

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147960.5 - phase: 0
(617 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
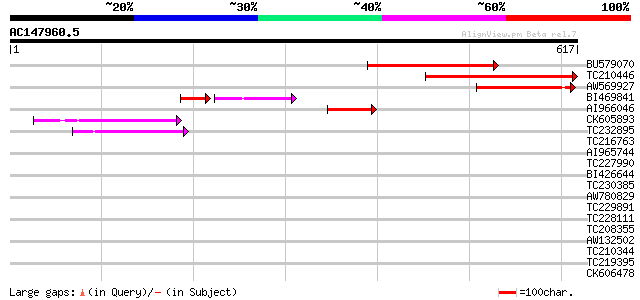
Score E
Sequences producing significant alignments: (bits) Value
BU579070 similar to GP|11994399|db contains similarity to recept... 186 2e-47
TC210446 similar to UP|Q7EZR1 (Q7EZR1) Receptor protein kinase P... 151 7e-37
AW569927 weakly similar to GP|11994399|dbj contains similarity t... 88 1e-17
BI469841 similar to GP|11994399|dbj contains similarity to recep... 56 2e-10
AI966046 63 3e-10
CK605893 59 8e-09
TC232895 weakly similar to PRF|1604369A.0|226743|1604369A sulfat... 46 4e-05
TC216763 UP|Q9LKY3 (Q9LKY3) Pti1 kinase-like protein, complete 40 0.003
AI965744 similar to GP|11994399|dbj contains similarity to recep... 35 0.010
TC227990 weakly similar to UP|Q84XG7 (Q84XG7) Erwinia induced pr... 38 0.015
BI426644 37 0.020
TC230385 weakly similar to UP|Q9M019 (Q9M019) Receptor like prot... 37 0.020
AW780829 37 0.026
TC229891 similar to UP|Q40096 (Q40096) Receptor protein kinase, ... 36 0.058
TC228111 similar to UP|Q9XED4 (Q9XED4) Receptor-like protein kin... 36 0.058
TC208355 UP|Q9LKY4 (Q9LKY4) Pti1 kinase-like protein, complete 36 0.058
AW132502 similar to PIR|D86244|D86 protein Ser/Thr protein kinas... 35 0.13
TC210344 similar to UP|Q9AVE0 (Q9AVE0) SRKb, partial (21%) 34 0.17
TC219395 UP|Q9LKZ4 (Q9LKZ4) Receptor-like protein kinase 3, part... 34 0.22
CK606478 33 0.37
>BU579070 similar to GP|11994399|db contains similarity to receptor protein
kinase~gene_id:MIL23.20 {Arabidopsis thaliana}, partial
(19%)
Length = 437
Score = 186 bits (473), Expect = 2e-47
Identities = 94/143 (65%), Positives = 102/143 (70%)
Frame = +2
Query: 390 H**WKLRPTSTQFRWRTFVMVYQG*NCSGFSQRS*IHSRTYRPYIYSSRHKVRKYSIRQK 449
H* W+LR TST R+++F MVY NCSGFSQRS IHS Y IY S HKV K+ RQK
Sbjct: 2 H*EWQLRTTST*IRFQSFAMVYPSSNCSGFSQRSSIHS*AYGTCIYPS*HKVGKHFNRQK 181
Query: 450 LLCKGCRLRINQAD*CWNFISSHC*YGWHIWLHATRICIW*CFFKNRCICFWSSSL*TNF 509
L CKGC L INQ D*CW FI+SHC*Y HIWLHATRICIW CF +NRC+CFWS SL*TNF
Sbjct: 182 LRCKGCILWINQVD*CWKFITSHC*YEGHIWLHATRICIWQCFSQNRCLCFWSCSL*TNF 361
Query: 510 C*SSCNNG*RFWC*FKGPCSFV* 532
* S WC* GPC FV*
Sbjct: 362 W*RSIEQRWCLWC*TXGPCIFV* 430
