

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148348.6 + phase: 0
(552 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
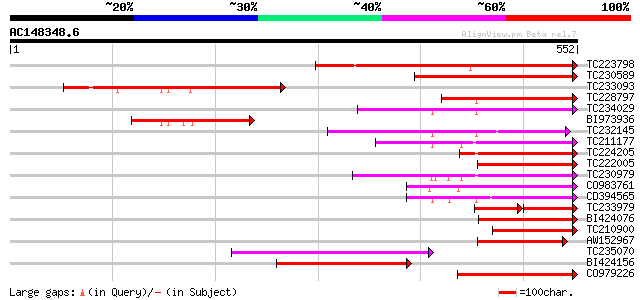
Sequences producing significant alignments: (bits) Value
TC223798 similar to UP|Q9ZQE5 (Q9ZQE5) Expressed protein (At2g15... 342 2e-94
TC230589 similar to UP|Q9ZQE5 (Q9ZQE5) Expressed protein (At2g15... 311 4e-85
TC233093 weakly similar to UP|Q9ZQE5 (Q9ZQE5) Expressed protein ... 237 9e-63
TC228797 177 1e-44
TC234029 similar to UP|Q9LN01 (Q9LN01) T6D22.15, partial (32%) 166 3e-41
BI973936 homologue to GP|4742001|gb| serine-threonine kinase 9 {... 160 1e-39
TC232145 159 2e-39
TC211177 weakly similar to UP|Q9LW63 (Q9LW63) Emb|CAB37460.1, pa... 156 3e-38
TC224205 148 7e-36
TC222005 weakly similar to UP|Q9FIB2 (Q9FIB2) Selenium-binding p... 146 3e-35
TC230979 weakly similar to UP|Q6Z8F8 (Q6Z8F8) Selenium-binding p... 145 5e-35
CO983761 137 1e-32
CD394565 137 2e-32
TC233979 90 1e-30
BI424076 weakly similar to PIR|D96516|D965 F16N3.14 [imported] -... 121 9e-28
TC210900 121 9e-28
AW152967 117 1e-26
TC235070 weakly similar to UP|Q9FJY7 (Q9FJY7) Selenium-binding p... 116 3e-26
BI424156 112 4e-25
CO979226 109 3e-24
>TC223798 similar to UP|Q9ZQE5 (Q9ZQE5) Expressed protein
(At2g15690/F9O13.24), partial (16%)
Length = 789
Score = 342 bits (878), Expect = 2e-94
Identities = 165/258 (63%), Positives = 210/258 (80%), Gaps = 3/258 (1%)
Frame = +3
Query: 298 QMNELGLEITSETMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQS 357
QM + G+ ET VL+AC AEAVE+ +++ ESMK ++GI P +EHY+ ++++LG +
Sbjct: 3 QMKQAGVPPDGETFELVLAACAQAEAVEEGFLHFESMK-EHGIVPSMEHYLEVINILGNA 179
Query: 358 GYLKEAEEFIEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDPSKAVANKIPT 417
G L EAEEFIE++P E V +E+L+N+A+ HGD+DLEDH EE++ LDPSKAVA+K+P
Sbjct: 180 GQLNEAEEFIEKIPIELGVEAWESLRNFAQKHGDLDLEDHAEEVLTCLDPSKAVADKLPP 359
Query: 418 PPPKKYTAISMLDGKNRIIEYKNPTLYKDD--EKLIAMNS-MKDAGYVPDTRYVLHDIDQ 474
PP KK + ++ML+ KNR+ EY+ YK++ EKL ++ M++AGYVPDTRYVLHDID+
Sbjct: 360 PPRKKQSDVNMLEEKNRVTEYRYSIPYKEEAHEKLGGLSGQMREAGYVPDTRYVLHDIDE 539
Query: 475 EAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDCHNAIKIMSRIVGRELIVRD 534
+ KE+AL YHSERLAIAYGLISTPPRT LRIIKNLR+CGDCHNAIKIMS+IVGRELIVRD
Sbjct: 540 DDKEKALQYHSERLAIAYGLISTPPRTTLRIIKNLRICGDCHNAIKIMSKIVGRELIVRD 719
Query: 535 NKRFHHFKDGKCSCGDYW 552
NKRFHHFKDGKCSCGDYW
Sbjct: 720 NKRFHHFKDGKCSCGDYW 773
>TC230589 similar to UP|Q9ZQE5 (Q9ZQE5) Expressed protein
(At2g15690/F9O13.24), partial (21%)
Length = 644
Score = 311 bits (798), Expect = 4e-85
