

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148236.2 - phase: 0 /pseudo
(362 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
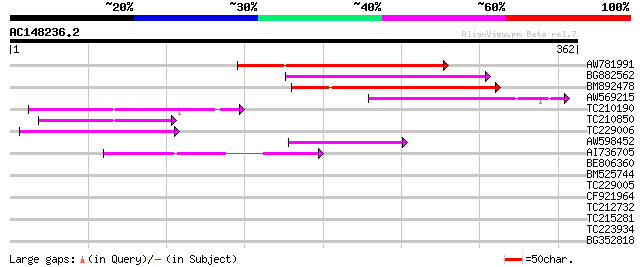
Score E
Sequences producing significant alignments: (bits) Value
AW781991 124 5e-29
BG882562 115 3e-26
BM892478 111 6e-25
AW569215 weakly similar to GP|4914326|gb| F14N23.12 {Arabidopsis... 72 4e-13
TC210190 similar to UP|Q93Z68 (Q93Z68) AT3g59470/T16L24_20, part... 59 3e-09
TC210850 similar to UP|Q93Z68 (Q93Z68) AT3g59470/T16L24_20, part... 56 3e-08
TC229006 51 7e-07
AW598452 weakly similar to GP|15982769|gb| At2g27110/T20P8.16 {A... 48 6e-06
AI736705 similar to GP|11994736|db far-red impaired response pro... 48 8e-06
BE806360 similar to GP|6175165|gb| Mutator-like transposase {Ara... 37 0.014
BM525744 33 0.26
TC229005 32 0.59
CF921964 29 2.9
TC212732 UP|Q93XG4 (Q93XG4) Phytase, complete 29 2.9
TC215281 29 3.8
TC223934 28 5.0
BG352818 28 8.5
>AW781991
Length = 419
Score = 124 bits (312), Expect = 5e-29
Identities = 59/135 (43%), Positives = 87/135 (63%)
Frame = +3
Query: 146 LSKEVGGRTNLGFIRLDQKNYLRKKRERSLVQGEAGYLLQYFQRKTVENPTFYHAYQLDI 205
L KE GG + +GF +D +NY+R R RSL +G+ +L Y ++ ENP F++A Q D
Sbjct: 18 LIKEYGGISKVGFTEVDCRNYMRNNRLRSL-EGDIQLVLDYLRQMHAENPNFFYAVQGDE 194
Query: 206 EDQITNVFWADARMLVDYSYFGDVVSLDSTYCTNSSHRPLAIISGFNHHRGAVIFGAALL 265
+ ITNVFWAD + ++Y++FGD V+ D+TY +N P A +G NHH V+FG A L
Sbjct: 195 DQSITNVFWADPKARMNYTFFGDTVTFDTTYRSNRYRLPFAFFTGVNHHGQPVLFGCAFL 374
Query: 266 YDETAESYEWLFETF 280
+E+ S+ WLF+T+
Sbjct: 375 INESEASFVWLFKTW 419
>BG882562
Length = 416
Score = 115 bits (288), Expect = 3e-26
Identities = 56/131 (42%), Positives = 78/131 (58%)
Frame = +3
Query: 177 QGEAGYLLQYFQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTY 236
+G+ + YF R + NP F++ L+ + Q+ NVFW D+R YSYFGDVV+ DST
Sbjct: 24 KGDPELISNYFCRIQLMNPNFFYVMDLNDDGQLRNVFWIDSRSRAAYSYFGDVVAFDSTC 203
Query: 237 CTNSSHRPLAIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFTDQ 296
+N+ PL G NHH +V+ G LL DET E+Y WLF +L + PQT+ T+
Sbjct: 204 LSNNYEIPLVAFVGVNHHGKSVLLGCGLLADETFETYIWLFRAWLTCMTGRPPQTIITNX 383
Query: 297 AKSMAKALAEV 307
K+M A+AEV
Sbjct: 384 CKAMQSAIAEV 416
>BM892478
Length = 429
Score = 111 bits (277), Expect = 6e-25
Identities = 54/133 (40%), Positives = 81/133 (60%)
Frame = +3
Query: 181 GYLLQYFQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTYCTNS 240
G LL+YFQ + E+ F++A ++D N+FWAD R S+FGDV+ LD++Y
Sbjct: 24 GILLEYFQSRQAEDTGFFYAMEVD-NGNCMNIFWADGRSRYSCSHFGDVLVLDTSYRKTV 200
