

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148241.11 + phase: 0
(390 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
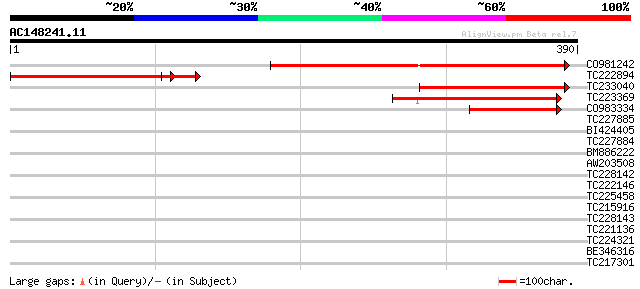
Score E
Sequences producing significant alignments: (bits) Value
CO981242 298 3e-81
TC222894 181 7e-46
TC233040 180 1e-45
TC223369 116 2e-26
CO983334 79 5e-15
TC227885 36 0.026
BI424405 35 0.076
TC227884 34 0.099
BM886222 weakly similar to PIR|S51591|S51 chitinase (EC 3.2.1.14... 33 0.29
AW203508 weakly similar to SP|P57758|CTNS_ Cystinosin homolog. [... 31 0.84
TC228142 similar to UP|YPL5_ARATH (Q9T096) Yippee-like protein A... 31 0.84
TC222146 weakly similar to UP|Q940Y1 (Q940Y1) AT4g22540/F7K2_120... 30 1.4
TC225458 UP|Q9SWB5 (Q9SWB5) Seed maturation protein PM37, complete 30 1.9
TC215916 similar to UP|Q9M011 (Q9M011) Light-inducible protein A... 30 1.9
TC228143 similar to UP|YPL5_ARATH (Q9T096) Yippee-like protein A... 30 2.4
TC221136 homologue to UP|LBD4_ARATH (Q9SHE9) LOB domain protein ... 30 2.4
TC224321 similar to UP|Q6H5X0 (Q6H5X0) Ripening-related protein-... 28 7.1
BE346316 similar to PIR|T07126|T0 magnesium chelatase (EC 4.99.1... 28 9.3
TC217301 similar to UP|Q9FGK9 (Q9FGK9) Dbj|BAA90625.1, partial (5%) 28 9.3
>CO981242
Length = 847
Score = 298 bits (763), Expect = 3e-81
Identities = 147/206 (71%), Positives = 172/206 (83%)
Frame = -2
Query: 180 EYYYMSARSLAGSATPPSFTHLRAAKSGPSALEFIHDSSDDDEASQVTSNISTTKPWSIP 239
E+YYMSARSLAGS TPP T++ AAKSGP+A+E I+DSS D+EA +SN S T+ IP
Sbjct: 843 EHYYMSARSLAGSGTPPWGTYMGAAKSGPAAMESINDSSSDNEAPPASSNNSATQAMPIP 664
Query: 240 RSVDGRYGTFLATAINLPLKGNSMRYGYIGFTGIKLLKVYVVVHSTYGQYLGWIMAAIYT 299
RSV G YGTFLA A+NLPL+GN++R GYIGF G KLL+ Y HST+GQ+LGW+MAAIY
Sbjct: 663 RSVAGSYGTFLAAAVNLPLRGNALREGYIGFGGRKLLQEYET-HSTFGQWLGWLMAAIYI 487
Query: 300 CSRIPQIWLNIKRGSVEGLNPFMFVFALIANTSYVGSILVRTTEFESIKANLPWLLDATV 359
R+PQIWLNIKRGSVEGLNPFMFVFAL+AN +YVGSILVRTTE+E IKAN+PWLLDA +
Sbjct: 486 SGRVPQIWLNIKRGSVEGLNPFMFVFALVANVTYVGSILVRTTEWERIKANMPWLLDAVI 307
Query: 360 CVALDFFIISQYIYYRYFRSSESSDD 385
CVALDFFIISQYIYYR F+ E+ DD
Sbjct: 306 CVALDFFIISQYIYYRCFQRREARDD 229
>TC222894
Length = 537
Score = 181 bits (458), Expect = 7e-46
Identities = 84/114 (73%), Positives = 92/114 (80%)
Frame = +3
Query: 1 MAPSYCVKENKECVKWVETYFKDCLCNLRDDISFSLGLMSLVSWGVAEIPQIITIFRNKS 60
M PSYCVKE K CV+WVE YFKDCLCNL+D+ISFS GL SLV WGVAEIPQIITIFR K
Sbjct: 150 MPPSYCVKERKPCVRWVEKYFKDCLCNLKDEISFSFGLTSLVFWGVAEIPQIITIFRTKK 329
Query: 61 SHGISLAFLLTWVAGDICNLVGCLLEPATLPTQFYTALTVINCSFMQALQSYYY 114
SHG+SL FLLTWVAGDICNL GC+LEPATLPTQ+YTAL + + L YY
Sbjct: 330 SHGVSLVFLLTWVAGDICNLTGCILEPATLPTQYYTALLYTITTIVLLLLIVYY 491
Score = 43.5 bits (101), Expect = 2e-04
Identities = 19/27 (70%), Positives = 22/27 (81%)
Frame = +1
Query: 105 FMQALQSYYYCKLYTMITSSDGANIVR 131
