

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148405.2 + phase: 0
(209 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
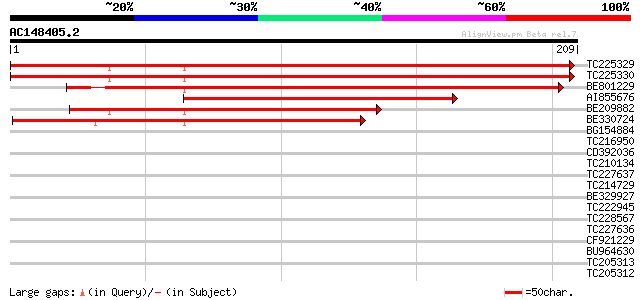
Score E
Sequences producing significant alignments: (bits) Value
TC225329 similar to UP|ATPX_SPIOL (P31853) ATP synthase B' chain... 267 3e-72
TC225330 similar to UP|ATPX_SPIOL (P31853) ATP synthase B' chain... 261 2e-70
BE801229 weakly similar to PIR|T05402|T05 H+-transporting two-se... 175 2e-44
AI855676 weakly similar to SP|P31853|ATPX_ ATP synthase B' chain... 112 1e-25
BE209882 weakly similar to SP|P31853|ATPX_ ATP synthase B' chain... 110 6e-25
BE330724 weakly similar to SP|P31853|ATPX_ ATP synthase B' chain... 102 1e-22
BG154884 38 0.004
TC216950 37 0.005
CD392036 similar to GP|29367413|gb| H+-transporting ATP synthase... 37 0.005
TC210134 34 0.042
TC227637 weakly similar to UP|Q9SYP0 (Q9SYP0) F9H16.4 protein, p... 33 0.093
TC214729 UP|LOXX_SOYBN (P24095) Seed lipoxygenase , complete 33 0.12
BE329927 32 0.16
TC222945 homologue to UP|Q8VAG3 (Q8VAG3) Wsv453 (WSSV513), parti... 32 0.27
TC228567 32 0.27
TC227636 weakly similar to UP|Q9SYP0 (Q9SYP0) F9H16.4 protein, p... 32 0.27
CF921229 32 0.27
BU964630 similar to GP|11994423|db helicase-like protein {Arabid... 31 0.46
TC205313 similar to UP|TL15_ARATH (O22160) Thylakoid lumenal 15 ... 31 0.46
TC205312 similar to UP|TL15_ARATH (O22160) Thylakoid lumenal 15 ... 31 0.46
>TC225329 similar to UP|ATPX_SPIOL (P31853) ATP synthase B' chain,
chloroplast precursor (Subunit II) , partial (67%)
Length = 1213
Score = 267 bits (682), Expect = 3e-72
Identities = 149/215 (69%), Positives = 174/215 (80%), Gaps = 7/215 (3%)
Frame = +3
Query: 1 MASMIMASSKPIIRVPTNSLPSTPKLPSLQTITPN-----LKLQLSKLKSMTLAATSLSF 55
MA+MIMAS+KP++ V T+S TPKLP LQ P LKL +SK + ++L
Sbjct: 135 MANMIMASTKPLVPVCTSSRSPTPKLPILQISLPKAPTLKLKLPISKPQMLSLLGGIAPL 314
Query: 56 VFAPPSLA--FEKAALFDFNLTLPIIVVEFLFLMFALDKLYFTPLGTFMDNRDAEIRGKL 113
V A PSLA FEKAALFDFNLTLPII+VEFL LM ALDK++FTPLG FMD RDA IR KL
Sbjct: 315 VLARPSLAEEFEKAALFDFNLTLPIIMVEFLLLMVALDKIWFTPLGKFMDERDAAIREKL 494
Query: 114 NSVTNTSEEVKELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVDAELQE 173
+SV +TSEEVK+LEE+ANAV+ AARAEIA ALN MKKETQAEVE+KIAEGRKKV+AELQE
