

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147714.2 - phase: 0 /pseudo
(971 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
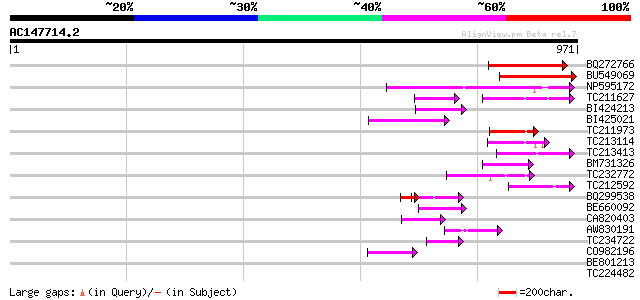
Sequences producing significant alignments: (bits) Value
BQ272766 weakly similar to GP|28558781|gb| pol protein {Cucumis ... 191 2e-48
BU549069 184 2e-46
NP595172 polyprotein [Glycine max] 182 6e-46
TC211627 85 1e-31
BI424213 81 3e-15
BI425021 78 2e-14
TC211973 weakly similar to UP|Q84ZV5 (Q84ZV5) Polyprotein, parti... 74 3e-13
TC213114 weakly similar to UP|Q8W150 (Q8W150) Polyprotein, parti... 69 1e-11
TC213413 weakly similar to UP|Q84ZV5 (Q84ZV5) Polyprotein, parti... 65 2e-10
BM731326 weakly similar to GP|21740635|em OSJNBb0043H09.2 {Oryza... 57 5e-08
TC232772 weakly similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial ... 55 1e-07
TC212592 weakly similar to UP|Q84ZV5 (Q84ZV5) Polyprotein, parti... 55 1e-07
BQ299538 54 3e-07
BE660092 weakly similar to GP|9884624|dbj retroelement pol polyp... 53 6e-07
CA820403 weakly similar to GP|13273463|gb| pol protein integrase... 51 3e-06
AW830191 46 7e-05
TC234722 similar to UP|Q6WAY9 (Q6WAY9) Pol (Fragment), partial (... 46 7e-05
CO982196 45 2e-04
BE801213 weakly similar to GP|6691193|gb| F7F22.17 {Arabidopsis ... 30 0.032
TC224482 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 36 0.072
>BQ272766 weakly similar to GP|28558781|gb| pol protein {Cucumis melo},
partial (9%)
Length = 410
Score = 191 bits (484), Expect = 2e-48
Identities = 92/136 (67%), Positives = 111/136 (80%)
Frame = -3
Query: 820 SYHDKRRKDLEFQEGDHVFLRVTPMTGVXRALKSKKLTPKFIGPYQILERVGTVAYRVGL 879
SYHDKRRKDLEF+ GDHVFLRVTP TGV RALKS KLTP FIGP+QIL++V VAY++ L
Sbjct: 408 SYHDKRRKDLEFEVGDHVFLRVTP*TGVGRALKS*KLTPHFIGPFQILKKVDFVAYQIAL 229
Query: 880 PPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDVQVRDNLTVETLPVRIDDRKVKMLRGKEI 939
PP L++LHNVFHVSQ+ KY+ DPSH++ D VQV++NL ETLP+RI D + K LRGKEI
Sbjct: 228 PPSLTSLHNVFHVSQIHKYINDPSHMVNLDVVQVKENLAHETLPLRIKDMRTKHLRGKEI 49
Query: 940 PLVRVVWTGATSESLT 955
LV+V+W GA+ E T
Sbjct: 48 LLVKVIWGGASGEEAT 1
>BU549069
Length = 615
Score = 184 bits (466), Expect = 2e-46
Identities = 87/132 (65%), Positives = 107/132 (80%)
Frame = -1
Query: 839 LRVTPMTGVXRALKSKKLTPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKY 898
L+VTP TGV +ALKS+KLTP FIG +QIL+R VAY++ LPP LSNLHNVFHVSQLR Y
