

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
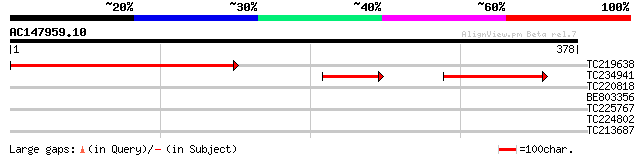
Query= AC147959.10 + phase: 1 /pseudo
(378 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC219638 195 2e-50
TC234941 97 6e-30
TC220818 37 0.015
BE803356 31 1.0
TC225767 similar to UP|Q8W590 (Q8W590) At2g32080/F22D22.17, part... 28 5.2
TC224802 similar to UP|O24320 (O24320) Lipoxygenase , partial (... 28 5.2
TC213687 28 8.9
>TC219638
Length = 968
Score = 195 bits (496), Expect = 2e-50
Identities = 94/153 (61%), Positives = 121/153 (78%), Gaps = 1/153 (0%)
Frame = +2
Query: 1 DGRRVQDFFLSMEVAIRNHPLWATATEEDIDCAMEGLEKYIMTKLFSRTFAASPEDAKID 60
D VQ F ME R HPLWA +EE+++ A EGLEKY+MTKLF+R FA+ P+D K D
Sbjct: 182 DSAAVQAFLAKMEADFRAHPLWAGCSEEELESAGEGLEKYVMTKLFARVFASLPDDVKFD 361
Query: 61 HEISEKISLLQTFLKPEHLDIPPVLHNEASWLLAEKELQKINAFKAPQEKLSTIMNCCRV 120
++SEK++L+Q F++PE+LDI PV NE+SWLLA+KELQKIN +KAP++KL I+NCCRV
Sbjct: 362 DQLSEKMALIQQFIRPENLDIKPVFQNESSWLLAQKELQKINMYKAPRDKLVCILNCCRV 541
Query: 121 INNLLLNAAM-SEYVPAGADDFIPVLIYVTIKA 152
I+NLLLNA++ S P GAD+F+PVLIYVTIKA
Sbjct: 542 ISNLLLNASVASRENPPGADEFLPVLIYVTIKA 640
>TC234941
Length = 811
Score = 97.1 bits (240), Expect(2) = 6e-30
Identities = 46/69 (66%), Positives = 58/69 (83%)
Frame = +2
Query: 290 VLQHGTNYPYMEAESKDLAMEDVDILLNHYKDLVAKYTIICKAINYLSMSEKEPLLHQLE 349
VL HG+NYPYMEA+SK+L + DVD+LL+ YKDLVAKYTI+CKAI LS +E+EPLL LE
Sbjct: 131 VLLHGSNYPYMEAKSKELTVGDVDMLLSDYKDLVAKYTILCKAIGCLSTAEREPLLRHLE 310
Query: 350 MQGEGSMLS 358
MQG ++L+
Sbjct: 311 MQGPETLLN 337
Score = 52.0 bits (123), Expect(2) = 6e-30
Identities = 26/41 (63%), Positives = 30/41 (72%)
Frame = +3
Query: 209 ACKQPNWPRKSQVNYTLHVKLNKK*RTRAMFQTKCITNLIL 249
ACKQP+ P KSQVNY LHVK K RT A+ Q KC TNL++
Sbjct: 3 ACKQPS*PIKSQVNYPLHVK*ASKRRTIAVVQRKCTTNLMI 125
>TC220818
Length = 816
Score = 37.0 bits (84), Expect = 0.015
Identities = 17/37 (45%), Positives = 24/37 (63%)
Frame = +2
Query: 297 YPYMEAESKDLAMEDVDILLNHYKDLVAKYTIICKAI 333
+PY+ A DL + DV+ LLN+YK LV KY + K +
Sbjct: 176 FPYLFASVGDLMVGDVEDLLNNYKQLVFKYVSLSKGL 286
>BE803356
Length = 559
Score = 30.8 bits (68), Expect = 1.0
Identities = 22/74 (29%), Positives = 31/74 (41%), Gaps = 5/74 (6%)
Frame = -2
Query: 22 WATATEEDIDC-----AMEGLEKYIMTKLFSRTFAASPEDAKIDHEISEKISLLQTFLKP 76
W T E DIDC + G L +R AS K++H SE+ SL F++
Sbjct: 261 WLTVLESDIDCSPLM*SFVGSRHSSGMVLKARMLCASTASLKLEHFTSEERSL*PVFIRR 82
Query: 77 EHLDIPPVLHNEAS 90
+ P H A+
Sbjct: 81 GNFW*PAPWHGHAT 40
>TC225767 similar to UP|Q8W590 (Q8W590) At2g32080/F22D22.17, partial (93%)
Length = 1586
Score = 28.5 bits (62), Expect = 5.2
Identities = 13/45 (28%), Positives = 26/45 (56%)
Frame = -2
Query: 213 PNWPRKSQVNYTLHVKLNKK*RTRAMFQTKCITNLILETK*TILG 257
PN+ + Q+N +H++L + R M + + TN+I ++ T+ G
Sbjct: 946 PNFILRRQINRPVHIQLRRSGRLSTMTRYEASTNIIRKSNKTL*G 812
>TC224802 similar to UP|O24320 (O24320) Lipoxygenase , partial (73%)
Length = 2213
Score = 28.5 bits (62), Expect = 5.2
Identities = 18/65 (27%), Positives = 35/65 (53%), Gaps = 3/65 (4%)
Frame = -2
Query: 91 WLLAEKELQKINAFKAPQEKLSTIMNC--CRVI-NNLLLNAAMSEYVPAGADDFIPVLIY 147
WL+ + + ++N+F P +L +NC C VI +++LN ++ + + I V++Y
Sbjct: 1396 WLVFQVTVTRLNSFLPPSLELRVFLNCIICWVIQGDIVLNPCLNSIPYF*SINSIRVVLY 1217
Query: 148 VTIKA 152
KA
Sbjct: 1216 HKAKA 1202
>TC213687
Length = 433
Score = 27.7 bits (60), Expect = 8.9
Identities = 14/19 (73%), Positives = 16/19 (83%)
Frame = +2
Query: 58 KIDHEISEKISLLQTFLKP 76
+I +E SEKISLL TFLKP
Sbjct: 197 QIVNETSEKISLL*TFLKP 253
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.328 0.139 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,292,107
Number of Sequences: 63676
Number of extensions: 209903
Number of successful extensions: 1259
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 1253
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1258
length of query: 378
length of database: 12,639,632
effective HSP length: 99
effective length of query: 279
effective length of database: 6,335,708
effective search space: 1767662532
effective search space used: 1767662532
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 60 (27.7 bits)
Medicago: description of AC147959.10


