNLRRLNKS 507
>TC220634
Length = 583
Score = 32.3 bits (72), Expect = 0.63
Identities = 33/125 (26%), Positives = 58/125 (46%), Gaps = 2/125 (1%)
Frame = +2
Query: 449 EVKSLRSSQTELKKELSESIKEKYEAEKLLLHEREKSEQAEIAWRKQLEKCGLLLKQLQE 508
E++S+ EL+ +ESIK+ E KLL ++ E EK LL ++ E
Sbjct: 26 ELQSMFQENDELRSREAESIKKLEELSKLLEEATTRNHTEENGDLTDSEKDYDLLPKVVE 205
Query: 509 CSVELPYEDEDRTFLQSSSSTDAFNKLKTSDDQIDILLA--EVENLEKDAGSAASNVDKT 566
S E ED + ++ S + + +LK ++ Q D +L+ + E +E S K
Sbjct: 206 FSEENGLVGEDISKVELSVNQE---ELKQNNMQEDSILSNDKAEKIESPKHEEVSGKRKE 376
Query: 567 NDIKD 571
++ K+
Sbjct: 377 DETKE 391
>TC207454
Length = 1270
Score = 32.3 bits (72), Expect = 0.63
Identities = 47/201 (23%), Positives = 87/201 (42%), Gaps = 7/201 (3%)
Frame = +3
Query: 60 SKNSDPIVMSAFQDASQQNSVKDPDSNNRDYSVQSSLHSGLSLTAGSSSVASDYGSDTPY 119
S+ + P++ AS N D ++D S +SS+ S LSL + + S D+P
Sbjct: 492 SEENSPVISCP---ASPPNEAND----HKDDSQKSSVGSNLSLDIINETSLSSSPIDSPL 650
Query: 120 ---DASEVGTPRIGQDDNSEVGTDDLTLDEDTTNPIEKLVKYGISNIDEGLFMGQTILEQ 176
D E G G+++ + + T E N + L K+G +N G FM + +
Sbjct: 651 PISDHPENGPQTPGRNNINNSAGNSTTNSERKLNKFQWLWKFGRNN---GEFMSEKGGDT 821
Query: 177 LEGLPRHKVNARHGNNVTGKDRSNGNSYDSSLLGNNTMELFSQPGH----AKVLGHIRKL 232
E + N + +N T +N + S+ + ++T + + + + GH R +
Sbjct: 822 SEA-AKPANNCNNQSNTTPSSTANNCNNHSNTIPSSTAKNCNNQSNIIPSSTANGHRRSV 998
Query: 233 SNESVGSDGSSIRGSDISNFG 253
S + +D ++ G+ I N G
Sbjct: 999 SCQGESTD-QNVMGT-IRNIG 1055
>TC217053
Length = 740
Score = 32.3 bits (72), Expect = 0.63
Identities = 27/102 (26%), Positives = 49/102 (47%), Gaps = 3/102 (2%)
Frame = +3
Query: 101 SLTAGSSSVASDYGSDTPYDASEVGTPR-IGQDDNSEVGTDDLTLDEDTTNPIEKL--VK 157
S+ A S V S YG PY + + +PR +G + ++ D L LD++ T ++ +K
Sbjct: 30 SMVANSHRVGSFYGGAAPYRSRDGLSPRPVGASEEIQLRIDPLDLDDEITGLHRQVRRLK 209
Query: 158 YGISNIDEGLFMGQTILEQLEGLPRHKVNARHGNNVTGKDRS 199
+ I + + LE+L+ + K A NN+ ++S
Sbjct: 210 HVAEEIGTEVKYQKNFLEELQ-MTMIKAQAGVKNNLRRLNKS 332
>TC229109 similar to UP|Q6LA42 (Q6LA42) Pseudo-response regulator 5, partial
(7%)
Length = 1011
Score = 32.0 bits (71), Expect = 0.82
Identities = 28/110 (25%), Positives = 54/110 (48%), Gaps = 10/110 (9%)
Frame = +2
Query: 181 PRHKVNARHGNNVTGKDRSNGNSYDSSLLGNNTME---LFSQPG---HAKVLGHIRKLSN 234
P +VNA + +N+ K+ S+ Y+S NT + +++Q HA+ GHI ++
Sbjct: 236 PNFQVNAFYQSNM--KESSSEQLYESRGPNGNTTQNHIVYTQEHKSEHAEDRGHISPTTD 409
Query: 235 ESVGSDGSSIRGSDISNFGIPNSSGDGS----VDLPGSASVSRETDIVGH 280
+SV S + S +++ G ++ G S V+ +AS + D+ +
Sbjct: 410 QSVSSSFCNGNASHLNSIGYGSNCGSSSNVDQVNTVWAASEGKHEDLTNN 559
>TC219317 similar to GB|AAP13385.1|30023704|BT006277 At3g56360 {Arabidopsis
thaliana;} , partial (33%)
Length = 829
Score = 30.8 bits (68), Expect = 1.8
Identities = 15/40 (37%), Positives = 20/40 (49%), Gaps = 3/40 (7%)