Sbjct: 615 LKVTPRTGVGQALKSRKLTPHFIGHFQILKRAXPVAYQIALPPSLSNLHNVFHVSQLRMY 436
Query: 899 VPDPSHVIQSDDVQVRDNLTVETLPVRIDDRKVKMLRGKEIPLVRVVWTGATSESLTWEL 958
+ DPSHV++ DDVQV++NLT ETLP+RI+DR+ K LR KE PLV+V+W G + E TWEL
Sbjct: 435 IHDPSHVVKLDDVQVKENLTYETLPLRIEDRRTKHLRRKENPLVKVIWGGTSGEDATWEL 256
Query: 959 ESKMLESYPELF 970
ES+M +YP LF
Sbjct: 255 ESQMRVAYPSLF 220
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 182 bits (462), Expect = 6e-46
Identities = 116/335 (34%), Positives = 171/335 (50%), Gaps = 12/335 (3%)
Frame = +1
Query: 645 ILXR*RKSAHFLXXNISFPVSQXXEIYIKEIVKCMGVPSSIVSDRDPRFTSRFWKSLQEA 704
++ R K AHF+ + E ++ IVK G+P SIVSDRD FTS FW+ L +
Sbjct: 3496 VIDRLTKYAHFIPLKADYNSKVVAEAFMSHIVKLHGIPRSIVSDRDRVFTSTFWQHLFKL 3675
Query: 705 LGSKLRLSSAYHPQTDGQSERTIQSLEDLLRICVLEQGGTWDSHLPLIEFTYNNSYHSSI 764
G+ L +SSAYHPQ+DGQSE + LE LR E W LP EF YN +YH S+
Sbjct: 3676 QGTTLAMSSAYHPQSDGQSEVLNKCLEMYLRCFTYEHPKGWVKALPWAEFWYNTAYHMSL 3855
Query: 765 GMAPFEALYGRRCRTPLCWFESGERVVLGPEIVQQTTEKVQMIQE---KMKASQSRQKSY 821
GM PF ALYGR P + E+ +Q T++ ++ + + +Q K
Sbjct: 3856 GMTPFRALYGRE---PPTLTRQACSIDDPAEVREQLTDRDALLAKLKINLTRAQQVMKRQ 4026
Query: 822 HDKRRKDLEFQEGDHVFLRVTPMTGVXRAL-KSKKLTPKFIGPYQILERVGTVAYRVGLP 880
DK+R D+ FQ GD V +++ P L K++KL+ ++ GP+++L ++G VAY++ L
Sbjct: 4027 ADKKRLDVSFQIGDEVLVKLQPYRQHSAVLRKNQKLSMRYFGPFKVLAKIGDVAYKLEL- 4203
Query: 881 PHLSNLHNVFHVSQLR--------KYVPDPSHVIQSDDVQVRDNLTVETLPVRIDDRKVK 932
P + +H VFHVSQL+ Y+P P V + V PV+I ++
Sbjct: 4204 PSAARIHPVFHVSQLKPFNGTAQDPYLPLPLTVTEMGPVM---------QPVKILASRII 4356
Query: 933 MLRGKEIPLVRVVWTGATSESLTWELESKMLESYP 967
+ +I + V W + TWE + SYP
Sbjct: 4357 IRGHNQIEQILVQWENGLQDEATWEDIEDIKASYP 4461
>TC211627
Length = 1034
Score = 85.1 bits (209), Expect(2) = 1e-31
Identities = 54/158 (34%), Positives = 80/158 (50%), Gaps = 1/158 (0%)
Frame = +3
Query: 810 KMKASQSRQKSYHDKRRKDLEFQEGDHVFLRVTPMTGVXRALKSKKLTPKFIGPYQILER 869
+++ Q K D R+DL F GD V++R+ P KL+ +F GPYQI R
Sbjct: 357 RLQKYQDSMKRIADSHRRDLTFNIGDWVYVRL*PYRQTSIQSTYTKLSKRFYGPYQIQAR 536
Query: 870 VGTVAYRVGLPPHLSNLHNVFHVSQLR-KYVPDPSHVIQSDDVQVRDNLTVETLPVRIDD 928
VG VAYR+ LPP S +H +FHVS L+ + P P ++ ++ V+ P++ D
Sbjct: 537 VGQVAYRLQLPP-TSKIHPIFHVSLLKVHHGPIPPELLALPPFSTTNHPLVQ--PLQFLD 707
Query: 929 RKVKMLRGKEIPLVRVVWTGATSESLTWELESKMLESY 966
K+ IP V V WT E TWE +++ + Y
Sbjct: 708 WKMDESTTPPIPQVLVQWTNLAPEDTTWESWTQLKDIY 821
