

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148154.9 + phase: 0 /pseudo
(481 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
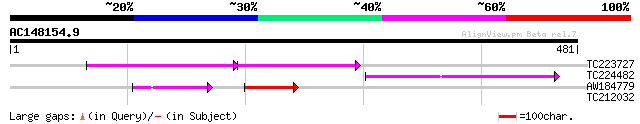
TC223727 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 52 2e-14
TC224482 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 61 1e-09
AW184779 43 9e-06
TC212032 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 32 0.82
>TC223727 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (9%)
Length = 843
Score = 52.4 bits (124), Expect(2) = 2e-14
Identities = 44/129 (34%), Positives = 67/129 (51%)
Frame = +3
Query: 66 *TLVLRHQTVLAEP*VSAGCFQARQEDLEKIGQ*IPVRWRYSVQEKL*HGIVKMC**T*S 125
* LV RHQ + + + A + ED+EK+G +++V EK H +C
Sbjct: 132 *ALVFRHQAICHKQRIPARDCRQ**EDIEKVGGRFLHERKHTV*EKPRHETSAVCGCQGG 311
Query: 126 GAVNA*CT*RYLRDPCYRAYYVKEVVTSRLLLDDHGA*LLPIRQKMPQMSNLC**DSCAS 185
+ + * + + RA Y +E SRLLL HG *LL ++MPQMS++ * C +
Sbjct: 312 KSHDRGSP*GLVWNARQRACYGQEDPKSRLLLAYHGK*LLCPCEEMPQMSSVRR*CQCPT 491
Query: 186 ACSQCYVIP 194
A S+C+V+P
Sbjct: 492 ASSECHVLP 518
Score = 44.7 bits (104), Expect(2) = 2e-14
Identities = 39/104 (37%), Positives = 47/104 (44%)
Frame = +2
Query: 194 PQGRSQCGAST*LEELNRRLQMVIVSS*WQLTTSPSGLKQHLIPM*PSKW*LSSSKTTSS 253
P G S CG L+ R +MVI SS + SPSG +Q IP W S + SS
Sbjct: 515 PLGLSPCGE*MSSGPLSPRPRMVIASSS*R*IISPSGSRQLPIPTS*GVWWSGSLRKRSS 694
Query: 254 VDMVFPARLLPTMVPT*TIMWCKLFVKNSKLSIITLLPIDLR*M 297
DMV RL T PT* F ++ K SI P R*+
Sbjct: 695 ADMVCQGRLSRTTAPT*ITR*WGKFARSLKSSITIPXPTGQR*I 826
>TC224482 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 669
Score = 60.8 bits (146), Expect = 1e-09
Identities = 60/165 (36%), Positives = 80/165 (48%), Gaps = 1/165 (0%)
Frame = +2
Query: 303 PTRISRELSKRW*PLTRTGMRCYPMLCMVTVLQCAVQPGQPLFLLYMVWKQFFLWK*RSR 362
P RIS+ LSKR TR G RC VT LQC Q GQ YM W+ + +*+SR
Sbjct: 8 PIRISKRLSKR*PCHTRIGTRCSHSRYTVTGLQCERQLGQRRSHWYMGWRLCYRLR*KSR 187
Query: 363 LSG*SWKQSYLR-LNGAKAGMIS*I*LRKNVWMPWLVDIHIKQE*RLLLTRKFILENSR* 421
G SW+ R +G K MIS LR + P ++ +E*R+ TR++ +S
Sbjct: 188 H*G-SWQNPD*RNQSGLKRAMISSTSLRVSA*RP*VMGACTSKE*RVHSTRRYACASSMR 364
Query: 422 GNLY*KEG*ASNPTQGASGHLTMKVLMLSRRPSPVVL*SLHTWMV 466
L *++ + T +G T K L+L R P L TWMV
Sbjct: 365 ETLC*RKCPMLSRTIEGNGPRTTKGLLL*RGLFPEEPWCLPTWMV 499
>AW184779
Length = 432
Score = 42.7 bits (99), Expect(2) = 9e-06
Identities = 27/46 (58%), Positives = 31/46 (66%)
Frame = +1
Query: 200 CGAST*LEELNRRLQMVIVSS*WQLTTSPSGLKQHLIPM*PSKW*L 245
CGA T* E L+ RLQM I S * QLTTSP+G KQ + +* W*L
Sbjct: 292 CGA*T*SEPLSPRLQMDITSF*SQLTTSPNGSKQFRMLV*LGVW*L 429
Score = 24.6 bits (52), Expect(2) = 9e-06
Identities = 24/69 (34%), Positives = 33/69 (47%), Gaps = 1/69 (1%)
Frame = +3
Query: 105 RYSVQEKL*HGIVKMC**T*SGAVNA-*CT*RYLRDPCYRAYYVKEVVTSRLLLDDHGA* 163
+Y +QE+ *HG+ MC * G NA * L + C + E S +LL +G
Sbjct: 9 KYPIQEEP*HGVASMC-GC*GG*ANAGRGA*GILWNTCQHTCHGPEDSESGVLLAHYGER 185
Query: 164 LLPIRQKMP 172
LL +MP
Sbjct: 186 LLHPCVEMP 212
>TC212032 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (3%)
Length = 803
Score = 31.6 bits (70), Expect = 0.82
Identities = 30/101 (29%), Positives = 48/101 (46%)
Frame = +1
Query: 15 NG*CSCYSIFHVPSKPLE*RAINQSTTPRKTFACVCYWGCDRSGW*KCG*L*TLVLRHQT 74
NG C+C+ HVP+ ++ +T + SG G * LV +Q
Sbjct: 397 NGRCACHFSVHVPANTAWGPTVH*V*VSWQTRTLL-------SGGRGTG-R*ALVF*YQA 552
Query: 75 VLAEP*VSAGCFQARQEDLEKIGQ*IPVRWRYSVQEKL*HG 115
+ + V G F+ RQ++ E+IG + +++VQEK *HG
Sbjct: 553 IR*KQRVPTGGFRQRQKEKEEIGSRLLHERKHTVQEKP*HG 675
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.364 0.161 0.620
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 22,571,793
Number of Sequences: 63676
Number of extensions: 336856
Number of successful extensions: 4613
Number of sequences better than 10.0: 8
Number of HSP's better than 10.0 without gapping: 2522
Number of HSP's successfully gapped in prelim test: 156
Number of HSP's that attempted gapping in prelim test: 2050
Number of HSP's gapped (non-prelim): 2768
length of query: 481
length of database: 12,639,632
effective HSP length: 101
effective length of query: 380
effective length of database: 6,208,356
effective search space: 2359175280
effective search space used: 2359175280
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC148154.9


