

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148404.10 + phase: 0
(400 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
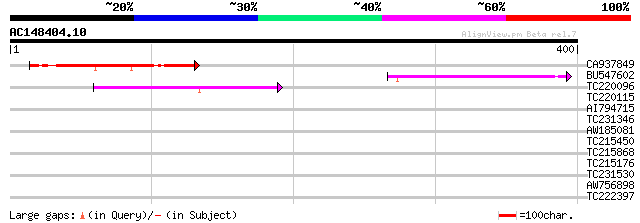
Score E
Sequences producing significant alignments: (bits) Value
CA937849 109 2e-24
BU547602 69 3e-12
TC220096 55 6e-08
TC220115 similar to UP|Q9LQX1 (Q9LQX1) T24P13.20, partial (10%) 29 3.3
AI794715 homologue to GP|1589778|gb| SPINDLY {Arabidopsis thalia... 29 4.3
TC231346 weakly similar to UP|Q6QM04 (Q6QM04) LRR protein WM1.2,... 28 5.6
AW185081 28 7.3
TC215450 similar to UP|P93488 (P93488) NAP1Ps, partial (30%) 28 7.3
TC215868 similar to UP|Q9LJ56 (Q9LJ56) Gb|AAD43149.1, partial (50%) 28 7.3
TC215176 similar to GB|BAC42264.1|26450296|AK117608 GPI-anchored... 28 7.3
TC231530 homologue to UP|C7DA_SOYBN (O48923) Cytochrome P450 71D... 28 7.3
AW756898 similar to GP|21595571|gb| thylakoid lumenal 16.5 kDa p... 28 9.6
TC222397 28 9.6
>CA937849
Length = 432
Score = 109 bits (273), Expect = 2e-24
Identities = 68/126 (53%), Positives = 85/126 (66%), Gaps = 6/126 (4%)
Frame = +1
Query: 15 FIIFFLQLFTAFSNASSSSSSTHSNITHIFQDILKAISSRQKWDL---NDVRVFNFDVAK 71
F IF LQ F AF+ SSSTHSN+THI QD+LKA+S++QKWD +DVRV FDV K
Sbjct: 82 FFIFLLQ-FIAFA-----SSSTHSNLTHILQDVLKAVSAKQKWDSSNNDDVRVTKFDVGK 243
Query: 72 IRFGTSQNYLFRI--GSSKN-NFTVKFSDEISSWNHNKFTTTPKPDLASLVDQLSSIAFL 128
+ FGTS +Y FRI G+ N NFT+KF D++++W NKF TP DL LV +L S L
Sbjct: 244 VMFGTSLSYEFRIRFGTDNNDNFTLKFVDQVATW--NKF-RTPFTDLPPLVHRLGSFPLL 414
Query: 129 DYIKLE 134
+KLE
Sbjct: 415 HTLKLE 432
>BU547602
Length = 600
Score = 69.3 bits (168), Expect = 3e-12
Identities = 47/133 (35%), Positives = 68/133 (50%), Gaps = 3/133 (2%)
Frame = -3
Query: 267 PVASLN---LRLILLEKILRSLLGHKILQDRLSGLIKANIKAYAGVKFPLELERDVGNNA 323
P+ASLN L+ EK+L S+LG K + L+KA++ A VK + E+ V
Sbjct: 499 PLASLNGSNANLLGFEKLLHSVLGDKAEKKGSFRLLKADVSAQTYVKIGFKAEKKVKEGE 320
Query: 324 TLSTLPDWRTRPSVERVWFEVMARVEDSRLKPLSIKKVKPFIESDSVSWANLMSNLSYTK 383
P WRT+P FEV+A+V+ ++ P + VKP I DSV+ L N S +K
Sbjct: 319 LEGYYPAWRTKPETVTTHFEVLAKVDGDKVVPEKVMPVKPVIAVDSVAENLLTRNASMSK 140
Query: 384 LRPVLLPPEALTL 396
+ V PP+ L
Sbjct: 139 TQ-VHPPPDPFYL 104
>TC220096
Length = 549
Score = 55.1 bits (131), Expect = 6e-08
Identities = 36/136 (26%), Positives = 60/136 (43%), Gaps = 3/136 (2%)
Frame = +2