Identities = 148/158 (93%), Positives = 154/158 (96%)
Frame = +1
Query: 395 EDHVEELIVSLDPSKAVANKIPTPPPKKYTAISMLDGKNRIIEYKNPTLYKDDEKLIAMN 454
ED+ EELIVSLDPSKAVANKIPTPPPKKYTAI+MLDG+NRIIEYKNPTLYKDDEKL A++
Sbjct: 1 EDYTEELIVSLDPSKAVANKIPTPPPKKYTAINMLDGRNRIIEYKNPTLYKDDEKLKALS 180
Query: 455 SMKDAGYVPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGD 514
MK+ GYVPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGD
Sbjct: 181 GMKETGYVPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGD 360
Query: 515 CHNAIKIMSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 552
CHNAIKIMSRIVGRELIVRDNKRFHHFKDGKCSCGDYW
Sbjct: 361 CHNAIKIMSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 474
>TC233093 weakly similar to UP|Q9ZQE5 (Q9ZQE5) Expressed protein
(At2g15690/F9O13.24), partial (16%)
Length = 712
Score = 237 bits (605), Expect = 9e-63
Identities = 128/239 (53%), Positives = 151/239 (62%), Gaps = 23/239 (9%)
Frame = +1
Query: 53 QHFNHHNTQNAPGRFPQQDWNPQNPNFRPPQPPQNPNYRQQPPPQNPNFRP---PPPPQN 109
Q + + P F Q +P NP PP Q P++ P Q+ RP N
Sbjct: 4 QQWQQQHQHQQPQHFNPQTHHPPNP---PPPSNQFPHHHNHQPRQHNVPRPFLEHGGNHN 174
Query: 110 PNFRP-PPPPQNPNFRQPIRQNPNFQPPNGPNRGINQN-----QWNPQNG---NLNQFQN 160
+ P PPPPQN NF+ P QNPNF+PP PNR NQN QWNP++ N NQFQN
Sbjct: 175 NQWNPQPPPPQNHNFQTPTSQNPNFRPPTTPNRWNNQNPAAPNQWNPRSQGFPNSNQFQN 354
Query: 161 PNNQFQTPNVQEQA-----------PPPPSIVDLTRFCQEGKVKEALELMEKGIKADANC 209
PNNQ QA P PPSI DLTR C+EGKVKEA+ELM+KG+KADA C
Sbjct: 355 PNNQLNNNQASIQAQTHPAPPPPPHPLPPSITDLTRLCKEGKVKEAIELMDKGVKADAGC 534
Query: 210 FEILFDLCGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHM 268
F++LFDLCG+SKS+EDAKK HD+FLQSTFRSD ++NKVIEMYGNCKSMTDA VF+HM
Sbjct: 535 FDLLFDLCGQSKSLEDAKKAHDHFLQSTFRSDLTLNNKVIEMYGNCKSMTDAHXVFNHM 711
>TC228797
Length = 900
Score = 177 bits (448), Expect = 1e-44
Identities = 81/138 (58%), Positives = 111/138 (79%), Gaps = 6/138 (4%)
Frame = +1
Query: 421 KKYTAISMLDGKNRIIEYK-NPTLYKDDEKLIAM-----NSMKDAGYVPDTRYVLHDIDQ 474
K + ++L+ ++R+ EY+ T + +++K+ A+ + MK+AGYVP+T++VLHDIDQ
Sbjct: 103 KNLASKNLLEVRSRVREYRAGDTSHPENDKIYALLRGLKSQMKEAGYVPETKFVLHDIDQ 282
Query: 475 EAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDCHNAIKIMSRIVGRELIVRD 534
E KE+ALL HSERLAIAYGL+++P R P+R+IKNLRVCGDCH A+KI+S++VGRELI+RD
Sbjct: 283 EGKEEALLAHSERLAIAYGLLNSPARAPMRVIKNLRVCGDCHTALKIISKLVGRELIIRD 462
Query: 535 NKRFHHFKDGKCSCGDYW 552
KRFHHF DG CSC DYW
Sbjct: 463 AKRFHHFNDGLCSCRDYW 516
>TC234029 similar to UP|Q9LN01 (Q9LN01) T6D22.15, partial (32%)
Length = 732
Score = 166 bits (419), Expect = 3e-41
Identities = 89/241 (36%), Positives = 134/241 (54%), Gaps = 27/241 (11%)