Score = 71.2 bits (173), Expect(2) = 1e-31
Identities = 38/78 (48%), Positives = 44/78 (55%)
Frame = +2
Query: 693 FTSRFWKSLQEALGSKLRLSSAYHPQTDGQSERTIQSLEDLLRICVLEQGGTWDSHLPLI 752
F S W L G+KLR S+AYHPQTDGQ+E + LE LR V + W L L
Sbjct: 5 FISGLWHELFHISGTKLRFSTAYHPQTDGQTEVINRILEQYLRAFVHDHPQHWFKFLSLA 184
Query: 753 EFTYNNSYHSSIGMAPFE 770
E YN S HS IG +PFE
Sbjct: 185 E*CYNTSVHSGIGFSPFE 238
>BI424213
Length = 426
Score = 80.9 bits (198), Expect = 3e-15
Identities = 39/87 (44%), Positives = 52/87 (58%)
Frame = +1
Query: 695 SRFWKSLQEALGSKLRLSSAYHPQTDGQSERTIQSLEDLLRICVLEQGGTWDSHLPLIEF 754
S FWK+L LG+KL S+ HPQTDGQ++ +SL LLR + +WD +LP +EF
Sbjct: 4 SHFWKTLWAKLGTKLLFSTTCHPQTDGQTKVVNRSLSTLLRALLKGNHKSWDEYLPHVEF 183
Query: 755 TYNNSYHSSIGMAPFEALYGRRCRTPL 781
YN H + +PFE +YG TPL
Sbjct: 184 AYNRGVHRTTKQSPFEVVYGFNPLTPL 264
>BI425021
Length = 426
Score = 77.8 bits (190), Expect = 2e-14
Identities = 48/139 (34%), Positives = 66/139 (46%)
Frame = -1
Query: 615 LDVPXWKWDSISMIL*RVAKYX*RDXRIGGILXR*RKSAHFLXXNISFPVSQXXEIYIKE 674
L VP W+ +SM I ++ R K H S +++
Sbjct: 417 LPVPQRPWEDLSMDFIVGLPPYHGHTTIFVVVNRFSKGIHLGTLPTSHTAHMVASLFLNI 238
Query: 675 IVKCMGVPSSIVSDRDPRFTSRFWKSLQEALGSKLRLSSAYHPQTDGQSERTIQSLEDLL 734
++K G P SIVSDRDP F S FW+ L G+ LR+SSAYHPQTDGQ+E + +E L
Sbjct: 237 VIKLHGFPRSIVSDRDPLFISHFWQDLFRLSGTVLRMSSAYHPQTDGQTEVLNRVIEQYL 58
Query: 735 RICVLEQGGTWDSHLPLIE 753
R V + +P +E
Sbjct: 57 RAFVHGRPRNLGRFIPWVE 1
>TC211973 weakly similar to UP|Q84ZV5 (Q84ZV5) Polyprotein, partial (4%)
Length = 730
Score = 73.9 bits (180), Expect = 3e-13
Identities = 36/84 (42%), Positives = 58/84 (68%), Gaps = 1/84 (1%)
Frame = +1
Query: 823 DKRRKDLEFQEGDHVFLRVTPMTGVXRALK-SKKLTPKFIGPYQILERVGTVAYRVGLPP 881
+KRR+D+E+ GD VFL++ P A + ++KL+P+F P+Q+ +VGT+AY++ LP
Sbjct: 112 NKRRRDIEYVVGD*VFLKMQPYRRRSLAKRINEKLSPRFYAPFQVFNKVGTIAYKLDLPS 291
Query: 882 HLSNLHNVFHVSQLRKYVPDPSHV 905
H+ +H VFHVS L+K P+ H+
Sbjct: 292 HI-KIHPVFHVSLLKKASPNLYHL 360
>TC213114 weakly similar to UP|Q8W150 (Q8W150) Polyprotein, partial (7%)
Length = 810
Score = 68.6 bits (166), Expect = 1e-11
Identities = 42/117 (35%), Positives = 68/117 (57%), Gaps = 11/117 (9%)
Frame = +3
Query: 819 KSYHDKRRKDLEFQEGDHVFLRVTPMTGVXRALK-SKKLTPKFIGPYQILERVGTVAYRV 877
K+Y D+ R+ + GD V+L++ P A K ++KL+P+F GPYQI +++G VA+ +
Sbjct: 6 KAYADRSRRAVTLSVGDWVYLKLQPYRLKSLAKKRNEKLSPRFYGPYQIKKQIGLVAFEL 185
