

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148236.3 + phase: 0 /pseudo
(116 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
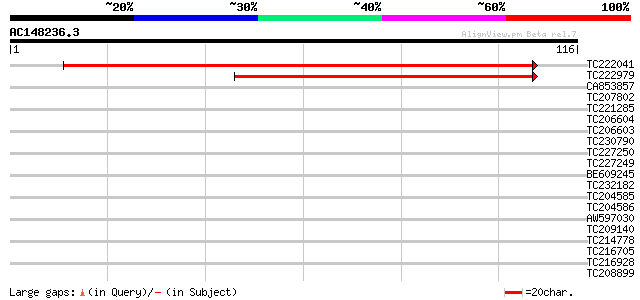
Score E
Sequences producing significant alignments: (bits) Value
TC222041 similar to UP|Q8LBQ6 (Q8LBQ6) Isp4-like protein, partia... 174 4e-45
TC222979 similar to UP|Q8H0V0 (Q8H0V0) Isp4 like protein, partia... 86 2e-18
CA853857 similar to PIR|H96681|H96 protein F1E22.10 [imported] -... 36 0.003
TC207802 similar to UP|Q6R3L0 (Q6R3L0) Metal-nicotianamine trans... 35 0.007
TC221285 similar to UP|Q6R3L0 (Q6R3L0) Metal-nicotianamine trans... 31 0.074
TC206604 similar to UP|Q9LSD4 (Q9LSD4) Gb|AAF23830.1, partial (37%) 31 0.097
TC206603 similar to UP|Q9LSD4 (Q9LSD4) Gb|AAF23830.1, partial (47%) 31 0.097
TC230790 similar to UP|Q9FJC8 (Q9FJC8) EspB-like protein, partia... 30 0.13
TC227250 similar to UP|Q9ZST7 (Q9ZST7) Asparagine synthetase , p... 28 0.82
TC227249 similar to UP|Q8LJT6 (Q8LJT6) Asparagine synthetase , ... 28 0.82
BE609245 similar to GP|6682869|dbj| sigma factor {Nicotiana taba... 27 1.1
TC232182 27 1.8
TC204585 similar to UP|Q9M611 (Q9M611) Soluble inorganic pyropho... 25 4.1
TC204586 homologue to UP|Q9LFF9 (Q9LFF9) Inorganic pyrophosphata... 25 4.1
AW597030 similar to SP|Q9LYT3|TT12 TRANSPARENT TESTA 12 protein.... 25 4.1
TC209140 weakly similar to UP|Q948R9 (Q948R9) RPS6-like protein ... 25 5.3
TC214778 homologue to UP|Q40512 (Q40512) Photosystem I light-har... 25 6.9
TC216705 25 6.9
TC216928 similar to UP|Q9SVA6 (Q9SVA6) GTP-binding-like protein,... 24 9.0
TC208899 similar to UP|Q9LHD1 (Q9LHD1) Multidrug resistance p-gl... 24 9.0
>TC222041 similar to UP|Q8LBQ6 (Q8LBQ6) Isp4-like protein, partial (38%)
Length = 1012
Score = 174 bits (442), Expect = 4e-45
Identities = 77/97 (79%), Positives = 92/97 (94%)
Frame = +2
Query: 12 LALIFTLPVAIITATTNQTPGLNVITEYIMGVILPGRPIANVCFKTYGYMSMSQAISFLS 71
LA FTLP++IITATTNQTPGLN+ITEY+ G+I PGRPIANVCFKTYGY+SM+QA+SFLS
Sbjct: 2 LAFGFTLPISIITATTNQTPGLNIITEYVFGLIYPGRPIANVCFKTYGYISMAQAVSFLS 181
Query: 72 DFKLGHYMKIPPRSMFIVQILGTLIAGTVDVGVAWWL 108
DFKLGHYMKIPPRSMF+VQ +GT++AGT+++GVAWWL
Sbjct: 182 DFKLGHYMKIPPRSMFLVQFIGTILAGTINIGVAWWL 292
>TC222979 similar to UP|Q8H0V0 (Q8H0V0) Isp4 like protein, partial (30%)
Length = 661
Score = 86.3 bits (212), Expect = 2e-18
Identities = 38/62 (61%), Positives = 47/62 (75%)
Frame = +2
Query: 47 GRPIANVCFKTYGYMSMSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTVDVGVAW 106
G+PIAN+ FK YG +S A+SFLSD KLGHYMKIPPR M+ Q+ GTL AG ++ VAW
