

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148289.16 + phase: 0
(200 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
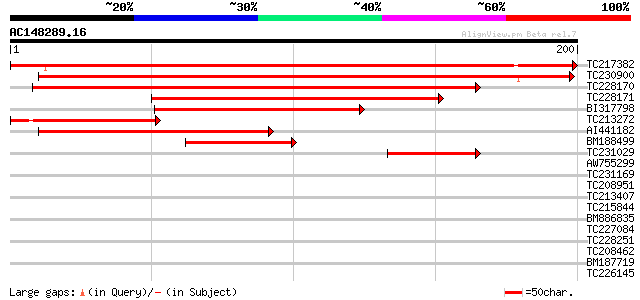
Sequences producing significant alignments: (bits) Value
TC217382 similar to UP|Q9FFD7 (Q9FFD7) Similarity to tyrosine ph... 346 5e-96
TC230900 similar to UP|Q9FFD7 (Q9FFD7) Similarity to tyrosine ph... 283 4e-77
TC228170 similar to UP|Q9ZVN4 (Q9ZVN4) T7A14.14 protein (At1g050... 224 2e-59
TC228171 similar to UP|Q9ZVN4 (Q9ZVN4) T7A14.14 protein (At1g050... 161 2e-40
BI317798 143 6e-35
TC213272 similar to UP|Q9FFD7 (Q9FFD7) Similarity to tyrosine ph... 86 9e-18
AI441182 83 7e-17
BM188499 59 2e-09
TC231029 similar to GB|AAL06938.1|15810022|AY054279 AT4g03960/T2... 44 4e-05
AW755299 39 0.002
TC231169 39 0.002
TC208951 29 1.3
TC213407 similar to UP|Q75QN6 (Q75QN6) PROPYZAMIDE-HTPERSENSITIV... 29 1.3
TC215844 UP|Q9SP48 (Q9SP48) Homeodomain-leucine zipper protein 5... 29 1.7
BM886835 similar to GP|26451280|db PTEN like protein {Arabidopsi... 28 3.7
TC227084 homologue to UP|LSM4_TOBAC (Q43582) Probable U6 snRNA-a... 28 3.7
TC228251 UP|CDPK_SOYBN (P28583) Calcium-dependent protein kinase... 28 3.7
TC208462 homologue to UP|LSM4_TOBAC (Q43582) Probable U6 snRNA-a... 27 4.8
BM187719 similar to GP|4249410|gb|A expressed protein {Arabidops... 27 6.3
TC226145 weakly similar to UP|Q9LEU7 (Q9LEU7) Serine/threonine p... 27 6.3
>TC217382 similar to UP|Q9FFD7 (Q9FFD7) Similarity to tyrosine phosphatase,
partial (81%)
Length = 1515
Score = 346 bits (887), Expect = 5e-96
Identities = 164/202 (81%), Positives = 184/202 (90%), Gaps = 2/202 (0%)
Frame = +2
Query: 1 MIVEVENIDED--DDVLIPPPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPY 58
MIVEVEN + D D VL+PPPNFSMVEDCI+RS LP PS+FPFLQTLNLRSIIYLCPEPY
Sbjct: 188 MIVEVENENGDLNDAVLVPPPNFSMVEDCIFRSGLPNPSNFPFLQTLNLRSIIYLCPEPY 367
Query: 59 PEENLDFLKEQNIRLFQFGIEGKTEVSLPALRDSIMEALKVLVDVRNHPILVHCKQGKHR 118
PEENLDFL+ QNIRLFQFGIEGKT++S+P L+DSIM+AL+VL+DVRNHP+L+HCK+GKHR
Sbjct: 368 PEENLDFLRSQNIRLFQFGIEGKTDISMPILKDSIMDALEVLIDVRNHPVLIHCKRGKHR 547
Query: 119 TGCLVGCFRKLQNWCLSSAFEEYQRFAGVKSRAADLTFIERFDLVSLRQCLYSIIYQYQG 178
TGCLVGC RKLQNWCLSS FEEYQRFAG KSR DLTFIE FD++SL QCLYSIIYQY G
Sbjct: 548 TGCLVGCLRKLQNWCLSSVFEEYQRFAGAKSRTTDLTFIEMFDVLSLSQCLYSIIYQYHG 727
