

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC148098.5 + phase: 0
(189 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
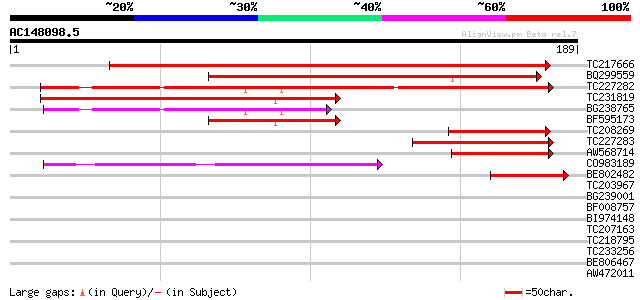
Score E
Sequences producing significant alignments: (bits) Value
TC217666 similar to GB|AAD14482.1|4249385|T2K10 T2K10.11 {Arabid... 284 2e-77
BQ299559 similar to GP|4249385|gb|A T2K10.11 {Arabidopsis thalia... 200 4e-52
TC227282 homologue to GB|AAD14482.1|4249385|T2K10 T2K10.11 {Arab... 191 2e-49
TC231819 similar to GB|AAD14482.1|4249385|T2K10 T2K10.11 {Arabid... 178 2e-45
BG238765 similar to GP|4249385|gb|A T2K10.11 {Arabidopsis thalia... 89 1e-18
BF595173 homologue to GP|4249385|gb|A T2K10.11 {Arabidopsis thal... 85 2e-17
TC208269 weakly similar to GB|AAD14482.1|4249385|T2K10 T2K10.11 ... 59 1e-09
TC227283 homologue to GB|AAD14482.1|4249385|T2K10 T2K10.11 {Arab... 57 4e-09
AW568714 homologue to GP|4249385|gb|A T2K10.11 {Arabidopsis thal... 49 2e-06
CO983189 47 4e-06
BE802482 41 3e-04
TC203967 homologue to UP|Q6UCJ3 (Q6UCJ3) Ubiquitin extension pro... 35 0.016
BG239001 similar to PIR|T52334|T52 ubiquitin extension protein [... 35 0.016
BF008757 homologue to GP|4249385|gb|A T2K10.11 {Arabidopsis thal... 35 0.027
BI974148 homologue to GP|4249385|gb|A T2K10.11 {Arabidopsis thal... 30 0.075
TC207163 33 0.10
TC218795 similar to UP|Q9JL05 (Q9JL05) BETA3 (Basic helix-loop-h... 32 0.18
TC233256 weakly similar to UP|HUL5_YEAST (P53119) Probable ubiqu... 31 0.30
BE806467 30 0.51
AW472011 similar to SP|P29530|OLE1 P24 oleosin isoform A (P89). ... 30 0.51
>TC217666 similar to GB|AAD14482.1|4249385|T2K10 T2K10.11 {Arabidopsis
thaliana;} , partial (37%)
Length = 1372
Score = 284 bits (727), Expect = 2e-77
Identities = 125/147 (85%), Positives = 140/147 (95%)
Frame = +1
Query: 34 AVFWRVVPRIFPPPRWEFGGTSLDRSKGNKRNWIIVWEDGFCDFNECEQRKSGCLNERFG 93
AVFWR+VPR FPPPRWEFGGT+LDRSKGNKRNWI+VWEDGFCDFNECEQ ++G LN +FG
Sbjct: 1 AVFWRIVPRNFPPPRWEFGGTALDRSKGNKRNWILVWEDGFCDFNECEQGRNGYLNYKFG 180
Query: 94 ADVFFKMSHEVYSYGEGLVGKVAADNGHKWVYSDTQNGCEANYVGPWNASIDHQPRAWEF 153
ADVFFKMSHEVYSYGEGL+GKVAADN HKWVYSDTQNGCE++Y+G WNAS+DHQP+ WEF
Sbjct: 181 ADVFFKMSHEVYSYGEGLMGKVAADNNHKWVYSDTQNGCESSYIGAWNASMDHQPKTWEF 360
Query: 154 QLNSGIQTIAVIAVKEGLVQLGSFDKV 180
QLNSGIQTIAVIAV+EGLVQLGSF+K+
Sbjct: 361 QLNSGIQTIAVIAVREGLVQLGSFNKI 441
>BQ299559 similar to GP|4249385|gb|A T2K10.11 {Arabidopsis thaliana}, partial
(10%)
Length = 422
Score = 200 bits (508), Expect = 4e-52
Identities = 95/140 (67%), Positives = 106/140 (74%), Gaps = 29/140 (20%)
Frame = +2
Query: 67 IIVWEDGFCDFNECEQRKSGCLNERFGADVFFKMSHEVYSYGEGLVGKVAADNGHKWVYS 126
I+VWEDGFCDFNECEQ ++GCLN +FGADVFFKMSHEVYSYGEGL+GKVAADN HKWVYS
Sbjct: 2 ILVWEDGFCDFNECEQGRNGCLNYKFGADVFFKMSHEVYSYGEGLMGKVAADNNHKWVYS 181
Query: 127 DTQNGCEANYVGPWNASIDH-----------------------------QPRAWEFQLNS 157
DTQNGCE++Y+G WNAS+DH QP+ WEFQLNS