Sbjct: 2 GKPIANLLFKIYGRISTVHALSFLSDLKLGHYMKIPPRCMYTAQLEGTLFAGVENLAVAW 181
Query: 107 WL 108
W+
Sbjct: 182 WM 187
>CA853857 similar to PIR|H96681|H96 protein F1E22.10 [imported] - Arabidopsis
thaliana, partial (24%)
Length = 531
Score = 35.8 bits (81), Expect = 0.003
Identities = 17/44 (38%), Positives = 23/44 (51%)
Frame = -2
Query: 63 MSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTVDVGVAW 106
+S A DFK G+ PRSMF+ Q+LGT + + V W
Sbjct: 353 VSXASDLTQDFKTGYMTLASPRSMFLSQVLGTAMGCVISPCVFW 222
>TC207802 similar to UP|Q6R3L0 (Q6R3L0) Metal-nicotianamine transporter YSL1,
partial (29%)
Length = 1137
Score = 34.7 bits (78), Expect = 0.007
Identities = 16/46 (34%), Positives = 28/46 (60%)
Frame = +1
Query: 63 MSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTVDVGVAWWL 108
+S + + DFK HY + PR+MFI Q++G + G V ++++L
Sbjct: 31 ISVSCILMQDFKTAHYTRTSPRAMFICQVIG-IAMGCVTAPLSFFL 165
>TC221285 similar to UP|Q6R3L0 (Q6R3L0) Metal-nicotianamine transporter YSL1,
partial (32%)
Length = 945
Score = 31.2 bits (69), Expect = 0.074
Identities = 13/34 (38%), Positives = 21/34 (61%)
Frame = +1
Query: 63 MSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLI 96
+S A + DFK +Y P++MFI Q++GT +
Sbjct: 76 VSVACXLMLDFKTAYYTCTSPKAMFICQLVGTAL 177
>TC206604 similar to UP|Q9LSD4 (Q9LSD4) Gb|AAF23830.1, partial (37%)
Length = 1091
Score = 30.8 bits (68), Expect = 0.097
Identities = 18/62 (29%), Positives = 30/62 (48%)
Frame = +2
Query: 47 GRPIANVCFKTYGYMSMSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTVDVGVAW 106
G IA V ++ A + DFK G+ +SMF+ Q++GT + G V + +
Sbjct: 143 GGVIAGVASCAVMMSIVATAADLMQDFKTGYLTLSSAKSMFVSQLIGTAM-GCVIAPLTF 319
Query: 107 WL 108
W+
Sbjct: 320 WM 325
>TC206603 similar to UP|Q9LSD4 (Q9LSD4) Gb|AAF23830.1, partial (47%)
Length = 1062
Score = 30.8 bits (68), Expect = 0.097
Identities = 18/62 (29%), Positives = 30/62 (48%)
Frame = +1
Query: 47 GRPIANVCFKTYGYMSMSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTVDVGVAW 106
G IA V ++ A + DFK G+ +SMF+ Q++GT + G V + +
Sbjct: 655 GGVIAGVAACAVMMSIVATAADLMQDFKTGYLTLSSAKSMFVSQLIGTAM-GCVIAPLTF 831
Query: 107 WL 108
W+
Sbjct: 832 WM 837
>TC230790 similar to UP|Q9FJC8 (Q9FJC8) EspB-like protein, partial (32%)
Length = 861
Score = 30.4 bits (67), Expect = 0.13
Identities = 15/38 (39%), Positives = 20/38 (52%)
Frame = +2
Query: 63 MSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTV 100
+S + + DFK GH PRSM + Q +GT I V
Sbjct: 71 VSISSDLMHDFKTGHLTFTSPRSMLLSQAIGTAIGCVV 184
>TC227250 similar to UP|Q9ZST7 (Q9ZST7) Asparagine synthetase , partial (96%)
