

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148347.8 - phase: 0 /pseudo
(97 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
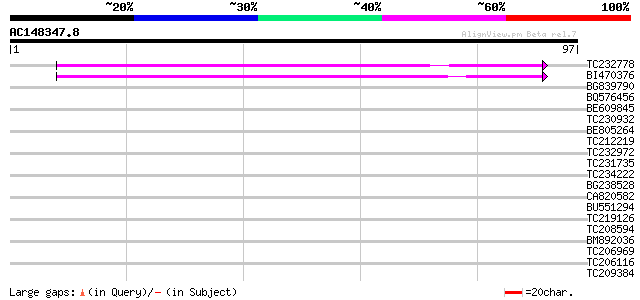
TC232778 similar to UP|Q9LNG5 (Q9LNG5) F21D18.16, partial (5%) 52 5e-08
BI470376 weakly similar to GP|14090338|dbj P0638D12.5 {Oryza sat... 49 4e-07
BG839790 similar to PIR|D96521|D96 protein F21D18.16 [imported] ... 34 0.013
BQ576456 34 0.013
BE609845 weakly similar to GP|3688284|emb lanatoside 15'-O-acety... 31 0.087
TC230932 29 0.43
BE805264 27 1.3
TC212219 27 1.6
TC232972 weakly similar to UP|O82681 (O82681) Lanatoside 15'-O-a... 27 1.6
TC231735 27 1.6
TC234222 27 2.1
BG238528 homologue to GP|6224712|gb| non-phototropic hypocotyl 3... 27 2.1
CA820582 27 2.1
BU551294 26 3.7
TC219126 similar to UP|Q7XYW9 (Q7XYW9) EXO70-G1 protein, partial... 26 3.7
TC208594 weakly similar to GB|AAS56346.1|45269930|AY558020 YNL23... 26 3.7
BM892036 26 3.7
TC206969 26 3.7
TC206116 UP|CC48_SOYBN (P54774) Cell division cycle protein 48 h... 25 4.8
TC209384 homologue to UP|Q71SQ1 (Q71SQ1) MYC1, partial (41%) 25 4.8
>TC232778 similar to UP|Q9LNG5 (Q9LNG5) F21D18.16, partial (5%)
Length = 746
Score = 52.0 bits (123), Expect = 5e-08
Identities = 31/84 (36%), Positives = 42/84 (49%)
Frame = -1
Query: 9 VMRVSGLQWLQHVSYLVIDQGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSIE 68
++R G W+ + YL I+ L+S ERW T +FHL E +TL D+ LL L +
Sbjct: 545 LLRQCGFYWIMKMGYLKINAALISAFIERWRPETHTFHLRCGEATITLQDVLILLGLRTD 366
Query: 69 SHMLYHNIYLTRSEAVDLMVELLG 92
L I T + DL ELLG
Sbjct: 365 GAPL---IGSTNLDWADLCEELLG 303
>BI470376 weakly similar to GP|14090338|dbj P0638D12.5 {Oryza sativa
(japonica cultivar-group)}, partial (5%)
Length = 429
Score = 48.9 bits (115), Expect = 4e-07
Identities = 27/84 (32%), Positives = 46/84 (54%)
Frame = -1
Query: 9 VMRVSGLQWLQHVSYLVIDQGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSIE 68
++++SGL + ++ L ++ L++ ERW T +FHL E +TL D+S LL + ++
Sbjct: 396 LLQISGLYPIMKLAQLKVNGALVNAFIERWRPETHTFHLKYGEATITLQDVSVLLGIPVD 217
Query: 69 SHMLYHNIYLTRSEAVDLMVELLG 92
L N T + +L ELLG
Sbjct: 216 GRPLIGN---TNIDWFELFHELLG 154
>BG839790 similar to PIR|D96521|D96 protein F21D18.16 [imported] -
Arabidopsis thaliana, partial (2%)
Length = 504
Score = 33.9 bits (76), Expect = 0.013
Identities = 14/39 (35%), Positives = 24/39 (60%)
Frame = -1
Query: 9 VMRVSGLQWLQHVSYLVIDQGLLSVIAERWHEVTSSFHL 47
++R G + +SYL I+ L++V+ ERW T +FH+
Sbjct: 261 LLRQCGFY*IMKMSYLKINASLMTVLIERWRPETHTFHM 145
>BQ576456
Length = 424
Score = 33.9 bits (76), Expect = 0.013
Identities = 20/41 (48%), Positives = 28/41 (67%)
Frame = +1
Query: 51 EMIVTLDDLSCLLHLSIESHMLYHNIYLTRSEAVDLMVELL 91
E+ +TLDD++CLLHL I + L+ L EAV L++ELL
Sbjct: 34 ELTITLDDMACLLHLPI-TGALHRFEPLGVDEAVLLLMELL 153
