

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
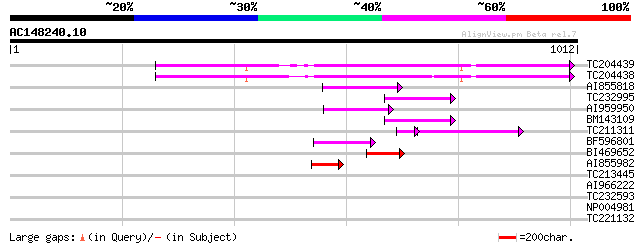
Query= AC148240.10 - phase: 0 /pseudo
(1012 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete 117 3e-26
TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, co... 107 2e-23
AI855818 weakly similar to GP|21741393|e OSJNBb0051N19.6 {Oryza ... 83 7e-16
TC232995 79 8e-15
AI959950 74 2e-13
BM143109 64 3e-10
TC211311 weakly similar to UP|O24587 (O24587) Pol protein, parti... 46 2e-08
BF596801 weakly similar to GP|29423270|g gag-pol polyprotein {Gl... 54 3e-07
BI469652 weakly similar to GP|18149115|dbj| reverse transcriptas... 52 2e-06
AI855982 45 2e-04
TC213445 39 0.011
AI966222 33 0.071
TC232593 weakly similar to UP|Q9XG91 (Q9XG91) Tpv2-1c protein (F... 30 4.1
NP004981 early nodulin 30 4.1
TC221132 weakly similar to UP|O23529 (O23529) RETROTRANSPOSON li... 29 9.1
>TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete
Length = 4731
Score = 117 bits (293), Expect = 3e-26
Identities = 207/759 (27%), Positives = 310/759 (40%), Gaps = 12/759 (1%)
Frame = +2
Query: 261 KSEVIMVENLKMSHLNFFVKNMGFSMNSLLLELHNKMGL*RERTEPYKKWPEP*SMKTI* 320
+S V M ENLK + M+SL HN+MG R +T K+ M
Sbjct: 2450 ESGVTMAENLKTAGSLNSAHLKASLMSSLQPLHHNRMG*LRGKTGLCKRLLGSCFMPKNF 2629
Query: 321 LNIFGQKQLILHVTFKIGSISDLCWRKLHMNSLKEEDPISLTFISLDVLVTS*TLKTI*R 380
I G K H T S + + M S K +S T SL+V VTS +++
Sbjct: 2630 PIISGLKP*TQHATSTTESH*EEGLQPPCMKSGKGGSHLSSTSTSLEVHVTSWQIESKEE 2809
Query: 381 NLMPRLKEESF*VTLKGQRHTECIIQKHIVLKNLSM*NLMI----ESLEIKLQSKVKVMQ 436
+PR+ +E TL+ H E I + N SM LMI E K S+ +
Sbjct: 2810 RWIPRVMQEYSWDTLQTAEHIEYSIPEPEQ*WNPSMWLLMICLQQERRMSKKMSEHRETM 2989
Query: 437 VQLILKMHQNLINLLILKSIQKLNLVQKLKSLQKLSLIQKQKLVQ*NRMKMLLRMFKITL 496
Q+ LK+ + L++L+ Q + L+ + L +
Sbjct: 2990 *QMQLKVEKMQKTLILLQMNQTSTNPTRDPPLESRRCTPRS*L*E--------------- 3124
Query: 497 SRLFSQNSNTNLHILRS*SLEAKIVLEEQDHISDKKSL**DCFQ*LSLKLLKKLSQMMVG 556
+ ++ S + LRS Q H+ K LS ++ K+ QM G
Sbjct: 3125 --IQTEGSLQDQGRLRS----------SQTHVLSPK---------LSPRM*KRH*QMSSG 3241
Query: 557 Y*PCKKN*ISFREMMCGI*YPNLFRRTLLEQNGYSETS*MKKEK*PETKPDLLLKVTVNK 616
CKKN + + M G *+ L L +G S T MKK *PET+PD LLK T+
Sbjct: 3242 SMLCKKNWSNSKGMKSGS*FLGLRELM*LAPSGSSRTKPMKKVS*PETRPDWLLKATLRL 3421
Query: 617 KALITLKHLLQLQDWKQSGYFYPMQLTMA*YCIKWMSKVLFLMVSLKKKYMSNNLLGLRI 676
K ++ L QL D S Y+ ++ C +WM + F M + KK M ++ L+