>TC210446 similar to UP|Q7EZR1 (Q7EZR1) Receptor protein kinase PERK1-like
protein, partial (29%)
Length = 741
Score = 151 bits (382), Expect = 7e-37
Identities = 88/166 (53%), Positives = 104/166 (62%), Gaps = 1/166 (0%)
Frame = +2
Query: 453 KGCRLRINQAD*CWNFISSHC*YGWHIWLHATRIC-IW*CFFKNRCICFWSSSL*TNFC* 511
+GC +Q D W I+SH W+IW+HAT +C IW CF ++R +CFWS SL*T F *
Sbjct: 17 QGC*FWFDQTDRSWKLIASHWSSCWNIWIHATGVCSIWRCFSQSRRVCFWSCSL*TYFG* 196
Query: 512 SSCNNG*RFWC*FKGPCSFV*RSF*SATSYRRS*KTGGS*AWR*LSN*SCIQDGTTCESM 571
S RF C KGP SFV* S *SA SY R+ +T S AW *L N* QDGTT +SM
Sbjct: 197 GSNCQDERFCCRLKGPRSFV*WSS*SARSYGRALQTS*SKAWG*LPN*FSSQDGTTSQSM 376
Query: 572 YK**SAATSKYEFCCSCSYHTNFNY*GLGYYFYIQKSKSCKSNVWK 617
Y S SKYE C CSY T FNY LG +F ++KSKSC+S V K
Sbjct: 377 YTRQSPTPSKYEIYCGCSYDTFFNYRRLGCWFLLRKSKSCESYVRK 514
>AW569927 weakly similar to GP|11994399|dbj contains similarity to receptor
protein kinase~gene_id:MIL23.20 {Arabidopsis thaliana},
partial (8%)
Length = 380
Score = 88.2 bits (217), Expect = 1e-17
Identities = 55/107 (51%), Positives = 67/107 (62%)
Frame = +1
Query: 509 FC*SSCNNG*RFWC*FKGPCSFV*RSF*SATSYRRS*KTGGS*AWR*LSN*SCIQDGTTC 568
FC C+ C* KGPCSFV*RS *S S+R+ +TGGS*AWR LSN* QD +T
Sbjct: 1 FCKECCSKDR*ICC*IKGPCSFV*RST*SE*SFRKYSQTGGS*AWRKLSN*FSSQDCSTW 180
Query: 569 ESMYK**SAATSKYEFCCSCSYHTNFNY*GLGYYFYIQKSKSCKSNV 615
ES+YK * T +YE CS+ T Y*GL + +++KS S KS V
Sbjct: 181 ESLYKR*PTTTPEYEVYSCCSHDTFITY*GLRH--FLRKSDSHKSTV 315
>BI469841 similar to GP|11994399|dbj contains similarity to receptor protein
kinase~gene_id:MIL23.20 {Arabidopsis thaliana}, partial
(6%)
Length = 396
Score = 56.2 bits (134), Expect(2) = 2e-10
Identities = 23/33 (69%), Positives = 30/33 (90%)
Frame = +1
Query: 186 DAELLQKYNPGVNFSQGSGLVYIPGKDQNRNYV 218
D+ LLQ+YNP VNF+QG+GLVY+PGKDQN ++V
Sbjct: 1 DSALLQRYNPDVNFNQGTGLVYVPGKDQNGSFV 99
Score = 27.7 bits (60), Expect(2) = 2e-10
Identities = 30/90 (33%), Positives = 43/90 (47%)
Frame = +3
Query: 223 RSIRWSYNWNICWSSSRTNSIIILHLCYVLPKEEDKEARISFGRIQRDLRSS*E**SLR* 282
RS SY N C + ++ I L++C++ P ED E IS R L S E *+ *
Sbjct: 120 RSYW*SYCRNSCRNCGSSSVIGSLYICWIFP-*EDTEG*ISSTRFHCTLCSRWEG*NFS* 296
Query: 283 CYIWNFRFCESC*YDWN*GGKIRRVFI*RT 312
NFR +C + + +I +FI* T