Query: 878 GLPPHLSNLHNVFHVSQLRKYV-----PDPSHVIQSDDV-----QVRDNLTVETLPV 924
LPP +H VFH S L+K V P P ++ S+D+ Q++ L++ L V
Sbjct: 186 DLPP-ARKIHPVFHASLLKKAVAATANPQPLPLMLSEDLSSEFFQLKSKLSITILMV 353
>TC213413 weakly similar to UP|Q84ZV5 (Q84ZV5) Polyprotein, partial (5%)
Length = 761
Score = 64.7 bits (156), Expect = 2e-10
Identities = 41/134 (30%), Positives = 64/134 (47%), Gaps = 1/134 (0%)
Frame = +1
Query: 834 GDHVFLRVTP-MTGVXRALKSKKLTPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHV 892
GD V +++ P G KLT ++ GP+++ ER+G V YR+ L H S +H VFHV
Sbjct: 7 GDWVLVKLRPHRQGSASETTYSKLTKRYYGPFEVQERLGKVVYRLKLTAH-SRIHPVFHV 183
Query: 893 SQLRKYVPDPSHVIQSDDVQVRDNLTVETLPVRIDDRKVKMLRGKEIPLVRVVWTGATSE 952
S L+ +V DP + + V P+ + D K+ +V V W A+ +
Sbjct: 184 SLLKAFVGDP-ETTHAGPLPVMRTEEATNTPLTVIDSKLVPADNGPRRMVLVQWPSASRQ 360
Query: 953 SLTWELESKMLESY 966
+WE + E Y
Sbjct: 361 DASWEDWQVLRERY 402
>BM731326 weakly similar to GP|21740635|em OSJNBb0043H09.2 {Oryza sativa
(japonica cultivar-group)}, partial (3%)
Length = 424
Score = 56.6 bits (135), Expect = 5e-08
Identities = 30/88 (34%), Positives = 51/88 (57%), Gaps = 1/88 (1%)
Frame = +1
Query: 811 MKASQSRQKSYHDKRRKDLEFQEGDHVFLRVTP-MTGVXRALKSKKLTPKFIGPYQILER 869
++ +QS + + R+ + GD V+L++ P G KLT +F GPY ++ +
Sbjct: 25 LERAQSLMVKHANNHRRPHDINVGDWVYLKIRPHRQGSMPPRLHPKLTARFYGPYLVMRQ 204
Query: 870 VGTVAYRVGLPPHLSNLHNVFHVSQLRK 897
VG VA+++ LP + +H VFHVSQL++
Sbjct: 205 VGAVAFQLQLPSE-ARIHPVFHVSQLKR 285
>TC232772 weakly similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial (7%)
Length = 729
Score = 55.5 bits (132), Expect = 1e-07
Identities = 38/155 (24%), Positives = 72/155 (45%), Gaps = 4/155 (2%)
Frame = +1
Query: 749 LPLIEFTYNNSYHSSIGMAPFEALYGRRCRTPLCWFESGERVVLGPEIVQQTTEKVQMIQ 808
LP +EF YN + HS+ PFE +Y TPL + E ++ +
Sbjct: 7 LPHVEFAYNRAVHSTTQHFPFEVVYDFNPLTPLDSLPLSNISGFKHKDAHAKVEYIKRLH 186
Query: 809 EKMKASQSRQKSYH----DKRRKDLEFQEGDHVFLRVTPMTGVXRALKSKKLTPKFIGPY 864
E+ K +++ + +K RK + + D V++ + + + KL P+ GP+
Sbjct: 187 EQAKTQIAKKNESYVKQTNKNRKKVVLEPSDWVWVHMRKERFPKQ--RMSKLQPRGDGPF 360
Query: 865 QILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYV 899
Q+LER+ AY++ +P + + F+++ L +V
Sbjct: 361 QVLERINYNAYKIDIPGEY-EVSSSFNIADLTPFV 462
>TC212592 weakly similar to UP|Q84ZV5 (Q84ZV5) Polyprotein, partial (4%)
Length = 664
Score = 55.5 bits (132), Expect = 1e-07
Identities = 36/112 (32%), Positives = 58/112 (51%)