Sbjct: 3422 KV*TLMRLLPQLLDLSPSDYYLV*LVSSNSSCTRWM*RAHF*MDT*MKKSMWSSQRDLQT 3601
Query: 677 LSILTMFINLRNHYMA*NKLPELGMID*VIS*LKMILKEDKLTQHSSERHLRRTF*LCKY 736
I M+ R M *+KL ELGM * S L + ++LT+ S + +T *L +Y
Sbjct: 3602 RLIQIMYTGSRRLSMD*SKLQELGMKG*QSSLLSKGIGREELTRPSLSNKMLKT**LHRY 3781
Query: 737 MLMI*YLVLLMHLFAKNSLS*CRMNLK*V*WEN*SSFLEFKSTKVKKEYMFIKQNIQRSF 796
MLM L + C +NL+*V E+* F +FK ++ + Y K +QR+
Sbjct: 3782 MLMTLCLEGCRMRCFDILFNRCNLNLR*VLLES*LIFWDFK*SRWRTPYSSHKAGMQRTL 3961
Query: 797 *RSSS*KI--------VK**TLQCIQPAP*AKKILEQ**TRSYTEV*LVLCYTSLHLDLI 848
RS ++ + *+ Q ++ AP K+ TE * Y D
Sbjct: 3962 SRSLGWRMPVIKGHLHLLT*SCQRMKQAPVLIKVC--------TEA**GAYYI*QLADPT 4117
Query: 849 SYSVYACVQDFNQILENLI*LLLRESSGI*KEQLILDSFIGNP*ITS*LDSVMLIMLVTG 908
S VQD I + *L RE +* + + I L VMLI L
Sbjct: 4118 SPMQ*VFVQDIKPIPR*VT*LK*REF*NM*MALVTMGLCTVIVQIQCWLGIVMLIGLEVQ 4297
Query: 909 LKENQPVEIVNSWEKI*YPGQAKDKQQLLCLQQKQNTFQLQVVVHNYFG*NISWKTIRLM 968
+ E + + WE + G A+ + LQQK + Q + VH+ FG*+ ++
Sbjct: 4298 MTEKALLVDASIWETTLFHGSARSRTVCPYLQQKPSILQQEAAVHS*FG*SRC*RSTMSN 4477
Query: 969 LTVFPFFVIILLLSVCQRIQFYIQEPSILRSNTILSETM 1007
V +L + +I F EPS L + +SE +
Sbjct: 4478 KMS*HCTVTT*VLLIFLKILFNTAEPSTLTLDITISEIL 4594
>TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, complete
Length = 4734
Score = 107 bits (268), Expect = 2e-23
Identities = 208/760 (27%), Positives = 310/760 (40%), Gaps = 13/760 (1%)
Frame = +2
Query: 261 KSEVIMVENLKMSHLNFFVKNMGFSMNSLLLELHNKMGL*RERTEPYKKWPEP*SMKTI* 320
+S V M E+LK + L M+SL HNKM + +T KK M
Sbjct: 2453 ESGVTMAESLKTASLLNSAHLKASLMSSLQPLHHNKMA*LKGKTGLCKKLLGSCFMPKNF 2632
Query: 321 LNIFGQKQLILHVTFKIGSISDLCWRKLHMNSLKEEDPISLTFISLDVLVTS*TLKTI*R 380
I G K H T S + + M S K +S T SL+V VT +++
Sbjct: 2633 PIISGLKP*TQHATSTTESHLEEGLQPHCMKSGKGGSQLSSTSTSLEVHVTFWQIESKGE 2812
Query: 381 NLMPRLKEESF*VTLKGQRHTECIIQKHIVLKNLSM*NLMI----ESLEIKLQSKVKVMQ 436
+PR+ +E TL+ H E I + + N SM LMI E K S+ +
Sbjct: 2813 RWIPRVMQEYSWDTLQTAEHIEYSIPEPEL*WNPSMWLLMI*LQQERRMSKKMSEHRETM 2992
Query: 437 VQLILKMHQNLINLLILKSIQKLNLVQKLKSLQKLSLIQKQKLVQ*NRMKMLLRMFKITL 496
Q+ LK+ + L++L+ Q + + L+ + L + + L ++ L
Sbjct: 2993 *QIQLKVQKMQKTLILLQMNQTSINLTRDPPLESRRCTPRS*L*EIQTEESLQDQGRLRL 3172
Query: 497 SRLFSQNSNTNLHILRS*SLEAKIVLEEQDHISDKKSL**DCFQ*LSLKLLKKLSQMMVG 556
S + H+ K LS ++ K+ M G
Sbjct: 3173 SPI---------------------------HVLSPK---------LSPRM*KRH*LMSSG 3244