Sbjct: 495 SSVKDTSEEVKQLEEKANAVMAAARAEIAAALNTMKKETQAEVEQKIAEGRKKVEAELQE 674
Query: 174 ALANLEKQKEETVKALDSQIAALSQDIVNKVLPVS 208
AL++LE QKEET+K+LDSQIAALSQ+IVNKVLP +
Sbjct: 675 ALSSLENQKEETIKSLDSQIAALSQEIVNKVLPTA 779
>TC225330 similar to UP|ATPX_SPIOL (P31853) ATP synthase B' chain,
chloroplast precursor (Subunit II) , partial (67%)
Length = 982
Score = 261 bits (666), Expect = 2e-70
Identities = 146/215 (67%), Positives = 172/215 (79%), Gaps = 7/215 (3%)
Frame = +1
Query: 1 MASMIMASSKPIIRVPTNSLPSTPKLPSLQTITPN-----LKLQLSKLKSMTLAATSLSF 55
MA+MI+AS+KP++ V T+S TPK+P LQ P LKL +SK + ++L
Sbjct: 133 MANMIVASTKPLVPVCTSSRSPTPKVPILQISLPKAPTLKLKLPISKPQMLSLLGGIAPL 312
Query: 56 VFAPPSLA--FEKAALFDFNLTLPIIVVEFLFLMFALDKLYFTPLGTFMDNRDAEIRGKL 113
V A PSLA EKAALFDFNLTLPII+VEFL LM ALDK++FTPLG FMD RDA IR KL
Sbjct: 313 VLARPSLAEEIEKAALFDFNLTLPIIMVEFLLLMVALDKIWFTPLGKFMDERDAAIREKL 492
Query: 114 NSVTNTSEEVKELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVDAELQE 173
+SV +TSEEVK+LEE+ANAV+ A+RAEIA ALN MKKETQAEVE+KIAEGRKKV+AELQE
Sbjct: 493 SSVKDTSEEVKQLEEKANAVMAASRAEIAAALNTMKKETQAEVEQKIAEGRKKVEAELQE 672
Query: 174 ALANLEKQKEETVKALDSQIAALSQDIVNKVLPVS 208
ALA LE QKEET+K+LDSQIAALSQ+IVNKVLP +
Sbjct: 673 ALAALENQKEETIKSLDSQIAALSQEIVNKVLPTA 777
>BE801229 weakly similar to PIR|T05402|T05 H+-transporting two-sector ATPase
(EC 3.6.3.14) chain 9 - Arabidopsis thaliana, partial
(52%)
Length = 553
Score = 175 bits (443), Expect = 2e-44
Identities = 102/185 (55%), Positives = 131/185 (70%), Gaps = 2/185 (1%)
Frame = +2
Query: 22 STPKLPSLQTITPNLKLQLSKLKSMTLAATSLSFVFAPPSLA--FEKAALFDFNLTLPII 79
S PK P+L+ LKL +SK + ++L V A PSL FEKA LFDFNLTLPII
Sbjct: 14 SLPKAPTLK-----LKLPISKPQMLSLLRGIAPLVLARPSLPEKFEKAPLFDFNLTLPII 178
Query: 80 VVEFLFLMFALDKLYFTPLGTFMDNRDAEIRGKLNSVTNTSEEVKELEEQANAVLRAARA 139
+VE L LM ALDK++ TPLG FMD RDA I+ KL S+ +TS++V L+E+ANA + AA A
Sbjct: 179 MVELLLLMVALDKIWLTPLGKFMDYRDAAIKEKLISMKDTSDDVNHLKEKANAAMAAAPA 358
Query: 140 EIAVALNQMKKETQAEVEEKIAEGRKKVDAELQEALANLEKQKEETVKALDSQIAALSQD 199
EIA LN MKK+TQA+VE+KI E RK VDAEL +A++ E Q E+T+K +D QIA L Q
Sbjct: 359 EIATQLNTMKKDTQADVEQKIDEERKTVDAELHDAMSI*ENQ*EDTLKCID*QIAPLIQQ 538
Query: 200 IVNKV 204
IV+++
Sbjct: 539 IVSRL 553
>AI855676 weakly similar to SP|P31853|ATPX_ ATP synthase B' chain
chloroplast precursor (EC 3.6.3.14) (Subunit II).