>BE609845 weakly similar to GP|3688284|emb lanatoside 15'-O-acetylesterase
{Digitalis lanata}, partial (27%)
Length = 430
Score = 31.2 bits (69), Expect = 0.087
Identities = 18/47 (38%), Positives = 24/47 (50%), Gaps = 3/47 (6%)
Frame = -3
Query: 31 LSVIAERWHE---VTSSFHLPIWEMIVTLDDLSCLLHLSIESHMLYH 74
L+ + R+H + FHL IW TL D SC +SI + LYH
Sbjct: 416 LNTNSRRYHHR**IAVRFHLYIWYCRKTLMDRSCSHPISIHNTYLYH 276
>TC230932
Length = 673
Score = 28.9 bits (63), Expect = 0.43
Identities = 14/35 (40%), Positives = 20/35 (57%)
Frame = +3
Query: 50 WEMIVTLDDLSCLLHLSIESHMLYHNIYLTRSEAV 84
W M++TL + SCLL+ + +L N L SE V
Sbjct: 558 WVMLITLPEQSCLLNFFLNLPLLR*NQLLQNSEGV 662
>BE805264
Length = 445
Score = 27.3 bits (59), Expect = 1.3
Identities = 22/82 (26%), Positives = 39/82 (46%), Gaps = 14/82 (17%)
Frame = +3
Query: 20 HVSYLVIDQGLLSVIAERWHEVTSSFH-------LPIWEMIVTLDDLSC-------LLHL 65
H+ L++ LL ++ H TSS H LPIW+++ ++ LS LLHL
Sbjct: 165 HLQVLLLQLMLLKIL----HLQTSSVH*RLLEIMLPIWKILDSILQLSLLLLVVCYLLHL 332
Query: 66 SIESHMLYHNIYLTRSEAVDLM 87
E H+ ++ ++ + L+
Sbjct: 333 LCEYHLPLIQLHRCQASPLTLL 398
>TC212219
Length = 630
Score = 26.9 bits (58), Expect = 1.6
Identities = 12/19 (63%), Positives = 14/19 (73%)
Frame = -2
Query: 69 SHMLYHNIYLTRSEAVDLM 87
SHMLY NI L +E VDL+
Sbjct: 584 SHMLYTNISLAATEGVDLV 528
>TC232972 weakly similar to UP|O82681 (O82681) Lanatoside
15'-O-acetylesterase precursor, partial (45%)
Length = 570
Score = 26.9 bits (58), Expect = 1.6
Identities = 13/30 (43%), Positives = 16/30 (53%)
Frame = -1
Query: 45 FHLPIWEMIVTLDDLSCLLHLSIESHMLYH 74
FHL +W L D SC L I + +LYH
Sbjct: 354 FHLSLWYCRKKLLDRSCSHPLRIHNTLLYH 265
>TC231735
Length = 707
Score = 26.9 bits (58), Expect = 1.6
Identities = 17/52 (32%), Positives = 26/52 (49%), Gaps = 3/52 (5%)
Frame = -3
Query: 26 IDQGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLS---IESHMLYH 74
+DQ L + + WH + L + + ++D SC LHL +E H LYH
Sbjct: 657 VDQKLETPV---WHSSQKPWLLKLVGGLPSVDSHSCPLHLH*VVLELHWLYH 511
>TC234222
Length = 456
Score = 26.6 bits (57), Expect = 2.1
Identities = 16/41 (39%), Positives = 19/41 (46%)
Frame = -3
Query: 8 DVMRVSGLQWLQHVSYLVIDQGLLSVIAERWHEVTSSFHLP 48
D+ L+WL H S I LLSV E + E S H P
Sbjct: 133 DLTAYGRLRWLHHDSIESIQGSLLSVRPENFSEH*QSTHRP 11
>BG238528 homologue to GP|6224712|gb| non-phototropic hypocotyl 3
{Arabidopsis thaliana}, partial (13%)
Length = 462
Score = 26.6 bits (57), Expect = 2.1
Identities = 9/24 (37%), Positives = 16/24 (66%)
Frame = -2
Query: 59 LSCLLHLSIESHMLYHNIYLTRSE 82
L+C++H S +H+ H +YL S+
Sbjct: 362 LNCIMHYSSTNHLKPHTLYLDSSD 291
>CA820582
Length = 451
Score = 26.6 bits (57), Expect = 2.1
Identities = 14/35 (40%), Positives = 22/35 (62%), Gaps = 4/35 (11%)
Frame = +1
Query: 7 KDVMRVSGLQWLQHVSYLVI----DQGLLSVIAER 37
K+ MR+S +W+ H SYL + Q ++SVI +R
