

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148405.1 + phase: 0
(149 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
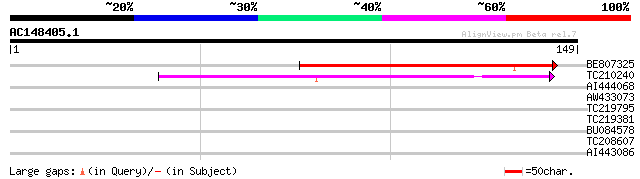
Score E
Sequences producing significant alignments: (bits) Value
BE807325 112 5e-26
TC210240 weakly similar to GB|AAT41846.1|48310596|BT014863 At3g0... 57 4e-09
AI444068 28 1.6
AW433073 27 4.7
TC219795 27 4.7
TC219381 26 8.0
BU084578 26 8.0
TC208607 similar to UP|Q7XIW7 (Q7XIW7) Myosin heavy chain-like, ... 26 8.0
AI443086 homologue to GP|21536945|gb| seven in absentia-like pro... 26 8.0
>BE807325
Length = 456
Score = 112 bits (281), Expect = 5e-26
Identities = 51/69 (73%), Positives = 58/69 (83%), Gaps = 1/69 (1%)
Frame = +3
Query: 77 PEKISWPLVWGQFCLCYEGQKLVTETDYLRDYGIKDGDQLRFIRHVSNVCNFQRK-PKKR 135
P KI WPLVW QFCLCY+GQKLV+E DY+RDYGIKDGDQL F+ HVS+ C+F+RK KKR
Sbjct: 27 PGKILWPLVWRQFCLCYQGQKLVSEMDYIRDYGIKDGDQLHFVYHVSDTCSFERKQSKKR 206
Query: 136 TVNLKQHGR 144
VNLKQH R
Sbjct: 207 VVNLKQHMR 233
>TC210240 weakly similar to GB|AAT41846.1|48310596|BT014863 At3g07860
{Arabidopsis thaliana;} , partial (91%)
Length = 820
Score = 56.6 bits (135), Expect = 4e-09
Identities = 33/110 (30%), Positives = 57/110 (51%), Gaps = 6/110 (5%)
Frame = +3
Query: 40 LTLTVLKLDSSSFQVEVAKTATVAVLKQAVEAAFRHKPEK------ISWPLVWGQFCLCY 93
+ ++VLKLD ++ V V +ATV LK A++ + ISW VW +CL
Sbjct: 285 MRISVLKLDGTTLDVIVMNSATVKDLKLAIKKKVNDMEQSSMGHRHISWKHVWANYCLSC 464
Query: 94 EGQKLVTETDYLRDYGIKDGDQLRFIRHVSNVCNFQRKPKKRTVNLKQHG 143
L+ + + LR++G+++ Q+RF+ +V + R+ KR + HG
Sbjct: 465 HNN*LLDDNEALRNFGVRNNSQVRFVSYV--MTKESRRHSKRRKHRFFHG 608
>AI444068
Length = 321
Score = 28.1 bits (61), Expect = 1.6
Identities = 19/42 (45%), Positives = 25/42 (59%), Gaps = 5/42 (11%)
Frame = +3
Query: 8 SNESPRRSLSIQLSSMIIVDCTLP-----YDKLPSQNLTLTV 44
+ +SPRRS + SS+ I+D YD+LPSQ L LTV
Sbjct: 201 NTKSPRRSSTS--SSLAIIDGIFARKSFFYDRLPSQPLRLTV 320
>AW433073
Length = 431
Score = 26.6 bits (57), Expect = 4.7
Identities = 12/33 (36%), Positives = 18/33 (54%)
Frame = -2
Query: 91 LCYEGQKLVTETDYLRDYGIKDGDQLRFIRHVS 123
LC++G+ VT + + I DGD LR + S
Sbjct: 223 LCFDGENPVTAAIFSSELIISDGDSLREVTRSS 125
>TC219795
Length = 695
Score = 26.6 bits (57), Expect = 4.7
Identities = 16/55 (29%), Positives = 26/55 (47%)
Frame = -2
Query: 56 VAKTATVAVLKQAVEAAFRHKPEKISWPLVWGQFCLCYEGQKLVTETDYLRDYGI 110
+A + +L QA+ A +K K SW + G + QKL +++ DY I
Sbjct: 679 IANAKFLHLLAQALTTAS*NKELKFSWQ*LTGDKLMSTRTQKLHYAIEFIMDYHI 515
>TC219381
Length = 1169
Score = 25.8 bits (55), Expect = 8.0
Identities = 14/37 (37%), Positives = 20/37 (53%)
Frame = -1
Query: 18 IQLSSMIIVDCTLPYDKLPSQNLTLTVLKLDSSSFQV 54
IQ S +++ LP + LPS NLT L L S ++
Sbjct: 1085 IQHSGSSLINFLLPSEPLPSSNLTPASLCLQLGSCEI 975
>BU084578
Length = 420
Score = 25.8 bits (55), Expect = 8.0
Identities = 12/23 (52%), Positives = 16/23 (69%), Gaps = 1/23 (4%)
Frame = -1
Query: 127 NFQRKPKKRTV-NLKQHGRYEFS 148
N Q++ KKR + N+K HG Y FS
Sbjct: 375 NTQKRTKKRHIKNIKSHGGYLFS 307
>TC208607 similar to UP|Q7XIW7 (Q7XIW7) Myosin heavy chain-like, partial (8%)
Length = 1625
Score = 25.8 bits (55), Expect = 8.0
Identities = 13/42 (30%), Positives = 25/42 (58%)
Frame = +3
Query: 31 PYDKLPSQNLTLTVLKLDSSSFQVEVAKTATVAVLKQAVEAA 72
P ++P ++ T+ K+ S FQ++ K + + +LK+ EAA
Sbjct: 714 PKAQIPKES---TLKKVIDSKFQIDETKISKLTILKKLEEAA 830
>AI443086 homologue to GP|21536945|gb| seven in absentia-like protein
{Arabidopsis thaliana}, partial (13%)
Length = 322
Score = 25.8 bits (55), Expect = 8.0
Identities = 9/19 (47%), Positives = 13/19 (68%)
Frame = +2
Query: 80 ISWPLVWGQFCLCYEGQKL 98
+S+PL W FCL +E +L
Sbjct: 263 VSFPLFWSDFCLHFEAFQL 319
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.133 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,675,297
Number of Sequences: 63676
Number of extensions: 84280
Number of successful extensions: 559
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 556
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 557
length of query: 149
length of database: 12,639,632
effective HSP length: 89
effective length of query: 60
effective length of database: 6,972,468
effective search space: 418348080
effective search space used: 418348080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 54 (25.4 bits)
Medicago: description of AC148405.1