[Spinach], partial (33%)
Length = 322
Score = 112 bits (280), Expect = 1e-25
Identities = 57/101 (56%), Positives = 77/101 (75%)
Frame = +2
Query: 65 EKAALFDFNLTLPIIVVEFLFLMFALDKLYFTPLGTFMDNRDAEIRGKLNSVTNTSEEVK 124
EKA L DFNLTLPII+ EFL LM AL+ ++ TPL FMD+RD IR KL SV +TS++VK
Sbjct: 20 EKAGLCDFNLTLPIIMEEFLLLMVALNNIWITPLEKFMDDRDTTIREKLISVNDTSKDVK 199
Query: 125 ELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRK 165
LE++ANA++ A+++ I+ +LN +KKET A+VE IA +K
Sbjct: 200 HLEDKANAIITASQSYISTSLNTIKKETHADVEHNIAHEKK 322
>BE209882 weakly similar to SP|P31853|ATPX_ ATP synthase B' chain
chloroplast precursor (EC 3.6.3.14) (Subunit II).
[Spinach], partial (36%)
Length = 370
Score = 110 bits (274), Expect = 6e-25
Identities = 65/122 (53%), Positives = 81/122 (66%), Gaps = 7/122 (5%)
Frame = +2
Query: 23 TPKLPSLQTITPN-----LKLQLSKLKSMTLAATSLSFVFAPPSLA--FEKAALFDFNLT 75
TPKLP LQ P LKL +SK + ++L V+A P L EKA LFDFNLT
Sbjct: 5 TPKLPILQISLPKAPTWKLKLPISKPQMLSLLGGIAPLVWAGPLLTEELEKAGLFDFNLT 184
Query: 76 LPIIVVEFLFLMFALDKLYFTPLGTFMDNRDAEIRGKLNSVTNTSEEVKELEEQANAVLR 135
LPII+VE L LM ALDK++ TPLG MD RD IR KL+SV +T++EVK L+++ANA +
Sbjct: 185 LPIIMVECLLLMVALDKIWSTPLGKLMDERDTAIREKLSSVKDTADEVKHLDDKANAWMA 364
Query: 136 AA 137
AA
Sbjct: 365 AA 370
>BE330724 weakly similar to SP|P31853|ATPX_ ATP synthase B' chain
chloroplast precursor (EC 3.6.3.14) (Subunit II).
[Spinach], partial (33%)
Length = 421
Score = 102 bits (255), Expect = 1e-22
Identities = 66/137 (48%), Positives = 87/137 (63%), Gaps = 7/137 (5%)
Frame = +2
Query: 2 ASMIMASSKPIIRVPTNSLPSTPKLPSLQ-----TITPNLKLQLSKLKSMTLAATSLSFV 56
A+MI+AS+KP++ V T+S TPK+P LQ T LKLQ+ ++L V
Sbjct: 2 ANMIVASTKPLVPVCTSSRSPTPKVPILQM*LR*APTLKLKLQM**PHMLSLLGGIAPLV 181
Query: 57 FAPPSLA--FEKAALFDFNLTLPIIVVEFLFLMFALDKLYFTPLGTFMDNRDAEIRGKLN 114
A PSLA EKA+L D +LT II+VE L + ALD ++FTPLG FMD+RDA I K +
Sbjct: 182 *ARPSLAEEIEKASLLDSDLTRSIIMVELLLMKVALDNIWFTPLGKFMDDRDAAITEKQS 361
Query: 115 SVTNTSEEVKELEEQAN 131
+ +T EEVK LEE+AN
Sbjct: 362 NEKDT*EEVKHLEEEAN 412
>BG154884
Length = 475
Score = 37.7 bits (86), Expect = 0.004
Identities = 28/75 (37%), Positives = 42/75 (55%), Gaps = 1/75 (1%)
Frame = +3
Query: 110 RGKLNSVTNTSEEVKELEEQ-ANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVD 168
R L S + EE+K LEE+ A + A R + LN +E + E+E ++AEG KK+
Sbjct: 264 RNLLESKRHHEEELKLLEEETARRLEEAIRKNVEEKLNT--EEVKLEIERRVAEGVKKLF 437