F Q+L+SYYYC YTM TS+DG NIVR
Sbjct: 445 FTQSLRSYYYC*SYTMTTSADGTNIVR 525
>TC233040
Length = 725
Score = 180 bits (456), Expect = 1e-45
Identities = 82/103 (79%), Positives = 93/103 (89%)
Frame = +2
Query: 283 HSTYGQYLGWIMAAIYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIANTSYVGSILVRTT 342
HST+GQ+LGW+MAAIY R+PQIWLNIKR +VEGLNPFMFVFAL+AN +YVGSILVRTT
Sbjct: 14 HSTFGQWLGWLMAAIYISGRVPQIWLNIKRSTVEGLNPFMFVFALVANVTYVGSILVRTT 193
Query: 343 EFESIKANLPWLLDATVCVALDFFIISQYIYYRYFRSSESSDD 385
E+ESIKAN+PWLLDA +CVALD FIISQYIYYRYFR E+ DD
Sbjct: 194 EWESIKANMPWLLDAVICVALDIFIISQYIYYRYFRRREAKDD 322
>TC223369
Length = 706
Score = 116 bits (290), Expect = 2e-26
Identities = 59/122 (48%), Positives = 79/122 (64%), Gaps = 6/122 (4%)
Frame = +3
Query: 264 RYGYIGFTGIKLLKVY------VVVHSTYGQYLGWIMAAIYTCSRIPQIWLNIKRGSVEG 317
R ++ + G KL +V V + G + GW M IY R+PQI LNI+RG VEG
Sbjct: 9 RQQFVIYVGRKLFQVSDDQLPKTDVSGSIGTFFGWAMTFIYLGGRLPQICLNIRRGHVEG 188
Query: 318 LNPFMFVFALIANTSYVGSILVRTTEFESIKANLPWLLDATVCVALDFFIISQYIYYRYF 377
LNP MF+FA+I N +YV SILV + ++ I+ NLPWL+DA CV LDFFI+ Q+IY+R +
Sbjct: 189 LNPLMFLFAVIGNATYVASILVISLDWSKIRPNLPWLVDAGGCVLLDFFILMQFIYFRCW 368
Query: 378 RS 379
S
Sbjct: 369 TS 374
>CO983334
Length = 807
Score = 78.6 bits (192), Expect = 5e-15
Identities = 35/63 (55%), Positives = 48/63 (75%)
Frame = -3
Query: 317 GLNPFMFVFALIANTSYVGSILVRTTEFESIKANLPWLLDATVCVALDFFIISQYIYYRY 376
GLNP MF+FA+I N +YV SI+V + ++ I+ NLPWL+DA CV LDFFI+ Q+IY+R
Sbjct: 616 GLNPLMFLFAVIGNATYVASIVVISLDWSKIRPNLPWLVDAGGCVLLDFFILMQFIYFRC 437
Query: 377 FRS 379
+ S
Sbjct: 436 WTS 428
>TC227885
Length = 656
Score = 36.2 bits (82), Expect = 0.026
Identities = 29/82 (35%), Positives = 40/82 (48%), Gaps = 2/82 (2%)
Frame = +1
Query: 296 AIYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIANTSYVGSILVRTTEFESIKANLP--W 353
AI+ +RIPQIW N S L + I + G +VR F +I+ N P
Sbjct: 97 AIFLFARIPQIWQNFSNKSTGEL-------SFITSFMNFGGSMVRV--FTTIQENAPKSV 249
Query: 354 LLDATVCVALDFFIISQYIYYR 375
LL + VA +F I+SQ I Y+
Sbjct: 250 LLGYAIGVATNFTILSQIIAYQ 315
>BI424405
Length = 421
Score = 34.7 bits (78), Expect = 0.076
Identities = 16/33 (48%), Positives = 22/33 (66%)
Frame = +1
Query: 36 LGLMSLVSWGVAEIPQIITIFRNKSSHGISLAF 68
LG ++ +SW VA PQ+I FR KS G+SL +
Sbjct: 112 LGWLAFLSWSVAGYPQLILNFRRKSVVGLSLDY 210
Score = 29.3 bits (64), Expect = 3.2
Identities = 13/32 (40%), Positives = 20/32 (61%)
Frame = +1
Query: 288 QYLGWIMAAIYTCSRIPQIWLNIKRGSVEGLN 319
Q LGW+ ++ + PQ+ LN +R SV GL+
Sbjct: 106 QVLGWLAFLSWSVAGYPQLILNFRRKSVVGLS 201
>TC227884
Length = 1152
Score = 34.3 bits (77), Expect = 0.099
Identities = 28/82 (34%), Positives = 40/82 (48%), Gaps = 2/82 (2%)
Frame = +3
Query: 296 AIYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIANTSYVGSILVRTTEFESIKANLP--W 353