Sbjct: 297 *CK*NFRTWWTCHHHRHNSEQISGIFI*GT 386
>AI966046
Length = 160
Score = 63.2 bits (152), Expect = 3e-10
Identities = 32/53 (60%), Positives = 39/53 (73%)
Frame = +2
Query: 347 KDGHESNKRISC*VEGVDACSSREPGTLDRILC*GIPFPCL*IH**WKLRPTS 399
+DG+ S KRISC*+E +D CS G +DRI *G+ FPCL*IH* WKL+ TS
Sbjct: 2 EDGYASIKRISC*IERLDTCSPS*SGAVDRI*Y*GLSFPCL*IH*EWKLKSTS 160
>CK605893
Length = 780
Score = 58.5 bits (140), Expect = 8e-09
Identities = 42/162 (25%), Positives = 73/162 (44%), Gaps = 1/162 (0%)
Frame = +3
Query: 27 SKTCNIALASYYLQDDTNLTYVSNIMQSNLVTKPEDIVSYNTDTITNKDFVQSFTRVNVP 86
+KTC L ++T +SN+ SN PE++ N T+ + V VP
Sbjct: 192 NKTCMSFLIFKSKPPFNSITTISNLTSSN----PEELARINDVTVLK--VFPTGKEVIVP 353
Query: 87 FPCDCIHDEFLGHIFQYQVATKDTYLSVASNNYSNLTTSEWLQNFNSYPSNDIPDTGTLN 146
C C+ E+ +Y + TY +VA++ + LTT + L NSY D+ L+
Sbjct: 354 LNCSCLTREYYQAETKYVLGQSPTYFTVANDTFEGLTTCDTLMRANSYGELDLLPGMELH 533
Query: 147 VTVNCSCGN-SDVSKDYGLFITYPLRPEDSLELISNKTEIDA 187
V + C+C ++ +TY + DS++ I+ + + A
Sbjct: 534 VPLRCACPTWHQITNGTKYLLTYSVNWGDSIKNIAARFNVAA 659
>TC232895 weakly similar to PRF|1604369A.0|226743|1604369A sulfated surface
glycoprotein SSG185. {Volvox carteri;} , partial (14%)
Length = 907
Score = 46.2 bits (108), Expect = 4e-05
Identities = 32/128 (25%), Positives = 59/128 (46%), Gaps = 2/128 (1%)
Frame = +2
Query: 69 DTITNKDFVQSFTRVNVPFPCDCIHDEFLGHIFQYQVATK-DTYLSVASNNYSNLTTSEW 127
+ IT+ + + T V VP C C + H Y + + +TY S+A+N Y LTT +
Sbjct: 338 NNITDVQTLPADTLVTVPVNCSC-SGPYYQHNASYTIKVQGETYFSIANNTYQALTTCQA 514
Query: 128 LQNFNSYPSNDIPDTGTLNVTVNCSCGNSDVSK-DYGLFITYPLRPEDSLELISNKTEID 186
L+ N+ D+ L+V + C+C + + +TY + +S+ I + +D
Sbjct: 515 LELQNTVGMRDLLKGQNLHVPLRCACPTQKQREAGFKYLLTYLVSQGESVSAIGDIFGVD 694
Query: 187 AELLQKYN 194
+ + N
Sbjct: 695 EQSILDAN 718
>TC216763 UP|Q9LKY3 (Q9LKY3) Pti1 kinase-like protein, complete
Length = 1634
Score = 40.0 bits (92), Expect = 0.003
Identities = 34/101 (33%), Positives = 47/101 (45%), Gaps = 2/101 (1%)
Frame = +3
Query: 404 WRTFVMVYQG*NCSGFSQRS*IHSRTYRPYIYSSRHKVRKYSIRQKLLCKGCRL-RINQA 462
W + +M + *NC SQ +*I S R + S H V ++ CKGC + +
Sbjct: 681 WSSSLMGSES*NCCWSSQGT*ISS*KGRDSYHPSLH*V**HTSF***CCKGC*F*FVKSS 860
Query: 463 D*CWNFISSHC*YGWHIWLHATRIC-IW*CFFKNRCICFWS 502
*C + SS W +WL +RIC W FK C+ WS