Length = 1916
Score = 27.7 bits (60), Expect = 0.82
Identities = 15/49 (30%), Positives = 25/49 (50%)
Frame = -2
Query: 52 NVCFKTYGYMSMSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTV 100
N+CF YG++S S A+ + I PRS I+ G+++ T+
Sbjct: 1381 NMCFGKYGFLSSSNALR-------NTHFSILPRSGLIIFHSGSILMATL 1256
>TC227249 similar to UP|Q8LJT6 (Q8LJT6) Asparagine synthetase , partial
(30%)
Length = 745
Score = 27.7 bits (60), Expect = 0.82
Identities = 15/49 (30%), Positives = 25/49 (50%)
Frame = -2
Query: 52 NVCFKTYGYMSMSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTV 100
N+CF YG++S S A+ + I PRS I+ G+++ T+
Sbjct: 195 NMCFGKYGFLSSSNALR-------NTHFSILPRSGLIIFHSGSILMATL 70
>BE609245 similar to GP|6682869|dbj| sigma factor {Nicotiana tabacum},
partial (6%)
Length = 375
Score = 27.3 bits (59), Expect = 1.1
Identities = 10/32 (31%), Positives = 19/32 (59%)
Frame = +3
Query: 57 TYGYMSMSQAISFLSDFKLGHYMKIPPRSMFI 88
+Y Y + + +S+ SDF HY +P +S+ +
Sbjct: 276 SYHYSDIIEKLSYGSDFGSTHYQIVPTKSLIV 371
>TC232182
Length = 682
Score = 26.6 bits (57), Expect = 1.8
Identities = 13/34 (38%), Positives = 17/34 (49%)
Frame = -2
Query: 18 LPVAIITATTNQTPGLNVITEYIMGVILPGRPIA 51
LP I T N PG + TEY + ++P P A
Sbjct: 366 LPSCYILNTGNHNPGNSF*TEYFVHTLIPHNPQA 265
>TC204585 similar to UP|Q9M611 (Q9M611) Soluble inorganic pyrophosphatase,
partial (44%)
Length = 523
Score = 25.4 bits (54), Expect = 4.1
Identities = 16/52 (30%), Positives = 25/52 (47%), Gaps = 13/52 (25%)
Frame = -2
Query: 78 YMKIPPR---SMFI----------VQILGTLIAGTVDVGVAWWLPRFNKEHM 116
Y+KI PR +MF ++ G LIAG + G+ WW + ++ M
Sbjct: 339 YLKIMPRVSSNMFPGNG*KNPLVKTRVWGVLIAGNLGWGLNWWSHFYTRQVM 184
>TC204586 homologue to UP|Q9LFF9 (Q9LFF9) Inorganic pyrophosphatase-like
protein, partial (94%)
Length = 1225
Score = 25.4 bits (54), Expect = 4.1
Identities = 14/48 (29%), Positives = 21/48 (43%), Gaps = 13/48 (27%)
Frame = -1
Query: 73 FKLGHYMKIPPR-------------SMFIVQILGTLIAGTVDVGVAWW 107
+ G Y+KI PR + ++ G LIAG + G+ WW
Sbjct: 346 WSFGSYLKIVPRVCSNVSPGNG*KNPLVKTRVWGLLIAGNLVWGLNWW 203
>AW597030 similar to SP|Q9LYT3|TT12 TRANSPARENT TESTA 12 protein. [Mouse-ear
cress] {Arabidopsis thaliana}, partial (25%)
Length = 401
Score = 25.4 bits (54), Expect = 4.1
Identities = 26/95 (27%), Positives = 42/95 (43%), Gaps = 4/95 (4%)
Frame = +1
Query: 6 LIFASGLALIFTLPVAIITATTNQTPGLNVITEYIMGV--ILPGRPIANVCFKTYGYMSM 63
LIF L+ +FT +I A +N TP L I+ + G+ IL G I + Y+++
Sbjct: 115 LIFRVSLSKLFTSDSDVIDAVSNLTP-LLAISVFFNGIQPILSGVAIGSGWQALVAYVNL 291