Sbjct: 182 DTQNGCESSYIGAWNASMDHVREVANNKCKKKNTWFIFLVL*LDI*S*QQPKTWEFQLNS 361
Query: 158 GIQTIAVIAVKEGLVQLGSF 177
GIQTIAVIAV+EGLVQLGSF
Sbjct: 362 GIQTIAVIAVREGLVQLGSF 421
>TC227282 homologue to GB|AAD14482.1|4249385|T2K10 T2K10.11 {Arabidopsis
thaliana;} , partial (61%)
Length = 1408
Score = 191 bits (484), Expect = 2e-49
Identities = 95/186 (51%), Positives = 128/186 (68%), Gaps = 15/186 (8%)
Frame = +2
Query: 11 LLQHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIFPPPRWEFGGTSLDRSKGNKRNWIIVW 70
LLQHTLR+LC +S+WVYAVFWR++PR +PPP+WE G + DRS+GN+RNWI+VW
Sbjct: 122 LLQHTLRSLCIHE----NSQWVYAVFWRILPRNYPPPKWE-GQGAYDRSRGNRRNWILVW 286
Query: 71 EDGFCDF--NECEQRKSGCLN-------------ERFGADVFFKMSHEVYSYGEGLVGKV 115
EDGFC+F + + SG + + ++FFKMSHE+Y+YGEGL+GKV
Sbjct: 287 EDGFCNFAASAAPEINSGDCSTPPAYGNCEFQPYQGLQPELFFKMSHEIYNYGEGLIGKV 466
Query: 116 AADNGHKWVYSDTQNGCEANYVGPWNASIDHQPRAWEFQLNSGIQTIAVIAVKEGLVQLG 175
AAD+ HKW+ + N E N++ W+ S D PR WE Q SGI+TIA+IAV+EG+VQLG
Sbjct: 467 AADHSHKWINKEP-NDQEINFLSAWHNSADSHPRTWEAQFLSGIKTIALIAVREGVVQLG 643
Query: 176 SFDKVL 181
+ KV+
Sbjct: 644 AVHKVI 661
>TC231819 similar to GB|AAD14482.1|4249385|T2K10 T2K10.11 {Arabidopsis
thaliana;} , partial (16%)
Length = 460
Score = 178 bits (451), Expect = 2e-45
Identities = 80/102 (78%), Positives = 89/102 (86%), Gaps = 2/102 (1%)
Frame = +1
Query: 11 LLQHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIFPPPRWEFGGTSLDRSKGNKRNWIIVW 70
LLQ TLR +C+ P SST SKWVYAVFWR++PR FPPPRWE GGT+LD SKGNKRNWI+VW
Sbjct: 1 LLQQTLRTICTSPNSSTPSKWVYAVFWRILPRNFPPPRWELGGTALDLSKGNKRNWILVW 180
Query: 71 EDGFCDFNECEQRKSGC--LNERFGADVFFKMSHEVYSYGEG 110
EDGFC FNECEQRKSG LN RFGA++FFKMSHEVY+YGEG
Sbjct: 181 EDGFCYFNECEQRKSGSGYLNGRFGAELFFKMSHEVYNYGEG 306
>BG238765 similar to GP|4249385|gb|A T2K10.11 {Arabidopsis thaliana}, partial
(18%)
Length = 349
Score = 89.0 bits (219), Expect = 1e-18
Identities = 49/111 (44%), Positives = 64/111 (57%), Gaps = 15/111 (13%)
Frame = +2
Query: 12 LQHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIFPPPRWEFGGTSLDRSKGNKRNWIIVWE 71
LQHTLR+LC +S+WVYAVF R++PR +PPP+WE G + D S+GN+ NWI+VWE
Sbjct: 32 LQHTLRSLCIHE----NSQWVYAVFLRILPRNYPPPKWE-GQRAYDISRGNRWNWILVWE 196
Query: 72 DGFCDF---NECEQRKSGCLN------------ERFGADVFFKMSHEVYSY 107
DGFC+F + E C +R FF MSH +Y Y
Sbjct: 197 DGFCNFAASSAPEINSRDCSTPPAYGNCEFHPYQRLQPYHFFYMSHYIYIY 349
>BF595173 homologue to GP|4249385|gb|A T2K10.11 {Arabidopsis thaliana},
partial (6%)
Length = 317
Score = 84.7 bits (208), Expect = 2e-17
Identities = 38/46 (82%), Positives = 42/46 (90%), Gaps = 2/46 (4%)
Frame = +3
Query: 67 IIVWEDGFCDFNECEQRKSGC--LNERFGADVFFKMSHEVYSYGEG 110
I+VWEDGFCDFNECEQRKSG LN RFGA++FFKMSHEVY+YGEG
Sbjct: 78 ILVWEDGFCDFNECEQRKSGSGYLNGRFGAELFFKMSHEVYNYGEG 215
>TC208269 weakly similar to GB|AAD14482.1|4249385|T2K10 T2K10.11 {Arabidopsis
thaliana;} , partial (9%)
Length = 800