Query: 169 AELQEALANLEKQKE 183
+++ A LEK KE
Sbjct: 438 DDVE---AQLEKGKE 473
>TC216950
Length = 873
Score = 37.4 bits (85), Expect = 0.005
Identities = 24/63 (38%), Positives = 37/63 (58%), Gaps = 1/63 (1%)
Frame = +2
Query: 122 EVKELEEQ-ANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVDAELQEALANLEK 180
E+K +EE+ A V A R + +L +E Q E+E ++ EGRK+++ E A LEK
Sbjct: 218 ELKLIEEETAKRVEEAIRKRVEESLKS--EEVQVEIERRLEEGRKRLN---DEVAAQLEK 382
Query: 181 QKE 183
+KE
Sbjct: 383 EKE 391
>CD392036 similar to GP|29367413|gb| H+-transporting ATP synthase chain
9-like protein {Oryza sativa (japonica cultivar-group)},
partial (7%)
Length = 181
Score = 37.4 bits (85), Expect = 0.005
Identities = 20/36 (55%), Positives = 23/36 (63%)
Frame = -3
Query: 94 YFTPLGTFMDNRDAEIRGKLNSVTNTSEEVKELEEQ 129
+FTPLG FMD RD IR + TSEEV LEE+
Sbjct: 179 WFTPLGKFMDERDPAIREN*LA*RTTSEEVN*LEEK 72
>TC210134
Length = 738
Score = 34.3 bits (77), Expect = 0.042
Identities = 21/60 (35%), Positives = 42/60 (70%)
Frame = +1
Query: 126 LEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVDAELQEALANLEKQKEET 185
LEE+ +A+LRA E A+ + +K++ + ++ K+ + ++V A+L++ALA +KQ++ET
Sbjct: 19 LEEKRSAILRAEELETAL-MEMVKQDNRRQLSAKVEQLDEEV-AQLRQALA--DKQEQET 186
>TC227637 weakly similar to UP|Q9SYP0 (Q9SYP0) F9H16.4 protein, partial (8%)
Length = 1330
Score = 33.1 bits (74), Expect = 0.093
Identities = 23/81 (28%), Positives = 42/81 (51%), Gaps = 2/81 (2%)
Frame = +3
Query: 121 EEVKELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEG--RKKVDAELQEALANL 178
EE +E+ +L+A +A + ++K++ + E EK E RKK +AE + A L
Sbjct: 645 EEPVRTKEEEELILKAEKARMEEEEAKLKEKRRLEEIEKAKEALLRKKRNAEKAQQRAAL 824
Query: 179 EKQKEETVKALDSQIAALSQD 199
+ QKE +K + + A ++
Sbjct: 825 KAQKEAELKEKEREKRAKKKE 887
>TC214729 UP|LOXX_SOYBN (P24095) Seed lipoxygenase , complete
Length = 2891
Score = 32.7 bits (73), Expect = 0.12
Identities = 18/48 (37%), Positives = 26/48 (53%), Gaps = 2/48 (4%)
Frame = -3
Query: 16 PTNSLPSTPKLPSLQTITPNLKLQLSKLKSMTLAA--TSLSFVFAPPS 61
P L PK+P +ITP L +++ LKS T T++ F+F P S
Sbjct: 234 PPTKLTPWPKMPVAVSITPPLPMEVMALKSNTFFGINTTVPFIFCPLS 91
>BE329927
Length = 464
Score = 32.3 bits (72), Expect = 0.16
Identities = 25/85 (29%), Positives = 43/85 (50%), Gaps = 6/85 (7%)
Frame = +3
Query: 99 GTFMDNRDAEIRGKLNSVTNTSEEVKELEEQANAVLRAARAEIAVALNQMKKET----QA 154
G F D+ E GK+ TN +++ + +EEQAN + + + A + + T QA