Query: 241 SHRPLAIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFTDQAKSM 300
P A G NHH+ V+ G AL+ DE+ ES+ WLF+T+L A ++P TV DQ ++
Sbjct: 201 YLVPFATFVGVNHHKQPVLLGCALIADESEESFTWLFQTWLRAMSGRLPLTVIADQDIAI 380
Query: 301 AKALAEVMPEAYH 313
+A+A+V P +H
Sbjct: 381 QRAIAKVFPVTHH 419
>AW569215 weakly similar to GP|4914326|gb| F14N23.12 {Arabidopsis thaliana},
partial (19%)
Length = 425
Score = 72.0 bits (175), Expect = 4e-13
Identities = 41/130 (31%), Positives = 65/130 (49%), Gaps = 2/130 (1%)
Frame = +1
Query: 230 VSLDSTYCTNSSHRPLAIISGFNHHRGAVIFGAALLYDETAESYEWLFETFLEAHKQKMP 289
V D+TY + L I G +++ F ALL DE +S+ W + FL K K P
Sbjct: 10 VVFDTTYRVEAYDMLLGIWLGVDNNGMTCFFSCALLRDENIQSFSWALKAFLGFMKGKAP 189
Query: 290 QTVFTDQAKSMAKALAEVMPEAYHGLCTWHLMQNGIKHLGNLMKGESY--FLSDFEKCMY 347
QT+ TD + +A+A +P+ H C WH++ +L+ G Y + ++F + +Y
Sbjct: 190 QTILTDHNMWLKEAIAVELPQTKHAFCIWHILSK-FSDWFSLLLGSQYDEWKAEFHR-LY 363
Query: 348 GYEDVEQFED 357
E VE FE+
Sbjct: 364 NLEQVEDFEE 393
>TC210190 similar to UP|Q93Z68 (Q93Z68) AT3g59470/T16L24_20, partial (65%)
Length = 866
Score = 58.9 bits (141), Expect = 3e-09
Identities = 48/159 (30%), Positives = 75/159 (46%), Gaps = 21/159 (13%)
Frame = +1
Query: 13 FCSLKNAWEFWNTYXGKXGFGVRKDYTHR-KKDGSVSSCRFVCCKEGMRKPDKRDYKTKN 71
F S A F+N Y + GF VR R ++DG+ VC KEG R DKR+ K
Sbjct: 7 FGSEAAAHAFYNAYATEVGFIVRVSKLSRSRRDGTAIGRTLVCNKEGFRMADKRE-KIVR 183
Query: 72 PRLETRTNCEARLGLKNV-DGKLMVHDFVEDHNHELL-------------------LPET 111
R ETR C A + ++ + GK +V FV++H H L + E
Sbjct: 184 QRAETRVGCRAMIMVRKLSSGKWVVAKFVKEHTHPLTPGKGRRDFYYEQYPTEQDKIREL 363
Query: 112 THMLSSQRKVSEIHCQQIELADSVGVQQKKSFDLLSKEV 150
+ L++++K S + + +EL + Q ++ D LSK++
Sbjct: 364 SQQLATEKKRSAAYKRHLEL---IFEQIEEQNDNLSKKI 471
>TC210850 similar to UP|Q93Z68 (Q93Z68) AT3g59470/T16L24_20, partial (63%)
Length = 1326
Score = 55.8 bits (133), Expect = 3e-08
Identities = 34/90 (37%), Positives = 47/90 (51%), Gaps = 2/90 (2%)
Frame = +1
Query: 19 AWEFWNTYXGKXGFGVRKDYTHR-KKDGSVSSCRFVCCKEGMRKPDKRDYKTKNPRLETR 77
A F+N Y GF VR R ++DG+ VC KEG R DKR+ K R ETR
Sbjct: 7 AHAFYNAYATDVGFIVRVSKLSRSRRDGTAIGRTLVCNKEGFRMADKRE-KIVRQRAETR 183
Query: 78 TNCEARLGLKNV-DGKLMVHDFVEDHNHEL 106
C A + ++ + GK ++ FV++H H L
Sbjct: 184 VGCRAMIMVRKLSSGKWVIAKFVKEHTHPL 273
>TC229006
Length = 991
Score = 51.2 bits (121), Expect = 7e-07
Identities = 35/105 (33%), Positives = 55/105 (52%), Gaps = 3/105 (2%)
Frame = +2
Query: 7 PKVDMFFCSLKNAWEFWNTYXGKXGFGVRKDYTHR-KKDGSVSSCRFVCCKEGMRKPDKR 65