AI+ +RIPQIW N S L + I + G +VR F +I+ + P
Sbjct: 615 AIFLFARIPQIWQNFSNKSTGEL-------SFITSFMNFGGSMVRV--FTTIQESAPKSV 767
Query: 354 LLDATVCVALDFFIISQYIYYR 375
LL + VA +F I+SQ I Y+
Sbjct: 768 LLGYAIGVATNFTILSQIIAYQ 833
>BM886222 weakly similar to PIR|S51591|S51 chitinase (EC 3.2.1.14) / lysozyme
(EC 3.2.1.17) PZ precursor pathogenesis-related -
common, partial (32%)
Length = 425
Score = 32.7 bits (73), Expect = 0.29
Identities = 26/99 (26%), Positives = 42/99 (42%), Gaps = 4/99 (4%)
Frame = +1
Query: 174 KEAARGEYYYMSARSLAGSATPPSFTHLRAAKS--GPSALEFIHDSSDDDEASQVTSNIS 231
K A +G Y+Y + + P FTHL A + P+ + SSD + S T +
Sbjct: 7 KAAIKGGYWYSESGLAVSNINPSHFTHLFCAFAHLDPNTNKVTISSSDSSQFSTFTQTLQ 186
Query: 232 TTKPWSIPR--SVDGRYGTFLATAINLPLKGNSMRYGYI 268
P S+ S+ G +G LA + + + R +I
Sbjct: 187 AKNP-SVKTLLSIGGGFGPSLAANFSRMARQANTRKSFI 300
>AW203508 weakly similar to SP|P57758|CTNS_ Cystinosin homolog. [Mouse-ear
cress] {Arabidopsis thaliana}, partial (35%)
Length = 348
Score = 31.2 bits (69), Expect = 0.84
Identities = 26/65 (40%), Positives = 33/65 (50%), Gaps = 3/65 (4%)
Frame = +2
Query: 36 LGLMSLVSWGVAEIPQIITIFRNKSSHGISLAFLLTWVAGDICNLV---GCLLEPATLPT 92
LG ++ +SW VA PQ+I R KS G SL + +I NL L+ ATLP
Sbjct: 77 LGWLAYLSWSVAGYPQLILNCRRKSVVGSSLDY-------EILNLTKHFSYLIYNATLP- 232
Query: 93 QFYTA 97
F TA
Sbjct: 233 -FVTA 244
Score = 28.1 bits (61), Expect = 7.1
Identities = 15/47 (31%), Positives = 25/47 (52%)
Frame = +2
Query: 288 QYLGWIMAAIYTCSRIPQIWLNIKRGSVEGLNPFMFVFALIANTSYV 334
Q LGW+ ++ + PQ+ LN +R SV G + + L + SY+
Sbjct: 71 QLLGWLAYLSWSVAGYPQLILNCRRKSVVGSSLDYEILNLTKHFSYL 211
>TC228142 similar to UP|YPL5_ARATH (Q9T096) Yippee-like protein At4g27740,
partial (72%)
Length = 624
Score = 31.2 bits (69), Expect = 0.84
Identities = 13/25 (52%), Positives = 17/25 (68%)
Frame = -3
Query: 80 LVGCLLEPATLPTQFYTALTVINCS 104
L+GC + LPTQ +TA+T IN S
Sbjct: 349 LLGCFVSSFILPTQHFTAITTINIS 275
>TC222146 weakly similar to UP|Q940Y1 (Q940Y1) AT4g22540/F7K2_120, partial
(7%)
Length = 535
Score = 30.4 bits (67), Expect = 1.4
Identities = 11/35 (31%), Positives = 22/35 (62%)
Frame = +1
Query: 115 CKLYTMITSSDGANIVRMLRVNCYFYNCIILQDNE 149
C+++ ++ S N VR++ + C +Y+C+ DNE
Sbjct: 418 CEVHVYLSLSGPPNEVRIVMMTC*YYSCVADNDNE 522
>TC225458 UP|Q9SWB5 (Q9SWB5) Seed maturation protein PM37, complete
Length = 1797
Score = 30.0 bits (66), Expect = 1.9
Identities = 26/95 (27%), Positives = 42/95 (43%), Gaps = 8/95 (8%)
Frame = -3
Query: 49 IPQIITIFRNKSSHGISLAFLL-----TWVAGDICNLVGCLLEPATLPTQFYTALTVINC 103
I II IF N S + + LL + L+ CLL LP + ++ C
Sbjct: 1426 INSIIIIFINSESSFHTKSLLLGALYPLCTTRHVIILIICLLSLLLLPPSLLLHVDIMEC 1247
Query: 104 SFMQALQSYYYCKLYTMI---TSSDGANIVRMLRV 135
SF+ +Q + C+L + S G N++R+ R+
Sbjct: 1246 SFLTFIQ-LHVCQL*RRLWWQNSLQGLNLIRV*RI 1145
