

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147741.8 - phase: 0
(402 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
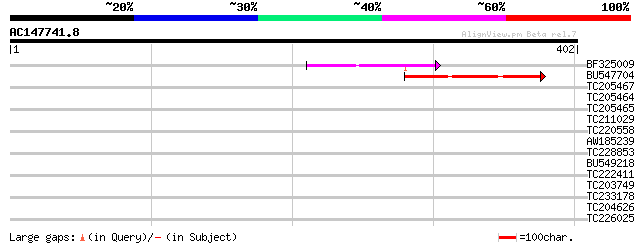
Sequences producing significant alignments: (bits) Value
BF325009 82 4e-16
BU547704 76 3e-14
TC205467 40 0.002
TC205464 weakly similar to GB|AAP42752.1|30984578|BT008739 At1g5... 34 0.13
TC205465 weakly similar to GB|AAP42752.1|30984578|BT008739 At1g5... 33 0.23
TC211029 similar to UP|Q96VI4 (Q96VI4) Protease 1, partial (3%) 33 0.30
TC220558 similar to UP|Q9M3H9 (Q9M3H9) Tubby-like protein, parti... 31 1.1
AW185239 weakly similar to GP|15010754|gb| AT3g08600/F17O14_7 {A... 30 2.5
TC228853 similar to UP|O04082 (O04082) Transcription factor RUSH... 29 3.3
BU549218 weakly similar to GP|19911193|db glucosyltransferase-5 ... 29 4.3
TC222411 28 5.7
TC203749 similar to UP|Q945N6 (Q945N6) AT5g10490/F12B17_160, par... 28 5.7
TC233178 similar to GB|AAO63380.1|28950913|BT005316 At5g23170 {A... 28 7.4
TC204626 UP|GLCB_SOYBN (P25974) Beta-conglycinin, beta chain pre... 28 7.4
TC226025 similar to UP|RK25_PEA (P11892) 50S ribosomal protein C... 28 9.6
>BF325009
Length = 382
Score = 82.0 bits (201), Expect = 4e-16
Identities = 49/109 (44%), Positives = 63/109 (56%), Gaps = 14/109 (12%)
Frame = +2
Query: 211 ICLFKGQSYAVDKIGKTVVVGPDSS-VQLVAEPLIGGGDMKFLVESEGDLLLADVYNCLF 269
+C+FKG Y D G+TV V PD + L AEP+ GG D KFLVESEG LLL D+Y +
Sbjct: 2 VCVFKGNLYGADSNGRTVRVRPDDGGLTLAAEPVFGG-DKKFLVESEGALLLVDMYLSYY 178
Query: 270 TDLCNPDHND-------------CVRIDLFKLNEKEKKWVKLTSLGDRV 305
+ H D V+ D+F+L+E+ KKWV+LT L DRV
Sbjct: 179 SCTQGLFHEDFDEEDVAGMGWERTVKFDVFRLDEEGKKWVELTDLRDRV 325
>BU547704
Length = 600
Score = 75.9 bits (185), Expect = 3e-14
Identities = 45/100 (45%), Positives = 61/100 (61%)
Frame = -2
Query: 281 VRIDLFKLNEKEKKWVKLTSLGDRVLFLGLGLVCSFSACASDLCVAKGNSVIFNNYIFES 340
V+ D+F L+E+ KKWV+LT LG+RVLFLG C+FSA A DL + +GN V +
Sbjct: 560 VKFDVFXLDEEGKKWVELTDLGERVLFLGDD--CAFSASAKDLNLGRGNCVAXRDDGLGF 387
Query: 341 HRPLESECKEYVLDLDLGRHSLLSDYPEYSNLFWPPPEWI 380
+R L V LD G+ S LS+ +S LF PPP+W+
Sbjct: 386 NRVLNG---MGVFRLDDGKISPLSECAGFSELFCPPPDWV 276
>TC205467
Length = 1504
Score = 40.0 bits (92), Expect = 0.002
Identities = 89/368 (24%), Positives = 131/368 (35%), Gaps = 22/368 (5%)