Score = 59.3 bits (142), Expect = 1e-09
Identities = 25/34 (73%), Positives = 32/34 (93%)
Frame = +3
Query: 147 QPRAWEFQLNSGIQTIAVIAVKEGLVQLGSFDKV 180
QP+AWEFQ NSGIQ+I +IAV+EG+VQLGSF+K+
Sbjct: 21 QPKAWEFQFNSGIQSIVIIAVREGVVQLGSFNKI 122
>TC227283 homologue to GB|AAD14482.1|4249385|T2K10 T2K10.11 {Arabidopsis
thaliana;} , partial (27%)
Length = 1192
Score = 57.4 bits (137), Expect = 4e-09
Identities = 26/47 (55%), Positives = 35/47 (74%)
Frame = +2
Query: 135 NYVGPWNASIDHQPRAWEFQLNSGIQTIAVIAVKEGLVQLGSFDKVL 181
N++ W+ S D PR WE Q SGI+TIA+IAV+EG+VQLG+ KV+
Sbjct: 2 NFLTAWHNSADSHPRTWEAQFLSGIKTIALIAVREGVVQLGAVHKVI 142
>AW568714 homologue to GP|4249385|gb|A T2K10.11 {Arabidopsis thaliana},
partial (8%)
Length = 393
Score = 48.5 bits (114), Expect = 2e-06
Identities = 23/34 (67%), Positives = 28/34 (81%)
Frame = +2
Query: 148 PRAWEFQLNSGIQTIAVIAVKEGLVQLGSFDKVL 181
PR WE Q SGI+TIA+IAV+EG+VQLG+ KVL
Sbjct: 242 PRTWEAQFLSGIKTIALIAVREGVVQLGAVHKVL 343
>CO983189
Length = 770
Score = 47.4 bits (111), Expect = 4e-06
Identities = 32/113 (28%), Positives = 47/113 (41%)
Frame = +2
Query: 12 LQHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIFPPPRWEFGGTSLDRSKGNKRNWIIVWE 71
+ H L+ C ++W YA FW++ WE G D K + +W
Sbjct: 380 IMHLLKGFCDH------TQWKYAGFWKLDQHFPMTLTWENGYQKRDEVKES------MWG 523
Query: 72 DGFCDFNECEQRKSGCLNERFGADVFFKMSHEVYSYGEGLVGKVAADNGHKWV 124
D + SG ++ + +MSH YS GEG+VGK+A H WV
Sbjct: 524 DLSFKSPDELYSSSGENSDYSARLLLIEMSHRXYSLGEGVVGKIALARDHCWV 682
>BE802482
Length = 405
Score = 41.2 bits (95), Expect = 3e-04
Identities = 21/26 (80%), Positives = 24/26 (91%)
Frame = +2
Query: 161 TIAVIAVKEGLVQLGSFDKVLLFFPS 186
TIAVIAV+EGLVQLGSF+KVL+ F S
Sbjct: 95 TIAVIAVREGLVQLGSFNKVLVSFIS 172
>TC203967 homologue to UP|Q6UCJ3 (Q6UCJ3) Ubiquitin extension protein,
partial (98%)
Length = 746
Score = 35.4 bits (80), Expect = 0.016
Identities = 15/27 (55%), Positives = 19/27 (69%)
Frame = -2
Query: 5 LPLLHSLLQHTLRNLCSFPTSSTSSKW 31
+P HS L H+LR+LC+FP SST W
Sbjct: 457 VPAPHSALGHSLRSLCTFPESSTL*NW 377
>BG239001 similar to PIR|T52334|T52 ubiquitin extension protein [imported]
- potato, partial (80%)
Length = 379
Score = 35.4 bits (80), Expect = 0.016
Identities = 15/27 (55%), Positives = 19/27 (69%)
Frame = +3
Query: 5 LPLLHSLLQHTLRNLCSFPTSSTSSKW 31
+P HS L H+LR+LC+FP SST W
Sbjct: 72 VPAPHSALGHSLRSLCTFPKSSTL*NW 152
>BF008757 homologue to GP|4249385|gb|A T2K10.11 {Arabidopsis thaliana},
partial (8%)
Length = 177
Score = 34.7 bits (78), Expect = 0.027
Identities = 15/33 (45%), Positives = 21/33 (63%)
Frame = +1
Query: 113 GKVAADNGHKWVYSDTQNGCEANYVGPWNASID 145
GKVAAD+ HKW+ + N E N++ W+ S D
Sbjct: 1 GKVAADHSHKWINKE-PNDQEINFLSAWHNSAD 96
>BI974148 homologue to GP|4249385|gb|A T2K10.11 {Arabidopsis thaliana},
partial (8%)
Length = 424
Score = 30.0 bits (66), Expect(2) = 0.075
Identities = 14/20 (70%), Positives = 18/20 (90%)
Frame = +1