Frame = -2
Query: 2 QRRSPPKHRHDGTSPLPLGMDW---SPAPRKWNGRDTEWP 38
QRR HR T+PLP +W +P+P W+ WP
Sbjct: 351 QRRFWLLHRVSRTAPLPSSSEW*SLAPSPWPWSSPPWPWP 232
>TC215927 similar to UP|Q8VWP8 (Q8VWP8) Acyltransferase-like protein
(Fragment), partial (82%)
Length = 1679
Score = 30.0 bits (66), Expect = 3.1
Identities = 13/33 (39%), Positives = 24/33 (72%), Gaps = 1/33 (3%)
Frame = +2
Query: 133 DNSEVGTDDLTLDEDTTNPIEKLVKY-GISNID 164
D++E+G DDLT+ E + +++L+ Y GI N++
Sbjct: 377 DDNEIGVDDLTVAEISNTNLKELIPYSGILNLE 475
>TC225678 similar to UP|Q6Y236 (Q6Y236) Kinase substrate HASPP28 (Fragment),
partial (54%)
Length = 912
Score = 30.0 bits (66), Expect = 3.1
Identities = 21/67 (31%), Positives = 35/67 (51%), Gaps = 7/67 (10%)
Frame = +1
Query: 451 KSLRSSQTELKKELSESIKEKYEAEKLLLHER-------EKSEQAEIAWRKQLEKCGLLL 503
KSL++ ++ K S +E+ E EK HER K+EQA +K LE+ L+
Sbjct: 415 KSLKARDVDVGKTTELSRREREEIEKQRAHERYMRLQEQGKTEQA----KKDLERLALIR 582
Query: 504 KQLQECS 510
+Q ++ +
Sbjct: 583 QQREDAA 603
>TC217912 weakly similar to UP|MFP1_ARATH (Q9LW85) MAR binding filament-like
protein 1, partial (16%)
Length = 756
Score = 29.6 bits (65), Expect = 4.1
Identities = 30/136 (22%), Positives = 55/136 (40%), Gaps = 14/136 (10%)
Frame = +2
Query: 346 ELETSKQKNKENLQQAILIERERFTQMQWDMEELRRKSLEMEMKLKSESGGNVGQDLTKE 405
+L S K+ + L+Q + ++E ++ D+EE + EM SG +
Sbjct: 8 QLVASLNKDLQALEQQVSKDKESRKSLERDLEEATKSLEEMNRNAVILSGELQRANSLVS 187
Query: 406 SIVQQNDVLLQDL--------------DAAREQLEILSKQYGELEAKSKADIKVLVKEVK 451
S+ ++ DVL++ L + A + L K+ LE K K +E+
Sbjct: 188 SLEKEKDVLIKSLTNQRNACKEAQDNIEDAHNLIMKLGKERENLEKKGKK----FEEELA 355
Query: 452 SLRSSQTELKKELSES 467
S + LK ++ S
Sbjct: 356 SAKGEILRLKSRINSS 403
>TC227230 similar to UP|P93839 (P93839) G/HBF-1, partial (72%)
Length = 1250
Score = 29.6 bits (65), Expect = 4.1
Identities = 40/161 (24%), Positives = 63/161 (38%), Gaps = 6/161 (3%)
Frame = +2
Query: 122 SEVGTPRIGQDDNSEVGTDDLTLDEDTTNPIE-KLVKYGISNIDEGLFMGQTILEQLEGL 180
S G+ R DD G + D T+P + K V+ +SN + + L L
Sbjct: 413 STSGSSREQSDDEDIEGETSMN---DNTDPADVKRVRRMLSNRESARRSRRRKQAHLTDL 583
Query: 181 PRHKVNARHGNNVTGKDRSN-GNSYDSSLLGNNTMELFSQPGHAKVLGHIRKLSNESV-- 237
R N+ K ++ Y S + N ++ + AKV K++ E+V
Sbjct: 584 ETQVSQLRGENSTLLKRLTDVSQKYSDSAVDNRVLKADVETLRAKV-----KMAEETVKR 748
Query: 238 --GSDGSSIRGSDISNFGIPNSSGDGSVDLPGSASVSRETD 276
G + SDIS+ G+P+ G D ASV + D
Sbjct: 749 ITGLNPMPHAMSDISSLGLPSFDGRSPSDTSADASVPVQDD 871
>TC227400 similar to UP|Q6R0G3 (Q6R0G3) MYB transcription factor, partial
(30%)
Length = 1488
Score = 29.3 bits (64), Expect = 5.3
Identities = 22/84 (26%), Positives = 44/84 (52%), Gaps = 1/84 (1%)
Frame = +1
Query: 76 QQNSVKDPDSNNRDYSVQSSLHSGLSLTAGSSSVASDYGSDTPYDA-SEVGTPRIGQDDN 134