Query: 557 Y*PCKKN*ISFREMMCGI*YPNLFRRTLLEQNGYSETS*MKKEK*PETKPDLLLKVTVNK 616
CKKN + + M G *+ + L +G S T MKK *PET+PDLLLK T+
Sbjct: 3245 SMLCKKNWSNSKGMKFGS*FLDPRELM*LAPSGSSRTKPMKKVL*PETRPDLLLKATLRL 3424
Query: 617 KALITLKHLLQLQDWKQSGYFYPMQLTMA*YCIKWMSKVLFLMVSLKKKYMSNNLLGLRI 676
K +K L D S + + C +WM + F M + KK M ++ L I
Sbjct: 3425 KV*TLMKLSPLLLDLSPSDCYLV*LASSNSSCTRWM*RARF*MDT*MKKPMWSSQRDL*I 3604
Query: 677 LSILTMFINLRNHYMA*NKLPELGMID*VIS*LKMILKEDKLTQHSSERHLRRTF*LCKY 736
I M+ R M *+KL ELGM * S L + ++LT+ S + +T * +Y
Sbjct: 3605 QLIQIMYTGSRRLSMD*SKLQELGMKG*QSSLLSKGIGREELTRLSLSNKMLKT***HRY 3784
Query: 737 MLMI*YLV-LLMHLFAKNSLS*CRMNLK*V*WEN*SSFLEFKSTKVKKEYMFIKQNIQRS 795
MLM L M F S + C +NL+*V E+* F + K ++ K Y K ++QR+
Sbjct: 3785 MLMTLCLEGCRMRCFDILS-NRCNLNLR*VLLES*LIFWDSK*SRWKTPYSSHKASMQRT 3961
Query: 796 F*RSSS*KI--------VK**TLQCIQPAP*AKKILEQ**TRSYTEV*LVLCYTSLHLDL 847
RS K+ + *+ Q ++ AP K+ TE *L Y DL
Sbjct: 3962 LSRSLGWKMPAIKEHLHLLT*SCQKMKLAPVLIKVC--------TEA*LGAYYI*QLADL 4117
Query: 848 ISYSVYACVQDFNQILENLI*LLLRESSGI*KEQLILDSFIGNP*ITS*LDSVMLIMLVT 907
S VQD IL + *+ RE +* + + I L VMLI L
Sbjct: 4118 TSPMQ*VFVQDIKPILR*VT*IK*REF*NM*MAPVTMGLCTVIVQIQCWLGIVMLIGLEV 4297
Query: 908 GLKENQPVEIVNSWEKI*YPGQAKDKQQLLCLQQKQNTFQLQVVVHNYFG*NISWKTIRL 967
+ E + V+ WE I + G A+ + L KQ+ Q + VHN FG*+ ++
Sbjct: 4298 QMTEKALLVDVSIWEPILFHGSARSRTVCPYLLLKQSILQQEAAVHN*FG*SRC*RSTMS 4477
Query: 968 MLTVFPFFVIILLLSVCQRIQFYIQEPSILRSNTILSETM 1007
V +L + +I F EPS L + + E +
Sbjct: 4478 NKMS*HCTVTT*VLLIFLKILFNTAEPSTLTLDITILEIL 4597
>AI855818 weakly similar to GP|21741393|e OSJNBb0051N19.6 {Oryza sativa
(japonica cultivar-group)}, partial (10%)
Length = 463
Score = 82.8 bits (203), Expect = 7e-16
Identities = 63/143 (44%), Positives = 75/143 (52%)
Frame = -1
Query: 559 PCKKN*ISFREMMCGI*YPNLFRRTLLEQNGYSETS*MKKEK*PETKPDLLLKVTVNKKA 618
PCKKN*I+ +E+MCG * NL EQNG+ E +*M E + D K + K+
Sbjct: 457 PCKKN*INLKEIMCGN**KNLKIILS*EQNGFLEIN*MNMA*LLEIRLD**QKGIIKKRE 278
Query: 619 LITLKHLLQLQDWKQSGYFYPMQLTMA*YCIKWMSKVLFLMVSLKKKYMSNNLLGLRILS 678
KH+LQLQD K F M IKWM KVLF MV KKKYM NN L+
Sbjct: 277 *TMKKHMLQLQD*KSLECF*HMYP**ILNSIKWMLKVLF*MV*FKKKYMLNNPQALKSRI 98
Query: 679 ILTMFINLRNHYMA*NKLPELGM 701
MFIN + +M *NK GM
Sbjct: 97 NQLMFINCKRLFMV*NKPLGRGM 29
>TC232995
Length = 1009
Score = 79.3 bits (194), Expect = 8e-15
Identities = 58/127 (45%), Positives = 72/127 (56%)
Frame = +3