Frame = +2
Query: 339 GIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPTVTVFETLKNYARIHGDVDLEDHV 398
GI P ++HY ++D+L +SG EA+ + + EP ++ +L N RIHG V+ ++V
Sbjct: 2 GISPKLQHYGCMIDLLARSGKFDEAKVLMGNMEMEPDGAIWGSLLNACRIHGQVEFGEYV 181
Query: 399 EELIVSLDPSKA---------------------VANKIPTPPPKKYTAISMLDGKNRIIE 437
E + L+P + + K+ KK + ++ + E
Sbjct: 182 AERLFELEPENSGAYVLLSNIYAGAGRWDDVAKIRTKLNDKGMKKVPGCTSIEIDGVVHE 361
Query: 438 YK-NPTLYKDDEKLIAM-----NSMKDAGYVPDTRYVLHDIDQEAKEQALLYHSERLAIA 491
+ + E + M +++ G+VPDT VL+D+D+E KE AL HSE+LAIA
Sbjct: 362 FLVGDKFHPQSENIFRMLDEVDRLLEETGFVPDTSEVLYDMDEEWKEGALTQHSEKLAIA 541
Query: 492 YGLISTPPRTPLRIIKNLRVCGDCHNAIKIMSRIVGRELIVRDNKRFHHFKDGKCSCGDY 551
+GLIST P + +RI+KNLRVC +CH+A K++S+I RE+I RD RFHHFKDG CSC D
Sbjct: 542 FGLISTKPGSTIRIVKNLRVCRNCHSATKLISKIFNREIIARDRNRFHHFKDGFCSCNDC 721
Query: 552 W 552
W
Sbjct: 722 W 724
>BI973936 homologue to GP|4742001|gb| serine-threonine kinase 9 {Takifugu
rubripes}, partial (1%)
Length = 405
Score = 160 bits (405), Expect = 1e-39
Identities = 84/135 (62%), Positives = 97/135 (71%), Gaps = 15/135 (11%)
Frame = +1
Query: 119 QNPNFRQPIRQNPNFQPPNGPNRGINQN-----QWNPQNG---NLNQFQNPNNQFQT--P 168
QN NF+ P QNPNF+ P PNR NQN QWNP++ N NQFQNPNNQ
Sbjct: 1 QNLNFQTPTSQNPNFRIPTTPNRWNNQNLATPNQWNPRSQGFPNPNQFQNPNNQLNNNQT 180
Query: 169 NVQEQAPP-----PPSIVDLTRFCQEGKVKEALELMEKGIKADANCFEILFDLCGKSKSV 223
+VQ QA P PPSI DLTR C+EGKVKEA+ELM+KG+KADA CF +LFD CG+SKS+
Sbjct: 181 SVQSQASPAPPPQPPSITDLTRLCREGKVKEAIELMDKGVKADAGCFALLFDSCGQSKSL 360
Query: 224 EDAKKVHDYFLQSTF 238
EDAKK HD+FLQSTF
Sbjct: 361 EDAKKAHDHFLQSTF 405
Score = 35.0 bits (79), Expect = 0.087
Identities = 25/74 (33%), Positives = 30/74 (39%), Gaps = 8/74 (10%)
Frame = +1
Query: 46 QPPSDHNQHF------NHHNTQNAPGRFPQQDWNPQNPNFRPPQPPQNPN--YRQQPPPQ 97
Q P+ N +F N N QN P Q WNP++ F P QNPN
Sbjct: 16 QTPTSQNPNFRIPTTPNRWNNQNLAT--PNQ-WNPRSQGFPNPNQFQNPNNQLNNNQTSV 186
Query: 98 NPNFRPPPPPQNPN 111
P PPPQ P+
Sbjct: 187 QSQASPAPPPQPPS 228
>TC232145
Length = 817
Score = 159 bits (403), Expect = 2e-39
Identities = 92/264 (34%), Positives = 141/264 (52%), Gaps = 27/264 (10%)
Frame = +3
Query: 310 TMLAVLSACGSAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQ 369
T ++VLSAC ++D + Y SM + + I+PG+EHY ++D+LG+ G L+EA FIE
Sbjct: 27 TFVSVLSACSHTGKIDDGFKYFNSMANVHNIKPGLEHYACMVDLLGRVGRLEEACRFIES 206
Query: 370 LPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDPSKA------------------- 410
+PFEP V+ L H +V++ V E + L+P
Sbjct: 207 MPFEPDSLVWGALLGACGKHANVEMGREVAERLFKLEPDNPGNYMLLSNIYIRHGMLEEA 386
Query: 411 --VANKIPTPPPKKYTAISMLDGKNRIIEYK-NPTLYKDDEKLIAM-----NSMKDAGYV 462