Query: 179 ASKKRRLMYQDENIQKPRLTSF 200
SKKRRL+Y+DEN+QKPRLTSF
Sbjct: 728 -SKKRRLLYKDENLQKPRLTSF 790
>TC230900 similar to UP|Q9FFD7 (Q9FFD7) Similarity to tyrosine phosphatase,
partial (77%)
Length = 770
Score = 283 bits (724), Expect = 4e-77
Identities = 134/190 (70%), Positives = 162/190 (84%), Gaps = 1/190 (0%)
Frame = +2
Query: 11 DDDVLIPPPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENLDFLKEQN 70
+D+V++ P NFSMVE+ IYRSS P+ S+F FL++LNLRSIIYLCPEPYP+ENL+FL+ QN
Sbjct: 8 EDEVVVAPTNFSMVEEGIYRSSFPRSSNFSFLESLNLRSIIYLCPEPYPQENLEFLQSQN 187
Query: 71 IRLFQFGIEGKTEVSLPALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCFRKLQ 130
I F FGIEGKT++S+ A+RD+I+EA+KVL+DVRNHP+L+HC QGKHRTGC+VGC RKLQ
Sbjct: 188 ILCFHFGIEGKTDLSVSAVRDNILEAVKVLIDVRNHPVLIHCNQGKHRTGCVVGCLRKLQ 367
Query: 131 NWCLSSAFEEYQRFAGVKSRAADLTFIERFDLVSLRQCLYSIIYQYQG-ASKKRRLMYQD 189
+WCLSS FEEY+RFAG K R DL FIE DL+SLRQCL SIIYQY G ASKK RL Y+D
Sbjct: 368 SWCLSSVFEEYKRFAGAKYRTTDLRFIETVDLLSLRQCLNSIIYQYLGYASKKLRLRYRD 547
Query: 190 ENIQKPRLTS 199
EN KP+LTS
Sbjct: 548 ENSIKPQLTS 577
>TC228170 similar to UP|Q9ZVN4 (Q9ZVN4) T7A14.14 protein (At1g05000), partial
(74%)
Length = 1012
Score = 224 bits (571), Expect = 2e-59
Identities = 100/158 (63%), Positives = 128/158 (80%)
Frame = +1
Query: 9 DEDDDVLIPPPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENLDFLKE 68
D+ +D+ IPP NF+MV++ I+RS P+P++F FLQTL LRSIIYLCPEPYPE N++FLK
Sbjct: 103 DDGEDLFIPPLNFAMVDNGIFRSGFPEPANFSFLQTLGLRSIIYLCPEPYPEANMEFLKS 282
Query: 69 QNIRLFQFGIEGKTEVSLPALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCLVGCFRK 128
I+LFQFGIEG E + D+I EALKV++DVRNHP+++HCK+GKHRTGCLVGC+RK
Sbjct: 283 NGIKLFQFGIEGHKEPFVNIPEDTIREALKVVLDVRNHPVIIHCKRGKHRTGCLVGCYRK 462
Query: 129 LQNWCLSSAFEEYQRFAGVKSRAADLTFIERFDLVSLR 166
LQ WCLSS F+EYQRFA K+R +D F+E FD+ SL+
Sbjct: 463 LQKWCLSSVFDEYQRFAAAKARVSDQRFVELFDISSLK 576
>TC228171 similar to UP|Q9ZVN4 (Q9ZVN4) T7A14.14 protein (At1g05000), partial
(48%)
Length = 743
Score = 161 bits (408), Expect = 2e-40
Identities = 70/103 (67%), Positives = 85/103 (81%)
Frame = +3
Query: 51 IYLCPEPYPEENLDFLKEQNIRLFQFGIEGKTEVSLPALRDSIMEALKVLVDVRNHPILV 110
IYLCPEPYPE N++FLK I+LFQFGIEG E + D+I EALKV++DVRNHP+++
Sbjct: 3 IYLCPEPYPEANMEFLKSNGIKLFQFGIEGHKEPFVNIPEDTIREALKVVLDVRNHPVII 182