Frame = +1
Query: 6 WSKLPTELLNLISQRIYEEVDLIHFRSVCSTWRSSSVPKHHHKFRFQFPLLKFPFNADFI 65
W+ LP ELL I R+ D I +VC W S + + +
Sbjct: 37 WADLPAELLESILSRLIL-ADNIRASAVCRRWHSVAS------------------DVRVV 159
Query: 66 NKNRSFCYITK-QNIFLIKPPQQEQTLTLR-PLL--IRICQNARGKSKLYHPLLQNGYKF 121
N++ Y K + + P Q +T T P L R+C G LY P + F
Sbjct: 160 NQSPWLMYFPKFGDCYEFYDPVQRKTHTFELPELNGSRVCYTKDGWLLLYRPRTHRVFFF 339
Query: 122 PHTFMLDFNKLSILHLGSNFFMIDFDFTFNDKLYNPDDYMYPKKVVVVTCHGKKPLIVGT 181
+ F + KL F M F+ +P ++ K V P +V
Sbjct: 340 -NPFTRELIKLP------RFEMTYQIVAFSCAPTSPGCVLFTVKHV-------SPTVVAI 477
Query: 182 LTSPPQPLLMKCGDEKW------NVIPDMSTKFGDICLFKGQSYAVDKIGKTVVVGPDS- 234
T P G +W N +P +S+ + + G Y + G V P
Sbjct: 478 STCYP-------GATEWTTVNYQNRLPFVSSIWNKLVFCNGLFYCLSLTGWLGVFDPVEC 636
Query: 235 --SVQLVAEPLIGGGDM-------KFLVESEGDLLLADVYNCLFTDLCNPDHNDCVRIDL 285
SV V P KF+ E EGD+L+ +Y C NP +
Sbjct: 637 TWSVLAVPPPKCPENFFAKNWWKGKFMAEHEGDILV--IYTCCSE---NPI--------I 777
Query: 286 FKLNEKEKKWVKLTSLGDRVLFLGLGLVCSF--SACASDLCVAKGNSVIFNNYIFESHRP 343
FKL++ KW ++T+L G+ L SF S +DL NSV F+ F R
Sbjct: 778 FKLDQTLMKWEEMTTLD------GVTLFASFLSSHSRTDLIGIMRNSVYFSKVRFYGKRC 939
Query: 344 LESECKEY 351
+ +Y
Sbjct: 940 ISFSLGDY 963
>TC205464 weakly similar to GB|AAP42752.1|30984578|BT008739 At1g57790
{Arabidopsis thaliana;} , partial (17%)
Length = 756
Score = 33.9 bits (76), Expect = 0.13
Identities = 28/114 (24%), Positives = 42/114 (36%), Gaps = 14/114 (12%)
Frame = +1
Query: 6 WSKLPTELLNLISQRIYEEVDLIHFRSVCSTWRSSSV--------------PKHHHKFRF 51
WS LPTELL LI R+ + D + VC W S + PK + F
Sbjct: 181 WSDLPTELLELILSRLSLD-DNVRASVVCKRWHSVATSVCVVNQSPWLMYFPKFGDWYEF 357
Query: 52 QFPLLKFPFNADFINKNRSFCYITKQNIFLIKPPQQEQTLTLRPLLIRICQNAR 105
P + ++ + S TK L+ P+ + P + I + R
Sbjct: 358 YDPAHRKTYSIELPELRGSRVCYTKDGWLLLYRPRTHRVFFFNPFTMEIIKLPR 519
>TC205465 weakly similar to GB|AAP42752.1|30984578|BT008739 At1g57790
{Arabidopsis thaliana;} , partial (15%)
Length = 490
Score = 33.1 bits (74), Expect = 0.23
Identities = 26/104 (25%), Positives = 40/104 (38%), Gaps = 14/104 (13%)
Frame = +3
Query: 6 WSKLPTELLNLISQRIYEEVDLIHFRSVCSTWRSSSV--------------PKHHHKFRF 51
WS LPTELL LI R+ + D + VC W S + PK + F
Sbjct: 177 WSDLPTELLELILSRLSLD-DNVRASVVCKRWHSVATSVCVVNQSPWLMYFPKFGDWYEF 353
Query: 52 QFPLLKFPFNADFINKNRSFCYITKQNIFLIKPPQQEQTLTLRP 95