Sbjct: 861 P*C-SSTSSFYSCSWDLWLSCSRICNDWTTHFKK*CL*LWS 980
>AI965744 similar to GP|11994399|dbj contains similarity to receptor protein
kinase~gene_id:MIL23.20 {Arabidopsis thaliana}, partial
(15%)
Length = 425
Score = 35.4 bits (80), Expect(2) = 0.010
Identities = 19/52 (36%), Positives = 28/52 (53%)
Frame = +2
Query: 415 NCSGFSQRS*IHSRTYRPYIYSSRHKVRKYSIRQKLLCKGCRLRINQAD*CW 466
NC S+R IH + ++ R+ + K+ RQKL CKGCR ++ D W
Sbjct: 269 NCP*CSKRIGIHP*AHCSCLHP*RY*ISKHFDRQKLSCKGCRFWSHKIDRIW 424
Score = 21.9 bits (45), Expect(2) = 0.010
Identities = 11/20 (55%), Positives = 15/20 (75%)
Frame = +3
Query: 372 GTLDRILC*GIPFPCL*IH* 391
G ++RILC*G+ L*+H*
Sbjct: 141 GEVNRILC*GLLIFGL*VH* 200
>TC227990 weakly similar to UP|Q84XG7 (Q84XG7) Erwinia induced protein 1,
partial (35%)
Length = 1386
Score = 37.7 bits (86), Expect = 0.015
Identities = 43/170 (25%), Positives = 72/170 (42%), Gaps = 4/170 (2%)
Frame = +3
Query: 29 TCNIALASYYLQDDTNLTYVSNIMQSNLVTKPEDIVSYNT--DTITNKDFVQSFTRVNVP 86
TC AL SY N T + +I + V DIV N T V V VP
Sbjct: 171 TCR-ALISY---SHPNTTTLGDIQKLFNVKHILDIVGANNLPSNATKTYAVGPNEVVKVP 338
Query: 87 FPCDCIHDEFLG-HIFQYQVATKDTYLSVASNNYSNLTTSEWLQNFNSYP-SNDIPDTGT 144
FPC C ++ L + Y++ DT +A+ ++ L +Q N+ +N+I
Sbjct: 339 FPCRCSNNTGLSDRVPLYRIKKGDTLYYIATTTFAGLMKWPQIQVANNIANANNITTGDM 518
Query: 145 LNVTVNCSCGNSDVSKDYGLFITYPLRPEDSLELISNKTEIDAELLQKYN 194
L + + CSC +V + + + P+ ++E I+ + ++L N
Sbjct: 519 LYIPLPCSC--DEVGGKSVVHYAHLVAPQSTVEGIAEEFGTTQQILLNLN 662
>BI426644
Length = 421
Score = 37.4 bits (85), Expect = 0.020
Identities = 30/86 (34%), Positives = 41/86 (46%), Gaps = 5/86 (5%)
Frame = +1
Query: 386 CL*IH**WKLRPTSTQFRWRT-----FVMVYQG*NCSGFSQRS*IHSRTYRPYIYSSRHK 440
CL*IH WK R S++ +W+T F + Y *N +* +R Y P SR+K
Sbjct: 115 CL*IHGKWKSRRLSSKKKWKTTTSLVFSISYSF*NGLWALILA*FKARAYCP----SRYK 282
Query: 441 VRKYSIRQKLLCKGCRLRINQAD*CW 466
+ +RQKL + CR CW
Sbjct: 283 TWQCLVRQKLCEQNCR--------CW 336
>TC230385 weakly similar to UP|Q9M019 (Q9M019) Receptor like protein kinase,
partial (23%)
Length = 699
Score = 37.4 bits (85), Expect = 0.020
Identities = 16/37 (43%), Positives = 22/37 (59%), Gaps = 1/37 (2%)
Frame = +3
Query: 466 WNFISSHC*YGWHIWLHATRI-CIW*CFFKNRCICFW 501
W+ ++ H GWH WLH TR+ F K+RC+C W