P M F S A F++ Y + GF +R R ++DG + + R C KEG +
Sbjct: 101 PYEGMEFESEDAAKIFYDEYARRLGFVMRVMSCRRPERDGRILARRLGCNKEGYCVSIRG 280
Query: 66 DYKT-KNPRLETRTNCEARLGLK-NVDGKLMVHDFVEDHNHELLL 108
+ + + PR TR C+A + +K N GK ++ FV+DHNH L++
Sbjct: 281 KFASVRKPRASTREGCKAMIHIKYNKSGKWVITKFVKDHNHPLVV 415
>AW598452 weakly similar to GP|15982769|gb| At2g27110/T20P8.16 {Arabidopsis
thaliana}, partial (7%)
Length = 256
Score = 48.1 bits (113), Expect = 6e-06
Identities = 27/76 (35%), Positives = 40/76 (52%)
Frame = +2
Query: 179 EAGYLLQYFQRKTVENPTFYHAYQLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTYCT 238
+A LL+Y +R EN ++A QLD +D+++NV A R Y + GD SLD+ Y
Sbjct: 17 DAHNLLEYCKRMQAENHGCFYAIQLDEDDRVSNVL*AGTRSRTSYKHMGDAESLDAAYGK 196
Query: 239 NSSHRPLAIISGFNHH 254
+ A +G N H
Sbjct: 197 HRYAVSYAPYTGRNCH 244
>AI736705 similar to GP|11994736|db far-red impaired response protein;
Mutator-like transposase-like protein; phytochrome A
signaling, partial (14%)
Length = 383
Score = 47.8 bits (112), Expect = 8e-06
Identities = 38/142 (26%), Positives = 65/142 (45%), Gaps = 2/142 (1%)
Frame = +3
Query: 61 KPDKRDYKTKNPRLE-TRTNCEARLGLKN-VDGKLMVHDFVEDHNHELLLPETTHMLSSQ 118
+ +K+D + R ++T+C+A + +K DGK ++H FV++HNHE LLP ++
Sbjct: 27 RQNKQDSENSTGRRSCSKTDCKASMHVKRRSDGKWVIHSFVKEHNHE-LLPAQAVSEQTR 203
Query: 119 RKVSEIHCQQIELADSVGVQQKKSFDLLSKEVGGRTNLGFIRLDQKNYLRKKRERSLVQG 178
R + + Q E VG++ ++KN K R L G
Sbjct: 204 RMYAAMARQFAEYKTVVGLK-----------------------NEKNPFDKGRNLGLESG 314
Query: 179 EAGYLLQYFQRKTVENPTFYHA 200
EA +L +F + N F++A
Sbjct: 315 EAKLMLDFFIQMQNMNSNFFYA 380
>BE806360 similar to GP|6175165|gb| Mutator-like transposase {Arabidopsis
thaliana}, partial (17%)
Length = 431
Score = 37.0 bits (84), Expect = 0.014
Identities = 19/70 (27%), Positives = 32/70 (45%), Gaps = 4/70 (5%)
Frame = +2
Query: 267 DETAESYEWLFE---TFLEAHKQKMPQ-TVFTDQAKSMAKALAEVMPEAYHGLCTWHLMQ 322
+E +++ W LE H + M + T+ +D+ K + + P A+HG C HL
Sbjct: 20 EENDDNWMWFLSELHNLLEIHTENMLRLTILSDRQKGIVDGVEASFPTAFHGFCMQHLSD 199
Query: 323 NGIKHLGNLM 332
+ K N M
Sbjct: 200 SFRKEFNNTM 229
>BM525744
Length = 411
Score = 32.7 bits (73), Expect = 0.26
Identities = 22/70 (31%), Positives = 31/70 (43%)
Frame = +3
Query: 202 QLDIEDQITNVFWADARMLVDYSYFGDVVSLDSTYCTNSSHRPLAIISGFNHHRGAVIFG 261
++D D I ++FWA + F V+ LD+TY N PL I G + F
Sbjct: 204 KVDDSDAIRDIFWAHPDAIKLLGSFHTVLFLDNTYKVNRYQLPLLEIVGVTSTE--LTFS 377
Query: 262 AALLYDETAE 271
A Y E+ E
Sbjct: 378 VAFAYMESDE 407
>TC229005
Length = 679