Query: 111 HCKQGKHRTGCLVGCFRKLQNWCLSSAFEEYQRFAGVKSRAAD 153
HCK+GKHRTGCLVGC+RKLQ WCLSS F+EYQRFA K+R +D
Sbjct: 183 HCKRGKHRTGCLVGCYRKLQKWCLSSVFDEYQRFAAAKARVSD 311
>BI317798
Length = 402
Score = 143 bits (360), Expect = 6e-35
Identities = 63/74 (85%), Positives = 71/74 (95%)
Frame = +3
Query: 52 YLCPEPYPEENLDFLKEQNIRLFQFGIEGKTEVSLPALRDSIMEALKVLVDVRNHPILVH 111
YLCPEPYPE NL+FL+ QNIRLFQFGIEGKT+VS+P L+DSIM+ALKVL+DVRNHPILVH
Sbjct: 180 YLCPEPYPEGNLEFLRSQNIRLFQFGIEGKTDVSMPVLKDSIMDALKVLIDVRNHPILVH 359
Query: 112 CKQGKHRTGCLVGC 125
CK+GKHRTGCLVGC
Sbjct: 360 CKRGKHRTGCLVGC 401
>TC213272 similar to UP|Q9FFD7 (Q9FFD7) Similarity to tyrosine phosphatase,
partial (20%)
Length = 433
Score = 86.3 bits (212), Expect = 9e-18
Identities = 40/53 (75%), Positives = 48/53 (90%)
Frame = +2
Query: 1 MIVEVENIDEDDDVLIPPPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYL 53
MIV+ EN D++D VL+PPPNF+MVEDC++RSS P PS+FPFLQTLNLRSIIYL
Sbjct: 278 MIVDFEN-DQNDAVLVPPPNFAMVEDCVFRSSFPTPSNFPFLQTLNLRSIIYL 433
>AI441182
Length = 417
Score = 83.2 bits (204), Expect = 7e-17
Identities = 37/83 (44%), Positives = 58/83 (69%)
Frame = +1
Query: 11 DDDVLIPPPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENLDFLKEQN 70
++++ +PP NF+MV++ I+R+ P ++F FL++L LRS++ LCPEPYPE +FLK
Sbjct: 166 EEEIFVPPLNFAMVDNGIFRAGFPDSANFGFLKSLRLRSVMCLCPEPYPETTSEFLKANG 345
Query: 71 IRLFQFGIEGKTEVSLPALRDSI 93
IRL+QF I+G E + D+I
Sbjct: 346 IRLYQFRIDGCKEPFVNIPNDTI 414
>BM188499
Length = 286
Score = 58.5 bits (140), Expect = 2e-09
Identities = 26/39 (66%), Positives = 36/39 (91%)
Frame = +2
Query: 63 LDFLKEQNIRLFQFGIEGKTEVSLPALRDSIMEALKVLV 101
L+FL+ QNIRLF FGIEGKT++S+ A+RD+I+EA+KVL+
Sbjct: 2 LEFLQSQNIRLFHFGIEGKTDLSVSAVRDNILEAVKVLI 118
>TC231029 similar to GB|AAL06938.1|15810022|AY054279 AT4g03960/T24M8_4
{Arabidopsis thaliana;} , partial (22%)
Length = 357
Score = 44.3 bits (103), Expect = 4e-05
Identities = 20/33 (60%), Positives = 25/33 (75%)
Frame = +3
Query: 134 LSSAFEEYQRFAGVKSRAADLTFIERFDLVSLR 166
LSS F+EYQRFAG K+R +D FIE FD+ L+
Sbjct: 15 LSSVFDEYQRFAGAKARVSDQRFIELFDISCLK 113
>AW755299
Length = 427
Score = 38.9 bits (89), Expect = 0.002
Identities = 19/36 (52%), Positives = 24/36 (65%)
Frame = +3
Query: 55 PEPYPEENLDFLKEQNIRLFQFGIEGKTEVSLPALR 90
P YP+ ++FL+ NIR FQF IEGKT +LP R