Sbjct: 78 GFFFDSD--EEAGKMQKSTNQTDQNENVEEQANGKSKGKTDKTSGAGKKRGRSTGGKPQA 251
Query: 155 EVEEKIAEGRKKVDAE--LQEALAN 177
++ +K G KK + ++EAL N
Sbjct: 252 KIAKKKTPGSKKAKTKK*VKEALGN 326
>TC222945 homologue to UP|Q8VAG3 (Q8VAG3) Wsv453 (WSSV513), partial (7%)
Length = 831
Score = 31.6 bits (70), Expect = 0.27
Identities = 30/94 (31%), Positives = 42/94 (43%), Gaps = 26/94 (27%)
Frame = -1
Query: 116 VTNTSEEVKEL-------EEQANAVLRAARAEIAVA----------LNQMKKETQAEVE- 157
VT T EE EL EEQAN + AA ++I +A L ++ +E E
Sbjct: 621 VTLTLEEYHELSRRAYKAEEQANMRIEAANSQIQIARESELRSLEKLKELNEELSVRRES 442
Query: 158 --------EKIAEGRKKVDAELQEALANLEKQKE 183
+K EG+ V+ EL+ A EKQK+
Sbjct: 441 LSIATENFKKANEGKLAVEHELRTWRAEQEKQKK 340
>TC228567
Length = 1531
Score = 31.6 bits (70), Expect = 0.27
Identities = 30/96 (31%), Positives = 44/96 (45%), Gaps = 26/96 (27%)
Frame = +2
Query: 116 VTNTSEEVKEL-------EEQANAVLRAARAEIAVA----------LNQMKKETQAEVE- 157
VT T EE EL EEQANA + AA ++I +A L ++ +E E
Sbjct: 887 VTLTLEEYHELSRRAYKAEEQANARIEAATSQIQIARESELRSLEKLEELNEELSVRRES 1066
Query: 158 --------EKIAEGRKKVDAELQEALANLEKQKEET 185
EK EG+ V+ EL+ A ++Q++ T
Sbjct: 1067LKIATGNSEKANEGKLAVEHELRTWRAEQKQQEKAT 1174
>TC227636 weakly similar to UP|Q9SYP0 (Q9SYP0) F9H16.4 protein, partial (8%)
Length = 1200
Score = 31.6 bits (70), Expect = 0.27
Identities = 23/81 (28%), Positives = 41/81 (50%), Gaps = 2/81 (2%)
Frame = +3
Query: 121 EEVKELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEG--RKKVDAELQEALANL 178
EE +E+ +L+A +A ++K++ + E EK E RKK +AE + A L
Sbjct: 402 EEPVRTKEEEELILKAEKARKEEEEAKLKEKRRLEEIEKAKEALQRKKRNAEKAQQRAAL 581
Query: 179 EKQKEETVKALDSQIAALSQD 199
+ QKE +K + + A ++
Sbjct: 582 KAQKEAELKEKEREKRAKKKE 644
>CF921229
Length = 680
Score = 31.6 bits (70), Expect = 0.27
Identities = 17/57 (29%), Positives = 33/57 (57%)
Frame = -3
Query: 116 VTNTSEEVKELEEQANAVLRAARAEIAVALNQMKKETQAEVEEKIAEGRKKVDAELQ 172
VT +EV+ +EE+ + + E+ V + + KKE++ + EEK G++ V E++
Sbjct: 444 VTEGKKEVEVIEEKKETEVTEEKKEVEVEVREEKKESEVKEEEK---GQEVVPEEVE 283
Score = 27.3 bits (59), Expect = 5.1
Identities = 25/67 (37%), Positives = 38/67 (56%), Gaps = 5/67 (7%)
Frame = +3
Query: 1 MASMIMASSKPIIRVPTN---SLPSTPKLPSLQ--TITPNLKLQLSKLKSMTLAATSLSF 55
+ S+ + SS P +++P++ SLP LPSLQ + P+L L S L S+ L ++ L
Sbjct: 435 LRSLPLPSSLPSLQLPSSLLLSLPLPSSLPSLQLPSSLPSLPLP-SSLLSLLLPSSLLLL 611
Query: 56 VFAPPSL 62
F P SL
Sbjct: 612 QF-PSSL 629