Sbjct: 99 WSIVT-HNKCGWHYWLHCTRVNSHRKGFHKHRCVCIW 206
>AW780829
Length = 424
Score = 37.0 bits (84), Expect = 0.026
Identities = 24/98 (24%), Positives = 47/98 (47%), Gaps = 1/98 (1%)
Frame = +1
Query: 86 PFPCDCIHDEFLGHIFQYQVATKDTYLSVASNNYSNLTTSEWLQNFNSY-PSNDIPDTGT 144
P C C+ Y V DT S+ S Y L ++E ++ N+ +N + GT
Sbjct: 10 PISCSCVDGIRRSMSTIYTVHAADTLASI-SEGYGGLVSAEQIKIVNAINATNPLTYRGT 186
Query: 145 LNVTVNCSCGNSDVSKDYGLFITYPLRPEDSLELISNK 182
L + + C+C ++ + ++++Y ++ +SL I+ K
Sbjct: 187 LVIPLPCTCFDNVNNGGNAIYMSYVVQRRESLXSIATK 300
>TC229891 similar to UP|Q40096 (Q40096) Receptor protein kinase, partial
(15%)
Length = 731
Score = 35.8 bits (81), Expect = 0.058
Identities = 27/83 (32%), Positives = 35/83 (41%), Gaps = 12/83 (14%)
Frame = +3
Query: 431 RPYIYSSRHKVRKYSIRQKLLCKGCRLRINQAD*CWNFISS-----------HC*YGWHI 479
R Y+ SSR + Y R K K C R + + F S H W+I
Sbjct: 30 RAYVPSSRFSIEDYP*RSK--SK*CFTRRHFESQNFRFWSG*NFWRREY*RKHNQNCWNI 203
Query: 480 WLHATRICI-W*CFFKNRCICFW 501
W+H TRIC W F+K C+ W
Sbjct: 204 WVHGTRICY*WTIFYKI*CLQLW 272
>TC228111 similar to UP|Q9XED4 (Q9XED4) Receptor-like protein kinase homolog
RK20-1, partial (28%)
Length = 738
Score = 35.8 bits (81), Expect = 0.058
Identities = 13/29 (44%), Positives = 20/29 (68%), Gaps = 1/29 (3%)
Frame = +2
Query: 477 WHIWLHATRIC-IW*CFFKNRCICFWSSS 504
W++W++ TRIC W F + RC+ FW +S
Sbjct: 92 WNLWIYGTRICNAWTIFSEIRCLQFWCTS 178
>TC208355 UP|Q9LKY4 (Q9LKY4) Pti1 kinase-like protein, complete
Length = 1536
Score = 35.8 bits (81), Expect = 0.058
Identities = 43/137 (31%), Positives = 59/137 (42%), Gaps = 11/137 (8%)
Frame = +1
Query: 377 ILC*-GIPFPCL*IH**--------WKLRPTSTQFRWRTFVMVYQG*NCSGFSQRS*IHS 427
+LC* P PCL*+ * W R W + +M + *NC SQ +*I S
Sbjct: 586 LLC*WSFPCPCL*VCS*RIPS*YSTWTQRCQGCT-TWSSSLMGSES*NCCWSSQGT*ISS 762
Query: 428 RTYRPYIYSSRHKVRKYSIRQKLLCKGCRL-RINQAD*CWNFISSHC*YGWHIWLHATRI 486
R + S H V ++ C+ C + + *C + SS W +WL +RI
Sbjct: 763 *KGRDSYHPSLH*V**HTSF***RCEDC*F*FVKSSP*C-SSTSSFYPCSWDLWLPCSRI 939
Query: 487 C-IW*CFFKNRCICFWS 502
C W FK C+ WS
Sbjct: 940 CNDWTTHFKK*CL*LWS 990
>AW132502 similar to PIR|D86244|D86 protein Ser/Thr protein kinase homolog
[imported] - Arabidopsis thaliana, partial (17%)
Length = 407
Score = 34.7 bits (78), Expect = 0.13