Query: 669 NNLLGLRILSILTMFINLRNHYMA*NKLPELGMID*VIS*LKMILKEDKLTQHSSERHLR 728
NN L L+ L TMFIN + +M *NK GM D*VI LK E K H S R
Sbjct: 6 NNPLVLKFLINQTMFINYKRLFMV*NKPLGHGMND*VIFFLKKNSPEVKWIPHYS*RESI 185
Query: 729 RTF*LCKYMLMI*YLVLLMHLFAKNSLS*CRMNLK*V*WEN*SSFLEFKSTKVKKEYMFI 788
F KYMLMI*+L LM A++ C++NLK *WEN*S+F ++KS+K+ K Y I
Sbjct: 186 MIFCWFKYMLMI*FLDPLMIHCARSFPLICKVNLKCQ*WEN*STFWDYKSSKLNKVYSSI 365
Query: 789 KQNIQRS 795
N R+
Sbjct: 366 NPNTARN 386
>AI959950
Length = 466
Score = 74.3 bits (181), Expect = 2e-13
Identities = 52/126 (41%), Positives = 69/126 (54%)
Frame = -2
Query: 560 CKKN*ISFREMMCGI*YPNLFRRTLLEQNGYSETS*MKKEK*PETKPDLLLKVTVNKKAL 619
CKKN ISF+ +M R LE NGY T+* + + +TK D LLKVT N+K
Sbjct: 387 CKKNLISFKRIMSRSSLNYQKERR*LE*NGYFVTN*TRMVRL*DTKQD*LLKVTHNRKV* 208
Query: 620 ITLKHLLQLQDWKQSGYFYPMQLTMA*YCIKWMSKVLFLMVSLKKKYMSNNLLGLRILSI 679
T K L L K ++ +Q + * CIKWM KV F M K+K+M NN L L++
Sbjct: 207 TTQKPLHLLHV*K*YASYFHLQPIVI*SCIKWM*KVHF*MA*SKRKFMLNNRLDLKMKPF 28
Query: 680 LTMFIN 685
+ MF+N
Sbjct: 27 INMFLN 10
>BM143109
Length = 415
Score = 63.9 bits (154), Expect = 3e-10
Identities = 49/126 (38%), Positives = 71/126 (55%)
Frame = +2
Query: 670 NLLGLRILSILTMFINLRNHYMA*NKLPELGMID*VIS*LKMILKEDKLTQHSSERHLRR 729
NLL + L M +N + YM *NK LGM *V + ++ +L +
Sbjct: 5 NLL*GKTQKSLIMSLN*KRFYMD*NKPLGLGMNF*VNFF*TRVFQKVRLILTFLFKRN*M 184
Query: 730 TF*LCKYMLMI*YLVLLMHLFAKNSLS*CRMNLK*V*WEN*SSFLEFKSTKVKKEYMFIK 789
+ +YMLMI +LV LM LFAKN L C+MNLK * +*+ FL++KS+K + EY+ +
Sbjct: 185 IYS*YRYMLMILFLVQLMILFAKNFLKICKMNLKCQ*CVS*TFFLDYKSSKQRMEYLSVN 364
Query: 790 QNIQRS 795
QNI ++
Sbjct: 365 QNIAKT 382
>TC211311 weakly similar to UP|O24587 (O24587) Pol protein, partial (15%)
Length = 1213
Score = 45.8 bits (107), Expect(2) = 2e-08
Identities = 65/190 (34%), Positives = 90/190 (47%)
Frame = +1
Query: 728 RRTF*LCKYMLMI*YLVLLMHLFAKNSLS*CRMNLK*V*WEN*SSFLEFKSTKVKKEYMF 787
R+ F L MLM * LV A++ LS* RM+LK V* +*SS +FKS + ++
Sbjct: 478 RKPFLLFISMLMT*SLVQPQKGCARSFLS**RMDLKRV*KVS*SSS*DFKSFRKFMGFLS 657
Query: 788 IKQNIQRSF*RSSS*KIVK**TLQCIQPAP*AKKILEQ**TRSYTEV*LVLCYTSLHLDL 847
IK+NIQ *+ S CI P + + V*L+L + L +D
Sbjct: 658 IKRNIQSPI*KGSEWMKPNLWQPLCIVPQSLTRMRKVIILHKRSIVV*LILYHI*LLVDQ 837
Query: 848 ISYSVYACVQDFNQILENLI*LLLRESSGI*KEQLILDSFIGNP*ITS*LDSVMLIMLVT 907
I +A VQDF+ I + L+ L L+ S I E LI+ + D VM I+LV
Sbjct: 838 ILCLSFAFVQDFSLIQKFLMLLQLKGS*DILLELLIIVYGLRKGLSLIF*DIVMFILLVI 1017