Query: 64 SQ--AISFLSDFKLGHYMKIPPRSMFIVQILGTLI 96
+ + LG + ++ ILG LI
Sbjct: 292 ASYYVVGLTVGCVLGFKTSLGVAGIWWGMILGVLI 396
>TC209140 weakly similar to UP|Q948R9 (Q948R9) RPS6-like protein
(AT3g17170/K14A17_29), partial (59%)
Length = 1111
Score = 25.0 bits (53), Expect = 5.3
Identities = 22/87 (25%), Positives = 38/87 (43%)
Frame = -3
Query: 4 WALIFASGLALIFTLPVAIITATTNQTPGLNVITEYIMGVILPGRPIANVCFKTYGYMSM 63
+A+I + LIFT+PV II + ++ + +I +C T M +
Sbjct: 854 FAIIAVIAILLIFTIPVIII-----------IFIKFFIIII--------ICICTQ-CMKL 735
Query: 64 SQAISFLSDFKLGHYMKIPPRSMFIVQ 90
+ + SD + Y K+P S +VQ
Sbjct: 734 RRRRAIFSDCLIPLYHKMPNNSFILVQ 654
>TC214778 homologue to UP|Q40512 (Q40512) Photosystem I light-harvesting
chlorophyll a/b-binding protein, partial (84%)
Length = 1202
Score = 24.6 bits (52), Expect = 6.9
Identities = 10/20 (50%), Positives = 14/20 (70%)
Frame = +3
Query: 44 ILPGRPIANVCFKTYGYMSM 63
ILP + +N+CF +Y Y SM
Sbjct: 810 ILP*KKKSNICFNSYLYNSM 869
>TC216705
Length = 637
Score = 24.6 bits (52), Expect = 6.9
Identities = 15/52 (28%), Positives = 26/52 (49%)
Frame = +3
Query: 54 CFKTYGYMSMSQAISFLSDFKLGHYMKIPPRSMFIVQILGTLIAGTVDVGVA 105
CF +G++ +S L ++ K+P R +F+ LG L+ T + G A
Sbjct: 426 CFSPFGHLVISDLKQMLLVISWAYFRKMPFR-LFV--YLGILLITTKNTGAA 572
>TC216928 similar to UP|Q9SVA6 (Q9SVA6) GTP-binding-like protein, partial
(25%)
Length = 354
Score = 24.3 bits (51), Expect = 9.0
Identities = 9/21 (42%), Positives = 15/21 (70%)
Frame = -2
Query: 3 WWALIFASGLALIFTLPVAII 23
+W L +S ++LIF + VAI+
Sbjct: 140 FWVLAISSSMSLIFCITVAIL 78
>TC208899 similar to UP|Q9LHD1 (Q9LHD1) Multidrug resistance
p-glycoprotein, partial (14%)
Length = 983
Score = 24.3 bits (51), Expect = 9.0
Identities = 9/20 (45%), Positives = 15/20 (75%)
Frame = -2
Query: 63 MSQAISFLSDFKLGHYMKIP 82
++ A+ FLSDF+L +M +P
Sbjct: 76 LTSAMCFLSDFRLYDFMSMP 17
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.328 0.142 0.459
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,855,847
Number of Sequences: 63676
Number of extensions: 49391
Number of successful extensions: 343
Number of sequences better than 10.0: 44
Number of HSP's better than 10.0 without gapping: 342
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 343
length of query: 116
length of database: 12,639,632
effective HSP length: 92
effective length of query: 24
effective length of database: 6,781,440
effective search space: 162754560
effective search space used: 162754560
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 51 (24.3 bits)
Medicago: description of AC148236.3