Sbjct: 138 PNQYPQ*TIEFLRSHNIRPFQFEIEGKTH-NLPLTR 242
>TC231169
Length = 884
Score = 38.9 bits (89), Expect = 0.002
Identities = 19/36 (52%), Positives = 24/36 (65%)
Frame = +1
Query: 55 PEPYPEENLDFLKEQNIRLFQFGIEGKTEVSLPALR 90
P YP+ ++FL+ NIR FQF IEGKT +LP R
Sbjct: 100 PNQYPQ*TIEFLRSHNIRPFQFEIEGKTH-NLPLTR 204
>TC208951
Length = 653
Score = 29.3 bits (64), Expect = 1.3
Identities = 13/45 (28%), Positives = 23/45 (50%)
Frame = +3
Query: 14 VLIPPPNFSMVEDCIYRSSLPKPSSFPFLQTLNLRSIIYLCPEPY 58
+++PP +V+ Y PSS P L ++ LR I+ + E +
Sbjct: 402 LIVPPATVIIVDHTTYPCRFLSPSSLPSLSSVFLRGIVGMASESF 536
>TC213407 similar to UP|Q75QN6 (Q75QN6) PROPYZAMIDE-HTPERSENSITIVE 1, partial
(19%)
Length = 572
Score = 29.3 bits (64), Expect = 1.3
Identities = 24/92 (26%), Positives = 38/92 (41%), Gaps = 3/92 (3%)
Frame = +1
Query: 42 LQTLNLRSIIYLCPEPYPEENLDFLKEQNIRLFQFGIEGKTEVSLPALRDSIMEALKVLV 101
LQ L + I+ LC + + F + F + + +S SI E +
Sbjct: 235 LQRLGITHILCLCTNEIGQSDSQFPDLFTYKNFSVCDDEDSNIS------SIFEEACDFI 396
Query: 102 DV---RNHPILVHCKQGKHRTGCLVGCFRKLQ 130
D H +LVHC +GK R+ LV + L+
Sbjct: 397 DYVEQAGHSVLVHCFEGKSRSATLVLAYLMLR 492
>TC215844 UP|Q9SP48 (Q9SP48) Homeodomain-leucine zipper protein 56, complete
Length = 1482
Score = 28.9 bits (63), Expect = 1.7
Identities = 17/43 (39%), Positives = 23/43 (52%), Gaps = 1/43 (2%)
Frame = -3
Query: 14 VLIPPPNFSMVEDC-IYRSSLPKPSSFPFLQTLNLRSIIYLCP 55
VL+P F V C + RSS PKPS P L ++ +S + P
Sbjct: 370 VLVPASTFGQVRVCSLLRSSSPKPSPDPRLVVVSSQSGALVLP 242
>BM886835 similar to GP|26451280|db PTEN like protein {Arabidopsis thaliana},
partial (29%)
Length = 422
Score = 27.7 bits (60), Expect = 3.7
Identities = 14/37 (37%), Positives = 20/37 (53%)
Frame = +1
Query: 109 LVHCKQGKHRTGCLVGCFRKLQNWCLSSAFEEYQRFA 145
++HC GK RTG +V + +C SA E Q +A
Sbjct: 181 VIHCMAGKGRTGLMVSAY---LTYCGMSADEALQLYA 282
>TC227084 homologue to UP|LSM4_TOBAC (Q43582) Probable U6 snRNA-associated
Sm-like protein LSm4 (Glycine-rich protein 10) (GRP 10),
partial (83%)
Length = 926
Score = 27.7 bits (60), Expect = 3.7
Identities = 15/39 (38%), Positives = 20/39 (50%)
Frame = +3
Query: 87 PALRDSIMEALKVLVDVRNHPILVHCKQGKHRTGCLVGC 125
P + D M L +L + HP+LV K G+ G LV C
Sbjct: 135 PPIND--MLPLSLLKTAQGHPMLVELKNGETYNGHLVNC 245
>TC228251 UP|CDPK_SOYBN (P28583) Calcium-dependent protein kinase SK5 (CDPK)