Query: 908 GLKENQPVEI 917
K VE+
Sbjct: 1018K*KGRALVEM 1047
Score = 31.6 bits (70), Expect(2) = 2e-08
Identities = 21/42 (50%), Positives = 25/42 (59%)
Frame = +3
Query: 690 YMA*NKLPELGMID*VIS*LKMILKEDKLTQHSSERHLRRTF 731
+M *NKL ELGM *V * +M E+ T H SER + TF
Sbjct: 363 FMV*NKL*ELGMKG*VHF*FQMDSPEE*RTPHYSERLKKETF 488
>BF596801 weakly similar to GP|29423270|g gag-pol polyprotein {Glycine max},
partial (7%)
Length = 336
Score = 53.9 bits (128), Expect = 3e-07
Identities = 44/111 (39%), Positives = 57/111 (50%)
Frame = +1
Query: 542 LSLKLLKKLSQMMVGY*PCKKN*ISFREMMCGI*YPNLFRRTLLEQNGYSETS*MKKEK* 601
++LK+ KK M++G CKKN* + +E M G * NL LLEQNG+ E +*M
Sbjct: 1 VNLKI*KKP**MIIGSLSCKKN*TNLKETMYGN**KNLKIILLLEQNGFLEIN*MNMV*L 180
Query: 602 PETKPDLLLKVTVNKKALITLKHLLQLQDWKQSGYFYPMQLTMA*YCIKWM 652
E KP K + K+ KH+L LQD K F+ M IKWM
Sbjct: 181 LEIKPG**RKDIIKKRE*TMKKHMLLLQD*KPLECFWHMHP**TLNFIKWM 333
>BI469652 weakly similar to GP|18149115|dbj| reverse transcriptase {Silene
noctiflora}, partial (60%)
Length = 427
Score = 51.6 bits (122), Expect = 2e-06
Identities = 30/67 (44%), Positives = 41/67 (60%)
Frame = +1
Query: 638 YPMQLTMA*YCIKWMSKVLFLMVSLKKKYMSNNLLGLRILSILTMFINLRNHYMA*NKLP 697
+P+QL * KW+ KV+F M LK+K MS+NL L + T+F N + +M *N+
Sbjct: 220 FPLQLIKT*IFFKWILKVVF*MTLLKRKCMSDNLQTL*TIHF*TIFSNFKRLHMV*NRHL 399
Query: 698 ELGMID* 704
LGMID*
Sbjct: 400 MLGMID* 420
>AI855982
Length = 484
Score = 45.1 bits (105), Expect = 2e-04
Identities = 29/58 (50%), Positives = 37/58 (63%)
Frame = +3
Query: 539 FQ*LSLKLLKKLSQMMVGY*PCKKN*ISFREMMCGI*YPNLFRRTLLEQNGYSETS*M 596
+ *L+LK+ KK M+ G PCKKN*I+ +E+MCG * NL EQNG E +*M
Sbjct: 87 YL*LNLKI*KKP**MITG*LPCKKN*INLKEIMCGN**KNLIIILSYEQNGSLEIN*M 260
>TC213445
Length = 705
Score = 38.9 bits (89), Expect = 0.011
Identities = 27/75 (36%), Positives = 41/75 (54%)
Frame = +2
Query: 924 I*YPGQAKDKQQLLCLQQKQNTFQLQVVVHNYFG*NISWKTIRLMLTVFPFFVIILLLSV 983
+*Y G K K L QKQN F L+V++H FG*+ ++ T+ L ++ V I + +
Sbjct: 449 L*YHGIVKSKIVLSYQLQKQNIFLLEVIMHKSFG*DNNFLTMV*NLIIYLSDVTIQVQLI 628
Query: 984 CQRIQFYIQEPSILR 998
+I F E SIL+
Sbjct: 629 YPKIIFCTLEQSILK 673
>AI966222
Length = 430
Score = 32.7 bits (73), Expect(2) = 0.071
Identities = 31/88 (35%), Positives = 43/88 (48%)
Frame = +2
Query: 309 KWPEP*SMKTI*LNIFGQKQLILHVTFKIGSISDLCWRKLHMNSLKEEDPISLTFISLDV 368
+W EP M T L+ K IL+ F+ I D + L MN ++E+P FI L V