Score = 31.6 bits (70), Expect = 0.59
Identities = 14/34 (41%), Positives = 23/34 (67%), Gaps = 1/34 (2%)
Frame = +1
Query: 76 TRTNCEARLGLK-NVDGKLMVHDFVEDHNHELLL 108
TR C+A + +K + GK ++ FV+DHNH L++
Sbjct: 7 TREGCKAMIHIKYDKSGKWVIAKFVKDHNHPLVV 108
>CF921964
Length = 836
Score = 29.3 bits (64), Expect = 2.9
Identities = 19/77 (24%), Positives = 35/77 (44%)
Frame = +1
Query: 47 VSSCRFVCCKEGMRKPDKRDYKTKNPRLETRTNCEARLGLKNVDGKLMVHDFVEDHNHEL 106
+S V C G K D +Y+ + E + ++ L ++ G++ V DF+ +
Sbjct: 322 ISVLLLVSCSNGQIK-DALNYQIEPFEFENQNG--QKVSLDDLKGEIWVADFIFTSCETI 492
Query: 107 LLPETTHMLSSQRKVSE 123
P T HM Q+++ E
Sbjct: 493 CPPMTAHMTELQKRLKE 543
>TC212732 UP|Q93XG4 (Q93XG4) Phytase, complete
Length = 1644
Score = 29.3 bits (64), Expect = 2.9
Identities = 13/37 (35%), Positives = 19/37 (51%)
Frame = +1
Query: 258 VIFGAALLYDETAESYEWLFETFLEAHKQKMPQTVFT 294
++ GA + YD+TAE Y+WL + P V T
Sbjct: 985 IMLGAYINYDKTAEQYKWLERDLENVDRSITPWLVVT 1095
>TC215281
Length = 733
Score = 28.9 bits (63), Expect = 3.8
Identities = 12/36 (33%), Positives = 22/36 (60%)
Frame = -2
Query: 163 QKNYLRKKRERSLVQGEAGYLLQYFQRKTVENPTFY 198
Q+ +LRKK++ + G LLQ+F ++ + P F+
Sbjct: 537 QRMHLRKKKQVKQI*GNKSLLLQFFAKRPPKPPFFF 430
>TC223934
Length = 889
Score = 28.5 bits (62), Expect = 5.0
Identities = 21/78 (26%), Positives = 34/78 (42%)
Frame = +2
Query: 261 GAALLYDETAESYEWLFETFLEAHKQKMPQTVFTDQAKSMAKALAEVMPEAYHGLCTWHL 320
G+ ++D+ E + + T + AH + D + A L + MPE + TW+
Sbjct: 419 GSRRVFDDMPERDVFAWTTMISAHVR--------DGDMASAGRLFDEMPEK--NVATWNA 568
Query: 321 MQNGIKHLGNLMKGESYF 338
M +G LGN E F
Sbjct: 569 MIDGYGKLGNAESAEFLF 622
>BG352818
Length = 265
Score = 27.7 bits (60), Expect = 8.5
Identities = 9/23 (39%), Positives = 13/23 (56%)
Frame = -1
Query: 304 LAEVMPEAYHGLCTWHLMQNGIK 326
L PEA+HG+ WH Q ++
Sbjct: 223 LPRAEPEAFHGVSDWHAWQRSVR 155
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.136 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,211,322
Number of Sequences: 63676
Number of extensions: 236429
Number of successful extensions: 1079
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 1068
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1075
length of query: 362
length of database: 12,639,632
effective HSP length: 98
effective length of query: 264
effective length of database: 6,399,384
effective search space: 1689437376
effective search space used: 1689437376
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 59 (27.3 bits)
Medicago: description of AC148236.2