>BU964630 similar to GP|11994423|db helicase-like protein {Arabidopsis
thaliana}, partial (3%)
Length = 452
Score = 30.8 bits (68), Expect = 0.46
Identities = 31/126 (24%), Positives = 54/126 (42%), Gaps = 12/126 (9%)
Frame = +1
Query: 52 SLSFVFAPPSLAFEKAALFDFNLTLPIIVVE---FLFLMFALDKLYFTP---LGTFMDNR 105
S++ +F P F F+ +P+I E F FL F+ DK+ F+P L T M ++
Sbjct: 94 SIATLFTPSFFFFSTDNTLFFSAPIPLIRCELQRFPFLKFSRDKICFSPCEVLATAMASK 273
Query: 106 ------DAEIRGKLNSVTNTSEEVKELEEQANAVLRAARAEIAVALNQMKKETQAEVEEK 159
D E R K E + + + VL + + K+ ++E + K
Sbjct: 274 GPRSRIDHESRAKRQKALEAPREPRRPKTHWDHVLE--------EMVWLSKDFESERKWK 429
Query: 160 IAEGRK 165
+A+ +K
Sbjct: 430 LAQAKK 447
>TC205313 similar to UP|TL15_ARATH (O22160) Thylakoid lumenal 15 kDa protein,
chloroplast precursor (p15), partial (76%)
Length = 945
Score = 30.8 bits (68), Expect = 0.46
Identities = 22/64 (34%), Positives = 35/64 (54%), Gaps = 4/64 (6%)
Frame = +2
Query: 6 MASSKPIIRVPTNSLPSTPKLPSLQTITPNLKLQLSKLKSMT----LAATSLSFVFAPPS 61
+++S P T LP LP L+++ P+ KL S +S+ +A S SF+F P+
Sbjct: 86 LSTSNPNKASSTFPLP----LPLLKSLQPSQKLTFSLGESVAGATLVALLSASFIFVDPA 253
Query: 62 LAFE 65
LAF+
Sbjct: 254 LAFK 265
>TC205312 similar to UP|TL15_ARATH (O22160) Thylakoid lumenal 15 kDa protein,
chloroplast precursor (p15), partial (76%)
Length = 950
Score = 30.8 bits (68), Expect = 0.46
Identities = 25/69 (36%), Positives = 36/69 (51%), Gaps = 12/69 (17%)
Frame = +1
Query: 9 SKPIIRVPT---NSLPSTPKLPSL-----QTITPNLKLQLSKLKSMT----LAATSLSFV 56
SKP + + T N ST LP L Q++ P+ KL S +S+ +A S SF+
Sbjct: 94 SKPHLSLSTSNPNKASSTLPLPILGLKTKQSLQPSQKLTFSLGESVAGATLVALLSASFI 273
Query: 57 FAPPSLAFE 65
F P+LAF+
Sbjct: 274 FVDPALAFK 300
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.314 0.128 0.329
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,805,315
Number of Sequences: 63676
Number of extensions: 64747
Number of successful extensions: 538
Number of sequences better than 10.0: 75
Number of HSP's better than 10.0 without gapping: 529
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 532
length of query: 209
length of database: 12,639,632
effective HSP length: 93
effective length of query: 116
effective length of database: 6,717,764
effective search space: 779260624
effective search space used: 779260624
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 57 (26.6 bits)
Medicago: description of AC148405.2