V + +K + S +D KNR + N + +++ M +K GYV
Sbjct: 387 DEVRRLMGINGVRKESGCSWIDVKNRTFVFNANDRSHSRTQEIYGMLQKLKELIKRRGYV 566
Query: 463 PDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDCHNAIKIM 522
+T++ + ++ ++EQ+L HSE+LA+A+GL+ PP +P+RI KNLR CGDCH +K
Sbjct: 567 AETQFATNSVEG-SEEQSLWCHSEKLALAFGLLVLPPGSPVRIKKNLRTCGDCHTVMKFA 743
Query: 523 SRIVGRELIVRDNKRFHHFKDGKC 546
S I RE+IVRD RFH F +G C
Sbjct: 744 SEIFQREIIVRDINRFHRFTNGSC 815
>TC211177 weakly similar to UP|Q9LW63 (Q9LW63) Emb|CAB37460.1, partial (17%)
Length = 823
Score = 156 bits (394), Expect = 3e-38
Identities = 90/226 (39%), Positives = 127/226 (55%), Gaps = 30/226 (13%)
Frame = +2
Query: 357 SGYLKEAEEFIEQLPFEPTVTVFETLKNYARIHGDVDLEDHVEELIVSLDPSKA------ 410
+G L+EA +FI++LP +PT + TL + HG+V++ V + I LD S
Sbjct: 2 AGRLEEACKFIDELPIKPTPXXWRTLLSSCSSHGNVEMAKLVIQRIFELDDSHGGDYVIL 181
Query: 411 ---------------VANKIPTPPPKKYTAISMLDGKNRIIEY--------KNPTLYKDD 447
+ + K S ++ N + E+ + L+
Sbjct: 182 SNLCARNGRWDDVNHLRKMMVDKGALKVPGCSSIEVNNVVHEFFSGDGVHSTSTILHHAL 361
Query: 448 EKLIAMNSMKDAGYVPDTRYVLH-DIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRII 506
++L+ +K AGYVPDT V + DI+ E KE L YHSE+LAI YGL++TPP T +R++
Sbjct: 362 DELV--KELKLAGYVPDTSLVFYADIEDEEKEIVLRYHSEKLAITYGLLNTPPGTTIRVV 535
Query: 507 KNLRVCGDCHNAIKIMSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 552
KNLRVC DCHNA K +S I GR++I+RD +RFHH KDGKCSCGDYW
Sbjct: 536 KNLRVCVDCHNAAKFISLIFGRQIILRDVQRFHHLKDGKCSCGDYW 673
>TC224205
Length = 464
Score = 148 bits (373), Expect = 7e-36
Identities = 64/114 (56%), Positives = 88/114 (77%)
Frame = +1
Query: 439 KNPTLYKDDEKLIAMNSMKDAGYVPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTP 498
+ P + K EKL + +++AGY PD +VLHD+D+E K +L YHSE+LA+AYGL+ P
Sbjct: 31 EQPIIMKMLEKLGGL--LREAGYCPDGSFVLHDVDEEEKTHSLGYHSEKLAVAYGLLKVP 204
Query: 499 PRTPLRIIKNLRVCGDCHNAIKIMSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 552
P+R++KNLRVCGDCH+AIK+++++ GRE+I+RD RFHHFKDG CSC DYW
Sbjct: 205 EGMPIRVMKNLRVCGDCHSAIKLIAKVTGREIILRDANRFHHFKDGHCSCKDYW 366
>TC222005 weakly similar to UP|Q9FIB2 (Q9FIB2) Selenium-binding protein-like,
partial (13%)
Length = 757
Score = 146 bits (368), Expect = 3e-35
Identities = 61/97 (62%), Positives = 81/97 (82%)
Frame = +2
Query: 456 MKDAGYVPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDC 515
+K+ GYVPDT +VLHD+++E KEQ L+YHSE+LA+A+G+ISTPP TP+++ KNLR C DC
Sbjct: 341 IKEEGYVPDTNFVLHDVEEEQKEQNLVYHSEKLAVAFGIISTPPGTPIKVFKNLRTCVDC 520
Query: 516 HNAIKIMSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 552
H AIK +S+IV R++ VRD+ RFH F+DG CSC DYW