Sbjct: 16 KEQMRMSLQKWMLHCSYLPLTCQPSQMMISVICKR 120
>BU551294
Length = 469
Score = 25.8 bits (55), Expect = 3.7
Identities = 10/17 (58%), Positives = 13/17 (75%)
Frame = +3
Query: 64 HLSIESHMLYHNIYLTR 80
HLS+ES + YHN +L R
Sbjct: 75 HLSLESILNYHNFFLMR 125
>TC219126 similar to UP|Q7XYW9 (Q7XYW9) EXO70-G1 protein, partial (20%)
Length = 1178
Score = 25.8 bits (55), Expect = 3.7
Identities = 10/26 (38%), Positives = 18/26 (68%)
Frame = -3
Query: 59 LSCLLHLSIESHMLYHNIYLTRSEAV 84
L C ++S ++ + + N++LTRS AV
Sbjct: 219 LVCFAYISSKTSLNFFNLFLTRSRAV 142
>TC208594 weakly similar to GB|AAS56346.1|45269930|AY558020 YNL231C
{Saccharomyces cerevisiae;} , partial (8%)
Length = 1478
Score = 25.8 bits (55), Expect = 3.7
Identities = 12/27 (44%), Positives = 15/27 (55%)
Frame = -3
Query: 29 GLLSVIAERWHEVTSSFHLPIWEMIVT 55
GLL ++E WH T F +W IVT
Sbjct: 207 GLLETLSEWWHGGTDRF--SVWRAIVT 133
>BM892036
Length = 422
Score = 25.8 bits (55), Expect = 3.7
Identities = 11/22 (50%), Positives = 15/22 (68%)
Frame = -1
Query: 6 VKDVMRVSGLQWLQHVSYLVID 27
+K +++V QWL HVS L ID
Sbjct: 164 LKKLIKVHAPQWLSHVS*LTID 99
>TC206969
Length = 726
Score = 25.8 bits (55), Expect = 3.7
Identities = 15/46 (32%), Positives = 24/46 (51%)
Frame = +2
Query: 28 QGLLSVIAERWHEVTSSFHLPIWEMIVTLDDLSCLLHLSIESHMLY 73
+G I E+W+ SS L I V +D L + + +I SH++Y
Sbjct: 431 EGQTKRIGEKWNNFFSSLTLYILLYQVKVDMLISIHNGTISSHLVY 568
>TC206116 UP|CC48_SOYBN (P54774) Cell division cycle protein 48 homolog
(Valosin containing protein homolog) (VCP), complete
Length = 2583
Score = 25.4 bits (54), Expect = 4.8
Identities = 14/37 (37%), Positives = 21/37 (55%), Gaps = 3/37 (8%)
Frame = +2
Query: 1 MQEQWVKDVMR---VSGLQWLQHVSYLVIDQGLLSVI 34
M+ W+K M VSG +WL+ VS+L G+ S +
Sbjct: 614 MKRDWMKSGMMMWVVSGNKWLRFVSWLSFH*GIHSFL 724
>TC209384 homologue to UP|Q71SQ1 (Q71SQ1) MYC1, partial (41%)
Length = 1054
Score = 25.4 bits (54), Expect = 4.8
Identities = 9/32 (28%), Positives = 18/32 (56%)
Frame = -1
Query: 15 LQWLQHVSYLVIDQGLLSVIAERWHEVTSSFH 46
LQ L + YL+ ++ + + WH++ +FH
Sbjct: 526 LQ*LDVIDYLI*NKCFTNYLVTPWHQILKNFH 431
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.323 0.136 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,300,422
Number of Sequences: 63676
Number of extensions: 74772
Number of successful extensions: 463
Number of sequences better than 10.0: 52
Number of HSP's better than 10.0 without gapping: 462
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 463
length of query: 97
length of database: 12,639,632
effective HSP length: 73
effective length of query: 24
effective length of database: 7,991,284
effective search space: 191790816
effective search space used: 191790816
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 51 (24.3 bits)
Medicago: description of AC148347.8