>TC215916 similar to UP|Q9M011 (Q9M011) Light-inducible protein ATLS1,
complete
Length = 630
Score = 30.0 bits (66), Expect = 1.9
Identities = 13/35 (37%), Positives = 20/35 (57%)
Frame = +3
Query: 342 TEFESIKANLPWLLDATVCVALDFFIISQYIYYRY 376
+EF I +PW+ D V + L+FFI +Y + Y
Sbjct: 384 SEFRVILVMMPWMSDELVILFLNFFICKKYSEF*Y 488
>TC228143 similar to UP|YPL5_ARATH (Q9T096) Yippee-like protein At4g27740,
partial (72%)
Length = 660
Score = 29.6 bits (65), Expect = 2.4
Identities = 12/25 (48%), Positives = 16/25 (64%)
Frame = -1
Query: 80 LVGCLLEPATLPTQFYTALTVINCS 104
L+GC + LPTQ +TA+ IN S
Sbjct: 366 LLGCFISSFILPTQHFTAIATINIS 292
>TC221136 homologue to UP|LBD4_ARATH (Q9SHE9) LOB domain protein 4, partial
(44%)
Length = 607
Score = 29.6 bits (65), Expect = 2.4
Identities = 14/33 (42%), Positives = 20/33 (60%)
Frame = -2
Query: 60 SSHGISLAFLLTWVAGDICNLVGCLLEPATLPT 92
+S+ I L L W +G+ CN++ LL P TL T
Sbjct: 111 ASYTIELTASLLWCSGNSCNILFTLLAPKTL*T 13
>TC224321 similar to UP|Q6H5X0 (Q6H5X0) Ripening-related protein-like,
partial (52%)
Length = 743
Score = 28.1 bits (61), Expect = 7.1
Identities = 21/67 (31%), Positives = 29/67 (42%)
Frame = -3
Query: 187 RSLAGSATPPSFTHLRAAKSGPSALEFIHDSSDDDEASQVTSNISTTKPWSIPRSVDGRY 246
RS S T P + R+++S P +L F + T+N T PW +P Y
Sbjct: 543 RSR*QSRTGPPLSPSRSSRSPP*SLSFSN-----------TANCR*TTPWRVPPHAPCCY 397
Query: 247 GTFLATA 253
GT TA
Sbjct: 396 GTSRRTA 376
>BE346316 similar to PIR|T07126|T0 magnesium chelatase (EC 4.99.1.-) chain
chlH - soybean, partial (9%)
Length = 625
Score = 27.7 bits (60), Expect = 9.3
Identities = 11/25 (44%), Positives = 16/25 (64%), Gaps = 2/25 (8%)
Frame = +2
Query: 135 VNCYFY--NCIILQDNEEEKRPLNP 157
+NC + N +QD EE K+P+NP
Sbjct: 413 INCVYEEANTTFIQDEEETKKPMNP 487
>TC217301 similar to UP|Q9FGK9 (Q9FGK9) Dbj|BAA90625.1, partial (5%)
Length = 1110
Score = 27.7 bits (60), Expect = 9.3
Identities = 15/46 (32%), Positives = 21/46 (45%)
Frame = +2
Query: 192 SATPPSFTHLRAAKSGPSALEFIHDSSDDDEASQVTSNISTTKPWS 237
+A F A S +E I +S +D A+ +ISTT WS
Sbjct: 143 AANAKFFVPTPAPSSNERTIEAIVESKQEDNATNEYPSISTTNEWS 280
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.322 0.137 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,021,930
Number of Sequences: 63676
Number of extensions: 304328
Number of successful extensions: 1554
Number of sequences better than 10.0: 39
Number of HSP's better than 10.0 without gapping: 1540
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1553
length of query: 390
length of database: 12,639,632
effective HSP length: 99
effective length of query: 291
effective length of database: 6,335,708
effective search space: 1843691028
effective search space used: 1843691028
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC148241.11