Sbjct: 521 HTAIKYISKIVQRKITVRDSNRFHCFEDGSCSCKDYW 631
>TC230979 weakly similar to UP|Q6Z8F8 (Q6Z8F8) Selenium-binding protein-like,
partial (61%)
Length = 837
Score = 145 bits (366), Expect = 5e-35
Identities = 82/248 (33%), Positives = 137/248 (55%), Gaps = 29/248 (11%)
Frame = +2
Query: 334 MKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPTVTVFETLKNYARIHGDVD 393
M+ ++ I+P ++HY ++D++G++G LKEA + I+ +P +P V+ +L + ++H +++
Sbjct: 14 MQFEHMIKPTIQHYGCMVDLMGRAGMLKEAYDLIKSMPIKPNDVVWRSLLSACKVHHNLE 193
Query: 394 LEDHVEELIVSLDPSK-----AVAN------------KIPTPPPKKYTA----ISMLDGK 432
+ + + I L+ +AN +I T +K S+++
Sbjct: 194 IGEIAADNIFKLNKHNPGDYLVLANMYARAQKWANVARIRTEMVEKNLVQTPGFSLVEAN 373
Query: 433 NRIIEY--------KNPTLYKDDEKLIAMNSMKDAGYVPDTRYVLHDIDQEAKEQALLYH 484
+ ++ K T+Y +++ +K GY PD VL D+D++ K Q L +H
Sbjct: 374 RNVYKFVSQDKSQPKCETIYDMIQQMEWQ--LKFEGYTPDMSQVLLDVDEDEKRQRLKHH 547
Query: 485 SERLAIAYGLISTPPRTPLRIIKNLRVCGDCHNAIKIMSRIVGRELIVRDNKRFHHFKDG 544
S++LAIA+ LI T +P+RI +NLR+C DCH K +S I RE+ VRD RFHHFKDG
Sbjct: 548 SQKLAIAFALIQTSEGSPIRISRNLRMCNDCHTYTKFISVIYEREITVRDRNRFHHFKDG 727
Query: 545 KCSCGDYW 552
CSC DYW
Sbjct: 728 TCSCKDYW 751
>CO983761
Length = 775
Score = 137 bits (345), Expect = 1e-32
Identities = 74/193 (38%), Positives = 108/193 (55%), Gaps = 27/193 (13%)
Frame = -2
Query: 387 RIHGDVDLEDHVEELIVSLDP---------------------SKAVANKIPTPPPKKYTA 425
RIH +V+L + L+ L+P +K+V + +K
Sbjct: 768 RIHCNVELAERASTLLFELEPRNAGNYVLLADIYAEAKMWSEAKSVMKLLEARGLQKLPG 589
Query: 426 ISMLDGKNRI-----IEYKNPTLYKDDEKLIAM-NSMKDAGYVPDTRYVLHDIDQEAKEQ 479
S ++ K ++ ++ NP + + L+ + N MK GYVP T VL+D+D+E KE+
Sbjct: 588 CSWIEVKRKVYSFVSVDEHNPQIEEIHALLVKLSNEMKAQGYVPQTNVVLYDLDEEEKER 409
Query: 480 ALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDCHNAIKIMSRIVGRELIVRDNKRFH 539
+L HSE+LA+A+GLI+T +RI KNLR+C DCH K +S+ RE++VRD RFH
Sbjct: 408 IVLGHSEKLAVAFGLINTVKGETIRIRKNLRLCEDCHAVTKFISKFANREILVRDVNRFH 229
Query: 540 HFKDGKCSCGDYW 552
HFKDG CSCGDYW
Sbjct: 228 HFKDGVCSCGDYW 190
>CD394565
Length = 682
Score = 137 bits (344), Expect = 2e-32
Identities = 78/193 (40%), Positives = 111/193 (57%), Gaps = 27/193 (13%)
Frame = -3
Query: 387 RIHGDVDLEDHVEELIVSLDPSKA----------VANKIPTPPPKKYTAI---------- 426
+IH +V+L + + LDP + +N + K TA+
Sbjct: 677 KIHKNVELGEKAAHKLFKLDPDEGGYHVLLANIYASNSMWDKVAKVRTAMEDKGLHKTPG 498
Query: 427 -SMLDGKNRIIE-YKNPTLYKDDEKLIAM-----NSMKDAGYVPDTRYVLHDIDQEAKEQ 479
S ++ +N I Y T + + +K+ A + +K AGYVPD + HD++++ K+Q
Sbjct: 497 CSWVELRNEIHTFYSGSTNHPESKKIYAFLETLGDEIKAAGYVPDPDSI-HDVEEDVKKQ 321