+ NS+K D + + S +S L S + GSS + YGS +P + S + T + ++
Sbjct: 517 KSNSLKSSDFDQENQSPKSVLSGVGSDSLGSSDSDTPYGSLSPMSSISGIHTSSFTRAEH 696
Query: 135 SEVGTDDLTLDEDTTNPIEKLVKY 158
+ +D+ +D D+ + + L+K+
Sbjct: 697 -KTTSDEAGMDTDSAHDEKPLMKF 765
>TC226973 similar to UP|Q6QU76 (Q6QU76) SMC3, partial (36%)
Length = 1707
Score = 29.3 bits (64), Expect = 5.3
Identities = 34/137 (24%), Positives = 65/137 (46%), Gaps = 16/137 (11%)
Frame = +3
Query: 387 EMKLKSESGGNVGQDLTKESIVQQNDVLLQDLDAAREQLEI-----LSKQYGELEAK-SK 440
E KL S+ + +DL ++ + + D + + +A R +L+ L ++ ELEA S
Sbjct: 99 EKKLLSDLNPEI-KDLKEKLVACKTDRI--ETEARRAELDTNLTTNLRRRKQELEAVISS 269
Query: 441 ADIKVLVKEVKSLRSSQTELK----------KELSESIKEKYEAEKLLLHEREKSEQAEI 490
AD LV + +S ++ K + ++ESI ++ K + E K + E
Sbjct: 270 ADADSLVADAESKWQELSDAKILVDDAIGQLRSVTESINDRTRQIKKIKDELNKLKSLED 449
Query: 491 AWRKQLEKCGLLLKQLQ 507
+ ++L++ L+QLQ
Sbjct: 450 EYERKLQEDAKELEQLQ 500
>TC230764 similar to UP|Q9FPT0 (Q9FPT0) Ubiquitin-specific protease 14,
partial (15%)
Length = 767
Score = 26.2 bits (56), Expect(2) = 6.0
Identities = 9/23 (39%), Positives = 14/23 (60%)
Frame = -3
Query: 43 TGWSYCVIIPSWVFVPKSKNSDP 65
T W +CV +P W+ +P S+ P
Sbjct: 261 T*WPHCVEVPLWLTIPISRYFPP 193
Score = 21.2 bits (43), Expect(2) = 6.0
Identities = 8/18 (44%), Positives = 9/18 (49%)
Frame = -1
Query: 10 RHDGTSPLPLGMDWSPAP 27
R + SP PL W P P
Sbjct: 362 RRNKDSPFPLEDQWMPRP 309
>TC225649 similar to UP|Q6UNT3 (Q6UNT3) Hypersensitive-induced response
protein, complete
Length = 1260
Score = 28.9 bits (63), Expect = 6.9
Identities = 28/110 (25%), Positives = 48/110 (43%), Gaps = 15/110 (13%)
Frame = +3
Query: 490 IAWRKQLEKCGLLLKQLQECSVELPYEDEDRTFLQ----------SSSSTDAFNKLKTSD 539
+ W + G L ++Q+ V + +D F+ S ++DAF +L +
Sbjct: 168 LPWCLGYQIAGSLSLRVQQLDVRCETKTKDNVFVNVVASVQYRAVSEKASDAFYRLTNTR 347
Query: 540 DQI-----DILLAEVENLEKDAGSAASNVDKTNDIKDGVICGDEMRKIIS 584
+QI D++ A V LE D S ++ NDI V +E+ K +S
Sbjct: 348 EQIQSYVFDVIRASVPKLELD-----SVFEQKNDIAKAV--EEELEKAMS 476
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.310 0.129 0.356
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 22,139,778
Number of Sequences: 63676
Number of extensions: 301146
Number of successful extensions: 1309
Number of sequences better than 10.0: 44
Number of HSP's better than 10.0 without gapping: 1299
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1306
length of query: 609
length of database: 12,639,632
effective HSP length: 103
effective length of query: 506
effective length of database: 6,081,004
effective search space: 3076988024
effective search space used: 3076988024
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 62 (28.5 bits)
Medicago: description of AC148405.4