, partial (16%)
Length = 445
Score = 27.7 bits (60), Expect = 3.7
Identities = 17/72 (23%), Positives = 31/72 (42%), Gaps = 16/72 (22%)
Frame = -2
Query: 78 IEGKTEVSLPALRDSIMEALKVLVDVRNHPILV----------------HCKQGKHRTGC 121
+EG E++LP L +++ VL +R +P + H K + C
Sbjct: 348 LEGGPELALPELPPNLVHLTDVLRALRENPRRLQSHHVCCCRTRTRFRSHSCCRKRKGDC 169
Query: 122 LVGCFRKLQNWC 133
+V ++ + NWC
Sbjct: 168 IVSAYQVVVNWC 133
>TC208462 homologue to UP|LSM4_TOBAC (Q43582) Probable U6 snRNA-associated
Sm-like protein LSm4 (Glycine-rich protein 10) (GRP 10),
partial (94%)
Length = 794
Score = 27.3 bits (59), Expect = 4.8
Identities = 13/32 (40%), Positives = 17/32 (52%)
Frame = +3
Query: 94 MEALKVLVDVRNHPILVHCKQGKHRTGCLVGC 125
M L +L + HP+LV K G+ G LV C
Sbjct: 129 MLPLSLLKTAQGHPMLVELKNGETYNGHLVNC 224
>BM187719 similar to GP|4249410|gb|A expressed protein {Arabidopsis
thaliana}, partial (14%)
Length = 423
Score = 26.9 bits (58), Expect = 6.3
Identities = 14/42 (33%), Positives = 20/42 (47%)
Frame = -2
Query: 34 PKPSSFPFLQTLNLRSIIYLCPEPYPEENLDFLKEQNIRLFQ 75
P+ S FLQ LN S+ ++ P YP L +E + Q
Sbjct: 326 PQNISVHFLQNLNHSSVTHVGPTTYPRGGLVVFREARLSASQ 201
>TC226145 weakly similar to UP|Q9LEU7 (Q9LEU7) Serine/threonine protein
kinase-like protein (CBL-interacting protein kinase 5),
partial (15%)
Length = 890
Score = 26.9 bits (58), Expect = 6.3
Identities = 14/41 (34%), Positives = 21/41 (51%)
Frame = +1
Query: 30 RSSLPKPSSFPFLQTLNLRSIIYLCPEPYPEENLDFLKEQN 70
R S+P P+ Q +R I + E Y E+N+DF +N
Sbjct: 16 RYSIPDIMRDPWFQVGFMRPIAFSIKESYVEDNIDFDDVEN 138
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.324 0.141 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,786,946
Number of Sequences: 63676
Number of extensions: 157025
Number of successful extensions: 942
Number of sequences better than 10.0: 46
Number of HSP's better than 10.0 without gapping: 932
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 940
length of query: 200
length of database: 12,639,632
effective HSP length: 92
effective length of query: 108
effective length of database: 6,781,440
effective search space: 732395520
effective search space used: 732395520
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 56 (26.2 bits)
Medicago: description of AC148289.16