Query: 60 NDVRVFNFDVAKIRFGTSQNYLFRIGSSKNNFTVKFSDEISSWNHNKFTTTPKPDLASLV 119
+DV + FD G S Y F + K +++ W++ + LV
Sbjct: 137 DDVTITGFDPRDAEVGHSVEYQFDLEIDHKVIPFKLLEDVKHWDYVDLPIFRAEAQSGLV 316
Query: 120 DQLSSIAFLDYIK---LEGPFELRVHESHHLSLSLPMNVSYNGLKHIIVGKGITVEVRRA 176
+ +S L + L GP EL +H+++ + LSLP +V LK +++ +G V V+ A
Sbjct: 317 PKRASADGLPVLAPFVLAGPMELWIHDANDMRLSLPHDVDAGVLKKVVLAEGAAVTVKGA 496
Query: 177 REISFYYQSDLDLQRN 192
R +S D L N
Sbjct: 497 RSVSLQQPLDFPLPLN 544
>TC220115 similar to UP|Q9LQX1 (Q9LQX1) T24P13.20, partial (10%)
Length = 623
Score = 29.3 bits (64), Expect = 3.3
Identities = 16/51 (31%), Positives = 25/51 (48%), Gaps = 1/51 (1%)
Frame = -3
Query: 17 IFFLQLFTAF-SNASSSSSSTHSNITHIFQDILKAISSRQKWDLNDVRVFN 66
+FFL +F + SS S H NI H+++ + A WDL + + N
Sbjct: 417 LFFLFIFCKL*PDQSSIISYYHHNILHVYRPEVSATKRASPWDLGNGKPHN 265
>AI794715 homologue to GP|1589778|gb| SPINDLY {Arabidopsis thaliana}, partial
(13%)
Length = 377
Score = 28.9 bits (63), Expect = 4.3
Identities = 21/58 (36%), Positives = 30/58 (51%), Gaps = 2/58 (3%)
Frame = +3
Query: 335 PSVERVWFEVMARVEDSRLKPLSIKKVKPFIESDSV--SWANLMSNLSYTKLRPVLLP 390
P V +VW ++ + +SRL + K KPF SDSV + + + L LR LLP
Sbjct: 147 PKVLQVWARILCAIPNSRL----VVKCKPFC-SDSVRQRFLSTLEQLGLEPLRVDLLP 305
>TC231346 weakly similar to UP|Q6QM04 (Q6QM04) LRR protein WM1.2, partial (14%)
Length = 1790
Score = 28.5 bits (62), Expect = 5.6
Identities = 14/39 (35%), Positives = 22/39 (55%)
Frame = +1
Query: 293 DRLSGLIKANIKAYAGVKFPLELERDVGNNATLSTLPDW 331
+R SG I + ++ + +KF D+GNN T+PDW
Sbjct: 1021 NRFSGYIPSTLQNCSTMKFI-----DMGNNQLSDTIPDW 1122
>AW185081
Length = 366
Score = 28.1 bits (61), Expect = 7.3
Identities = 11/27 (40%), Positives = 18/27 (65%)
Frame = +2
Query: 138 ELRVHESHHLSLSLPMNVSYNGLKHII 164
+L +H+ HHL +LP+ + N LK I+
Sbjct: 239 QLAIHQCHHLEGNLPVPSALNNLKPIL 319
>TC215450 similar to UP|P93488 (P93488) NAP1Ps, partial (30%)
Length = 672
Score = 28.1 bits (61), Expect = 7.3
Identities = 18/45 (40%), Positives = 27/45 (60%)
Frame = -3
Query: 12 LPFFIIFFLQLFTAFSNASSSSSSTHSNITHIFQDILKAISSRQK 56
LP F FFL L ++ S++SSSSSS+ S+ + I + SS +
Sbjct: 334 LPLF--FFLVLLSSSSSSSSSSSSSSSSSSSSSSSISSSSSSSSR 206
>TC215868 similar to UP|Q9LJ56 (Q9LJ56) Gb|AAD43149.1, partial (50%)
Length = 1472
Score = 28.1 bits (61), Expect = 7.3
Identities = 15/41 (36%), Positives = 23/41 (55%)