Sbjct: 2 RWLEPR*MIT*PLSTSRLK**ILYAIFRTKFI*DPS*KGLPMNYGRDENPTYHIFILLGV 181
Query: 369 LVTS*TLKTI*RNLMPRLKEESF*VTLK 396
V+ *T + I* L ++ E TLK
Sbjct: 182 SVSL*TQRII*EKLTQKVTVEYLLHTLK 265
Score = 22.3 bits (46), Expect(2) = 0.071
Identities = 14/39 (35%), Positives = 23/39 (58%)
Frame = +3
Query: 399 RHTECIIQKHIVLKNLSM*NLMIESLEIKLQSKVKVMQV 437
RH+EC + +LK LS+*+L SL S + ++Q+
Sbjct: 273 RHSECTTPEL*LLKKLSI*DLAKISLIKNY*S*MSLLQI 389
>TC232593 weakly similar to UP|Q9XG91 (Q9XG91) Tpv2-1c protein (Fragment),
partial (16%)
Length = 562
Score = 30.4 bits (67), Expect = 4.1
Identities = 20/47 (42%), Positives = 32/47 (67%)
Frame = +2
Query: 753 NSLS*CRMNLK*V*WEN*SSFLEFKSTKVKKEYMFIKQNIQRSF*RS 799
+S *C++ LK*+ +* FLE +S+KV+ + +K N+Q+ F*RS
Sbjct: 260 SSKK*CKL-LK*LILVS*LIFLELRSSKVRTKC*SVKGNMQKKF*RS 397
>NP004981 early nodulin
Length = 1458
Score = 30.4 bits (67), Expect = 4.1
Identities = 19/52 (36%), Positives = 31/52 (59%), Gaps = 4/52 (7%)
Frame = -1
Query: 148 FLKLASYNWSKDC----LTLIIIQMHFVVHVRKGKL*KALSNLKTLYQPLDP 195
+++ AS+NWS+DC L L +I+++FV + R K SN+ +Q L P
Sbjct: 927 WIQAASFNWSQDCNFVVLHLKLIKVYFVCNSR*SNNRKERSNV*QPHQFLVP 772
>TC221132 weakly similar to UP|O23529 (O23529) RETROTRANSPOSON like protein,
partial (5%)
Length = 799
Score = 29.3 bits (64), Expect = 9.1
Identities = 28/75 (37%), Positives = 35/75 (46%)
Frame = +2
Query: 873 ESSGI*KEQLILDSFIGNP*ITS*LDSVMLIMLVTGLKENQPVEIVNSWEKI*YPGQAKD 932
ESSGI KEQ ILD + I + +ML L T L V + + PG A+
Sbjct: 182 ESSGISKEQ*ILDFNCDHLQINIFVPFMMLTGLTTLLT*GPRVLT*FTLA*VSSPGAARS 361
Query: 933 KQQLLCLQQKQNTFQ 947
L LQQK NT +
Sbjct: 362 NPSLTNLQQKLNTIK 406
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.359 0.160 0.523
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 39,159,239
Number of Sequences: 63676
Number of extensions: 509422
Number of successful extensions: 5139
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 2855
Number of HSP's successfully gapped in prelim test: 191
Number of HSP's that attempted gapping in prelim test: 2203
Number of HSP's gapped (non-prelim): 3167
length of query: 1012
length of database: 12,639,632
effective HSP length: 107
effective length of query: 905
effective length of database: 5,826,300
effective search space: 5272801500
effective search space used: 5272801500
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 64 (29.3 bits)
Medicago: description of AC148240.10