Identities = 17/34 (50%), Positives = 20/34 (58%), Gaps = 1/34 (2%)
Frame = +2
Query: 469 ISSHC*YGWHIWLHATRIC-IW*CFFKNRCICFW 501
ISSH WHIWL TR+C IW + C+ FW
Sbjct: 236 ISSHYKGCWHIWLCGTRVCSIWETN*EK*CV*FW 337
>TC210344 similar to UP|Q9AVE0 (Q9AVE0) SRKb, partial (21%)
Length = 870
Score = 34.3 bits (77), Expect = 0.17
Identities = 13/30 (43%), Positives = 18/30 (59%)
Frame = +1
Query: 472 HC*YGWHIWLHATRICIW*CFFKNRCICFW 501
HC WHIWL+ +R+C F ++ CFW
Sbjct: 187 HCYNSWHIWLYGSRVCNGGIIF-HKV*CFW 273
>TC219395 UP|Q9LKZ4 (Q9LKZ4) Receptor-like protein kinase 3, partial (92%)
Length = 3037
Score = 33.9 bits (76), Expect = 0.22
Identities = 15/37 (40%), Positives = 22/37 (58%), Gaps = 1/37 (2%)
Frame = +2
Query: 472 HC*YGWHIWLHATRICIW-*CFFKNRCICFWSSSL*T 507
H Y W IW+H++R+C+ + RC+ FW SL T
Sbjct: 2315 HVCYCWFIWIHSSRVCLHIES**EKRCVQFWCGSLRT 2425
>CK606478
Length = 815
Score = 33.1 bits (74), Expect = 0.37
Identities = 31/126 (24%), Positives = 51/126 (39%), Gaps = 7/126 (5%)
Frame = +2
Query: 385 PCL*IH**WKLR--PTSTQFRWRTFVMVYQG*NCSGFSQRS*IHSRTYRPYIYSSRHKVR 442
PCL*+ *W L Q W + + +C G R+*+ S+ + Y +K +
Sbjct: 17 PCL*VCR*WGLE*MDLLPQREWEISELDAENADCIGCGHRT*LSSQFHFSSSYPQGYKQQ 196
Query: 443 KYSIRQKLLCKGCRLR----INQAD*CWNFISSHC*YGWHIWLHATRIC-IW*CFFKNRC 497
+S KG + + + +HC W LH +R+ W C +K C
Sbjct: 197 *HSSGW*FQGKGHEFKPC*VFGRRGRSTSRDEAHC---WDKRLHGSRVFGKWSCVYKA*C 367
Query: 498 ICFWSS 503
+C W +
Sbjct: 368 VCIWGT 385
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.351 0.153 0.577
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 35,202,078
Number of Sequences: 63676
Number of extensions: 625291
Number of successful extensions: 6192
Number of sequences better than 10.0: 66
Number of HSP's better than 10.0 without gapping: 4505
Number of HSP's successfully gapped in prelim test: 178
Number of HSP's that attempted gapping in prelim test: 1549
Number of HSP's gapped (non-prelim): 4856
length of query: 617
length of database: 12,639,632
effective HSP length: 103
effective length of query: 514
effective length of database: 6,081,004
effective search space: 3125636056
effective search space used: 3125636056
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.9 bits)
S2: 62 (28.5 bits)
Medicago: description of AC147960.5