P+ + ++ + + S TK L+ P+ + P
Sbjct: 354 YDPVHRKTYSIELPELSGSRVCYTKDGWLLLYRPRTHRVFFFNP 485
>TC211029 similar to UP|Q96VI4 (Q96VI4) Protease 1, partial (3%)
Length = 858
Score = 32.7 bits (73), Expect = 0.30
Identities = 29/101 (28%), Positives = 44/101 (42%), Gaps = 9/101 (8%)
Frame = +3
Query: 30 FRSVCSTWRSSSVPKHHHKFRFQFPLLKFPFNADFINKNRSFCYITKQNIFLIKPPQQEQ 89
F +TW P+HHH+ + Q + N FI N SF +IT N FL P
Sbjct: 357 FDPASNTWFLLPTPQHHHQHQHQ-----YQSNTSFIGTN-SFFFITAPN-FLYTPILHPS 515
Query: 90 TLTLRPL-------LIRICQNARGKSKLYHP--LLQNGYKF 121
PL L+ + +A+ ++ +HP ++ G KF
Sbjct: 516 WHPTPPLHFPRINPLLGVFHDAKDQNFGHHPKFIVVGGVKF 638
>TC220558 similar to UP|Q9M3H9 (Q9M3H9) Tubby-like protein, partial (42%)
Length = 629
Score = 30.8 bits (68), Expect = 1.1
Identities = 27/80 (33%), Positives = 37/80 (45%), Gaps = 16/80 (20%)
Frame = +2
Query: 6 WSKLPTELLNLISQRIYE-------EVDLIHFRSVCSTWRSSS---VPKHHHKFRFQFPL 55
W+ LP+ELL I QR+ E ++ SVC +WRS + V R FP+
Sbjct: 266 WANLPSELLLDIIQRVEESETSWPARAVVVFCASVCKSWRSITREIVKTPEQCGRITFPI 445
Query: 56 -LKFPFNAD-----FINKNR 69
LK P D FI +N+
Sbjct: 446 SLKQPGPRDSPIQCFIRRNK 505
>AW185239 weakly similar to GP|15010754|gb| AT3g08600/F17O14_7 {Arabidopsis
thaliana}, partial (26%)
Length = 431
Score = 29.6 bits (65), Expect = 2.5
Identities = 16/44 (36%), Positives = 24/44 (54%)
Frame = -3
Query: 68 NRSFCYITKQNIFLIKPPQQEQTLTLRPLLIRICQNARGKSKLY 111
NRS Y+T+Q I L++P +PL IR+ +N KL+
Sbjct: 390 NRSQIYVTRQRIKLLRPIA*VMVHKNQPLYIRLFKNPNWDLKLF 259
>TC228853 similar to UP|O04082 (O04082) Transcription factor RUSH-1alpha
isolog; 18684-24052, partial (24%)
Length = 1393
Score = 29.3 bits (64), Expect = 3.3
Identities = 19/59 (32%), Positives = 27/59 (45%), Gaps = 2/59 (3%)
Frame = -3
Query: 136 HLGSNFFMIDFDFTFNDKLYNPDDYMYPKKVVVVTCHGKKPLIVGTLTSPP--QPLLMK 192
H S F + F + Y Y +P+K C GK ++V T T+ P QPL +K
Sbjct: 671 HFDSREFHLRISFCQH*MSYQGHRY-FPQKCSAQNCFGKHSIVV*TTTATPQVQPLFLK 498
>BU549218 weakly similar to GP|19911193|db glucosyltransferase-5 {Vigna
angularis}, partial (22%)
Length = 632
Score = 28.9 bits (63), Expect = 4.3
Identities = 11/22 (50%), Positives = 13/22 (59%)
Frame = +1
Query: 172 HGKKPLIVGTLTSPPQPLLMKC 193
H PL+VG SPP P L+ C
Sbjct: 466 HDSTPLLVGPSFSPPTPFLLSC 531
>TC222411
Length = 968