Query: 480 ALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDCHNAIKIMSRIVGRELIVRDNKRFH 539
L HSERLAIA+GL++T P T L I KNLRVCGDCH+ K +S + GRE+IVRD +RFH
Sbjct: 320 LLSSHSERLAIAFGLLNTSPGTTLHIRKNLRVCGDCHDTTKYISLVTGREIIVRDLRRFH 141
Query: 540 HFKDGKCSCGDYW 552
HFK+G CSCGDYW
Sbjct: 140 HFKNGSCSCGDYW 102
>TC233979
Length = 825
Score = 90.1 bits (222), Expect(2) = 1e-30
Identities = 36/52 (69%), Positives = 44/52 (84%)
Frame = +1
Query: 501 TPLRIIKNLRVCGDCHNAIKIMSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 552
T LR+IKNLRVCGDCHNAIK +S++ RE+++RD RFHHF+ G CSCGDYW
Sbjct: 394 TTLRVIKNLRVCGDCHNAIKYISKVFEREVVLRDANRFHHFRSGVCSCGDYW 549
Score = 61.6 bits (148), Expect(2) = 1e-30
Identities = 26/47 (55%), Positives = 37/47 (78%)
Frame = +3
Query: 453 MNSMKDAGYVPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPP 499
M +++ GY+PDT + L D+++E KE +L YHSE+LAIAYGL+ TPP
Sbjct: 249 MKRIREEGYLPDTDFALVDVEEEDKECSLYYHSEKLAIAYGLMKTPP 389
>BI424076 weakly similar to PIR|D96516|D965 F16N3.14 [imported] - Arabidopsis
thaliana, partial (16%)
Length = 427
Score = 121 bits (303), Expect = 9e-28
Identities = 52/96 (54%), Positives = 66/96 (68%)
Frame = +1
Query: 457 KDAGYVPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDCH 516
K GY+ T++V H++ +E K Q L HSERLA+ YGL+ TP T +RI KNLR+C DCH
Sbjct: 58 KKGGYIAQTKFVFHNVSEEEKTQMLYRHSERLALGYGLLVTPKGTSIRITKNLRICDDCH 237
Query: 517 NAIKIMSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 552
KI S + R L+VRD RFHHF+ G CSCGD+W
Sbjct: 238 TFFKIASEVSQRALVVRDANRFHHFERGLCSCGDFW 345
>TC210900
Length = 654
Score = 121 bits (303), Expect = 9e-28
Identities = 51/82 (62%), Positives = 62/82 (75%)
Frame = +3
Query: 471 DIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDCHNAIKIMSRIVGREL 530
D++ E K L HSE++A+AY LISTP P+R+ KNLRVCGDCH AIK +S + GRE+
Sbjct: 3 DVEDEQKXXYLFQHSEKIAVAYALISTPKPKPIRVFKNLRVCGDCHTAIKYISIVTGREI 182
Query: 531 IVRDNKRFHHFKDGKCSCGDYW 552
+VRD RFHH KDGKCSC DYW
Sbjct: 183 VVRDANRFHHIKDGKCSCNDYW 248
>AW152967
Length = 323
Score = 117 bits (294), Expect = 1e-26
Identities = 51/88 (57%), Positives = 69/88 (77%)
Frame = +2
Query: 456 MKDAGYVPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLISTPPRTPLRIIKNLRVCGDC 515
MK AGYVPD L D+++E K + YHS+++A+A+GLI+ P + P+RI KNL +C DC
Sbjct: 44 MKKAGYVPDANLSLFDLEEEEKASKVWYHSKKIALAFGLITLPRKVPIRITKNLXICIDC 223
Query: 516 HNAIKIMSRIVGRELIVRDNKRFHHFKD 543
H+AIK +S+IVGRE+IV+DN RFHHFKD
Sbjct: 224 HSAIKFISKIVGREIIVKDNNRFHHFKD 307
>TC235070 weakly similar to UP|Q9FJY7 (Q9FJY7) Selenium-binding protein-like,
partial (22%)
Length = 875
Score = 116 bits (290), Expect = 3e-26