Frame = -1
Query: 3 TMISLRLHQLPFFIIFFLQLFTAFSNASSSSSSTHSNITHI 43
T S + P F + FL S++SSSSSS+ S++T +
Sbjct: 500 TTSSPNVFDRPLFFLAFLACLGLSSSSSSSSSSSSSSVTSL 378
>TC215176 similar to GB|BAC42264.1|26450296|AK117608 GPI-anchored protein
{Arabidopsis thaliana;} , partial (79%)
Length = 1570
Score = 28.1 bits (61), Expect = 7.3
Identities = 16/32 (50%), Positives = 21/32 (65%), Gaps = 2/32 (6%)
Frame = +3
Query: 10 HQLPFFIIFFLQLFT--AFSNASSSSSSTHSN 39
H L F++F L L + AFS+ SSSSS +SN
Sbjct: 132 HFLTLFLLFLLPLHSHSAFSHNSSSSSQINSN 227
>TC231530 homologue to UP|C7DA_SOYBN (O48923) Cytochrome P450 71D10 ,
complete
Length = 1661
Score = 28.1 bits (61), Expect = 7.3
Identities = 18/71 (25%), Positives = 32/71 (44%), Gaps = 4/71 (5%)
Frame = +1
Query: 146 HLSLSLPMNVSYNGLKHIIVGKGITVEVRRAREISFYYQSDLDLQR----NGSVICSNQK 201
HL L N+ I+ + E+ + +++F + D L R NGS I +Q
Sbjct: 226 HLKLGEVSNI-------IVTSPEMAQEIMKTHDLNFSDRPDFVLSRIVSYNGSGIVFSQH 384
Query: 202 NEFWPFLQSMC 212
++W L+ +C
Sbjct: 385 GDYWRQLRKIC 417
>AW756898 similar to GP|21595571|gb| thylakoid lumenal 16.5 kDa protein
chloroplast precursor {Arabidopsis thaliana}, partial
(31%)
Length = 319
Score = 27.7 bits (60), Expect = 9.6
Identities = 14/20 (70%), Positives = 18/20 (90%)
Frame = -3
Query: 17 IFFLQLFTAFSNASSSSSST 36
+FFL LFT FSN+SSSSS++
Sbjct: 179 LFFLSLFT-FSNSSSSSSAS 123
>TC222397
Length = 505
Score = 27.7 bits (60), Expect = 9.6
Identities = 14/59 (23%), Positives = 24/59 (39%)
Frame = +3
Query: 85 GSSKNNFTVKFSDEISSWNHNKFTTTPKPDLASLVDQLSSIAFLDYIKLEGPFELRVHE 143
G + +F DE W H + + ++ L S+ L+YIK + +HE
Sbjct: 54 GKQREDFKTTTIDENKKWMHRETEKQRRQEMTRLCTNFRSLLPLEYIKGKRSISDHMHE 230
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.322 0.137 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,666,234
Number of Sequences: 63676
Number of extensions: 264881
Number of successful extensions: 1723
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 1668
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1701
length of query: 400
length of database: 12,639,632
effective HSP length: 99
effective length of query: 301
effective length of database: 6,335,708
effective search space: 1907048108
effective search space used: 1907048108
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC148404.10