Query: 162 IAVIAVKEGLVQLGSFDKVL 181
IA+IAV+EG+VQLG+ KVL
Sbjct: 199 IALIAVREGVVQLGAVHKVL 258
Score = 21.9 bits (45), Expect(2) = 0.075
Identities = 8/15 (53%), Positives = 10/15 (66%)
Frame = +2
Query: 148 PRAWEFQLNSGIQTI 162
PR WE Q SGI+ +
Sbjct: 68 PRTWEAQFLSGIKVL 112
>TC207163
Length = 1351
Score = 32.7 bits (73), Expect = 0.10
Identities = 17/54 (31%), Positives = 24/54 (43%), Gaps = 11/54 (20%)
Frame = +1
Query: 24 TSSTSSKWV----------YAVFWRV-VPRIFPPPRWEFGGTSLDRSKGNKRNW 66
T ST+++W+ + FW VP P W F GTS +S G+ W
Sbjct: 1015 TESTTTRWISAKAATAATAFTAFWTPWVP*FLPISDWHFAGTSATKSSGSILGW 1176
>TC218795 similar to UP|Q9JL05 (Q9JL05) BETA3 (Basic helix-loop-helix domain
containing, class B5), partial (5%)
Length = 856
Score = 32.0 bits (71), Expect = 0.18
Identities = 17/54 (31%), Positives = 28/54 (51%)
Frame = +3
Query: 13 QHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIFPPPRWEFGGTSLDRSKGNKRNW 66
+ T R + + P + T++ W+ W +P + P W+F GTS SKG+ W
Sbjct: 384 ESTTRGVSAKPAALTTAFWIP---W--IP*LLPFSDWDFSGTSAAESKGSIPCW 530
>TC233256 weakly similar to UP|HUL5_YEAST (P53119) Probable
ubiquitin--protein ligase HUL5 , partial (3%)
Length = 732
Score = 31.2 bits (69), Expect = 0.30
Identities = 17/47 (36%), Positives = 22/47 (46%), Gaps = 1/47 (2%)
Frame = +2
Query: 120 GHKWVYSDTQNGC-EANYVGPWNASIDHQPRAWEFQLNSGIQTIAVI 165
G W+ T NGC + VG I H PRAW F ++T +I
Sbjct: 575 GRYWLLPATNNGCGQHXXVGASVIPIFHYPRAWRFCCMFILETAVII 715
>BE806467
Length = 355
Score = 30.4 bits (67), Expect = 0.51
Identities = 13/31 (41%), Positives = 18/31 (57%)
Frame = +1
Query: 12 LQHTLRNLCSFPTSSTSSKWVYAVFWRVVPR 42
L TLR +C +S+W Y+VFW + PR
Sbjct: 1 LHETLRRVC------LNSEWTYSVFWTIRPR 75
>AW472011 similar to SP|P29530|OLE1 P24 oleosin isoform A (P89). [Soybean]
{Glycine max}, partial (50%)
Length = 418
Score = 30.4 bits (67), Expect = 0.51
Identities = 23/71 (32%), Positives = 26/71 (36%), Gaps = 1/71 (1%)
Frame = +3
Query: 6 PLLHSLLQHTLRNLCSFPTSSTSSKWVYAVFWRVVPRIFPPPRWEFGGTSLDRSKGNK-R 64
P HS H L S+ S S Y WR P P P W DRS G+ R
Sbjct: 126 PFTHSRKVHLLST--SWADSPASQSAAYCSSWRD*PWQEPSPDWRLP----DRSVGSSAR 287
Query: 65 NWIIVWEDGFC 75
W +W C
Sbjct: 288 CWFQLWSPSAC 320
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.138 0.457
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,681,331
Number of Sequences: 63676
Number of extensions: 203100
Number of successful extensions: 1021
Number of sequences better than 10.0: 60
Number of HSP's better than 10.0 without gapping: 1009
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1012
length of query: 189
length of database: 12,639,632
effective HSP length: 92
effective length of query: 97
effective length of database: 6,781,440
effective search space: 657799680
effective search space used: 657799680
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 56 (26.2 bits)
Medicago: description of AC148098.5