Identities = 61/196 (31%), Positives = 106/196 (53%)
Frame = +2
Query: 217 CGKSKSVEDAKKVHDYFLQSTFRSDFKMHNKVIEMYGNCKSMTDARRVFDHMPNRNMDSW 276
C S + +H + +++ F+ + + +++ Y C + +A+R F + + N+ +W
Sbjct: 5 CSCLCSFRQGQLLHAHLIKTPFQVNVYVGTALVDFYSKCGHLAEAQRSFISIFSPNVAAW 184
Query: 277 HMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVLSACGSAEAVEDAYIYLESMKS 336
+I GYA +G E + LF M G+ + T + VLSAC A V + SM+
Sbjct: 185 TALINGYAYHGLGSEAILLFRSMLHQGIVPNAATFVGVLSACNHAGLVCEGLRIFHSMQR 364
Query: 337 KYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPTVTVFETLKNYARIHGDVDLED 396
YG+ P +EHY ++D+LG+SG+LKEAEEFI ++P E ++ L N + D+++ +
Sbjct: 365 CYGVTPTIEHYTCVVDLLGRSGHLKEAEEFIIKMPIEADGIIWGALLNASWFWKDMEVGE 544
Query: 397 HVEELIVSLDPSKAVA 412
E + SLDP+ A
Sbjct: 545 RAAEKLFSLDPNPIFA 592
>BI424156
Length = 406
Score = 112 bits (280), Expect = 4e-25
Identities = 49/132 (37%), Positives = 80/132 (60%)
Frame = +3
Query: 260 DARRVFDHMPNRNMDSWHMMIRGYANSTMGDEGLQLFEQMNELGLEITSETMLAVLSACG 319
DA +F++MP RN+ +W MI A + ++ + LF Q LGL+ + + VL AC
Sbjct: 9 DALNIFNNMPERNLTTWDTMITQLAKNGFAEDSIDLFTQFKNLGLKPDGQMFIGVLFACS 188
Query: 320 SAEAVEDAYIYLESMKSKYGIEPGVEHYMGLLDVLGQSGYLKEAEEFIEQLPFEPTVTVF 379
+++ ++ ESM YGI P + H++ ++D++G G+L EA EFIE++P EP+ +
Sbjct: 189 VLGDIDEGMLHFESMSKDYGIVPSMTHFVSVVDMIGSIGHLDEAFEFIERMPMEPSAETW 368
Query: 380 ETLKNYARIHGD 391
ETL N R+HG+
Sbjct: 369 ETLMNLCRVHGN 404
>CO979226
Length = 737
Score = 109 bits (273), Expect = 3e-24
Identities = 51/117 (43%), Positives = 75/117 (63%), Gaps = 1/117 (0%)
Frame = -3
Query: 437 EYKNPTLYKDDEKLIA-MNSMKDAGYVPDTRYVLHDIDQEAKEQALLYHSERLAIAYGLI 495
+Y +P K +KL M +++ G+V +T +LHD+D+E K + HSE++A+A+GL+
Sbjct: 693 DYSHPDSDKIHKKLEEIMRKIQEVGFVSETESILHDVDEEVKTDMINKHSEKIALAFGLL 514
Query: 496 STPPRTPLRIIKNLRVCGDCHNAIKIMSRIVGRELIVRDNKRFHHFKDGKCSCGDYW 552
+ PL IIKNL++ DC N K +S I + +RD+KRFHHF D CSCGDYW
Sbjct: 513 VSNADMPLVIIKNLQISQDCLNTAKFVSLIEKCTITIRDSKRFHHFSDASCSCGDYW 343
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.317 0.136 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 32,008,577
Number of Sequences: 63676
Number of extensions: 798698
Number of successful extensions: 32291
Number of sequences better than 10.0: 1527
Number of HSP's better than 10.0 without gapping: 8042
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 16003
length of query: 552
length of database: 12,639,632
effective HSP length: 102
effective length of query: 450
effective length of database: 6,144,680
effective search space: 2765106000
effective search space used: 2765106000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC148348.6