Score = 28.5 bits (62), Expect = 5.7
Identities = 15/43 (34%), Positives = 24/43 (54%)
Frame = -3
Query: 167 VVVTCHGKKPLIVGTLTSPPQPLLMKCGDEKWNVIPDMSTKFG 209
+++T +GK PLI+GT T PL ++ W I +S +G
Sbjct: 531 LLLTSYGKHPLILGTRTFEQSPLALR----TWKRISCISLVYG 415
>TC203749 similar to UP|Q945N6 (Q945N6) AT5g10490/F12B17_160, partial (12%)
Length = 1130
Score = 28.5 bits (62), Expect = 5.7
Identities = 17/35 (48%), Positives = 20/35 (56%), Gaps = 1/35 (2%)
Frame = +1
Query: 85 PQQEQTLTLRPLLIRICQ-NARGKSKLYHPLLQNG 118
P Q LT LIRICQ N++GK L H L+ G
Sbjct: 586 PPQLTMLTKLVDLIRICQ*NSKGKRNLLHRLMHPG 690
>TC233178 similar to GB|AAO63380.1|28950913|BT005316 At5g23170 {Arabidopsis
thaliana;} , partial (34%)
Length = 1056
Score = 28.1 bits (61), Expect = 7.4
Identities = 16/41 (39%), Positives = 22/41 (53%)
Frame = +1
Query: 288 LNEKEKKWVKLTSLGDRVLFLGLGLVCSFSACASDLCVAKG 328
LNE+E+ W G+ V+ + + L C F A LCV KG
Sbjct: 820 LNEEERPWA-----GEVVMGMEMELNCFFRVPAWRLCVLKG 927
>TC204626 UP|GLCB_SOYBN (P25974) Beta-conglycinin, beta chain precursor,
complete
Length = 1653
Score = 28.1 bits (61), Expect = 7.4
Identities = 16/32 (50%), Positives = 18/32 (56%)
Frame = -3
Query: 40 SSVPKHHHKFRFQFPLLKFPFNADFINKNRSF 71
SSVP H +F F F LL FNA+ N SF
Sbjct: 1015 SSVPLHFQRFLFLFLLLLLFFNANKFNVCISF 920
>TC226025 similar to UP|RK25_PEA (P11892) 50S ribosomal protein CL25,
chloroplast precursor, partial (79%)
Length = 726
Score = 27.7 bits (60), Expect = 9.6
Identities = 12/28 (42%), Positives = 18/28 (63%)
Frame = -3
Query: 32 SVCSTWRSSSVPKHHHKFRFQFPLLKFP 59
S+ S ++++ HH+F FQFPLL P
Sbjct: 361 SLSSAA*TNTLSSIHHRFSFQFPLLATP 278
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.324 0.142 0.452
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 24,050,257
Number of Sequences: 63676
Number of extensions: 424151
Number of successful extensions: 2025
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 2010
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2020
length of query: 402
length of database: 12,639,632
effective HSP length: 99
effective length of query: 303
effective length of database: 6,335,708
effective search space: 1919719524
effective search space used: 1919719524
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 60 (27.7 bits)
Medicago: description of AC147741.8