Frame = +2
Query: 855 KLTPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDVQVR 914
KL+ +F GP+Q++ERVG VAYR+ LP + LH VFH S L+ + +P +
Sbjct: 74 KLSKRFFGPFQVVERVGKVAYRLQLPVD-AKLHPVFHCSLLKPFQGNPPDTAAPLPPTLF 250
Query: 915 DNLTVETLPVRIDDRKVKMLRGKEIPLVRVVWTGATSESLTWELESKMLESY 966
D+ +V V + R V + V V W G + + TWE +++ + +
Sbjct: 251 DHQSVIAPLVILATRTV----NDDDIEVLVQWQGLSPDDATWEKWTELCKEF 394
>BQ299538
Length = 426
Score = 53.9 bits (128), Expect = 3e-07
Identities = 30/88 (34%), Positives = 46/88 (52%)
Frame = +3
Query: 689 RDPRFTSRFWKSLQEALGSKLRLSSAYHPQTDGQSERTIQSLEDLLRICVLEQGGTWDSH 748
R F + + + G+ L++S++YHP DGQ+ LE LR V +Q
Sbjct: 93 RGSNFCEFILEGIVKLPGTYLKMSTSYHP*IDGQTVN--HCLETFLRCFVADQPKM*VQW 266
Query: 749 LPLIEFTYNNSYHSSIGMAPFEALYGRR 776
L E+ YN ++H+S G PFE +YGR+
Sbjct: 267 LSWAEYWYNTNFHASTGTTPFEVVYGRK 350
Score = 45.1 bits (105), Expect = 2e-04
Identities = 19/33 (57%), Positives = 24/33 (72%)
Frame = +1
Query: 669 EIYIKEIVKCMGVPSSIVSDRDPRFTSRFWKSL 701
EI+ KE+V GVP+S++SD DP F S FWK L
Sbjct: 34 EIFTKEVVHLHGVPASVLSDEDPIFVSSFWKEL 132
>BE660092 weakly similar to GP|9884624|dbj retroelement pol polyprotein-like
{Arabidopsis thaliana}, partial (13%)
Length = 378
Score = 53.1 bits (126), Expect = 6e-07
Identities = 24/82 (29%), Positives = 46/82 (55%)
Frame = -3
Query: 700 SLQEALGSKLRLSSAYHPQTDGQSERTIQSLEDLLRICVLEQGGTWDSHLPLIEFTYNNS 759
+L + G R+S+ YHPQT+GQ+E + + ++ +L V W + L + + +
Sbjct: 370 ALLKKYGVVHRVSTPYHPQTNGQAEISNREIKRILEKIVQPSRKDWSTRLDDALWAHRTA 191
Query: 760 YHSSIGMAPFEALYGRRCRTPL 781
Y + IGM+P+ ++G+ C P+
Sbjct: 190 YKAPIGMSPYRVVFGKACHLPV 125
>CA820403 weakly similar to GP|13273463|gb| pol protein integrase region
{Ginkgo biloba}, partial (52%)
Length = 421
Score = 50.8 bits (120), Expect = 3e-06
Identities = 31/76 (40%), Positives = 41/76 (53%), Gaps = 1/76 (1%)
Frame = -3
Query: 671 YIKEIVKCMGVPSSIVSDRDPRFT-SRFWKSLQEALGSKLRLSSAYHPQTDGQSERTIQS 729
+IKE VK G SSIVSD D F S FW L + G+KL+ S AYHPQ D ++ +
Sbjct: 335 FIKEAVKLHGCSSSIVSDWDRLFLIS*FWTELFKMEGTKLKFSLAYHPQPDSHTKVVNRC 156
Query: 730 LEDLLRICVLEQGGTW 745
+E L+ + W
Sbjct: 155 IEMNLQCLTTSKRKQW 108
>AW830191
Length = 372
Score = 46.2 bits (108), Expect = 7e-05
Identities = 28/101 (27%), Positives = 47/101 (45%), Gaps = 2/101 (1%)
Frame = +3
Query: 745 WDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCRTPLCWFESGERVVLGPEIVQQTTEKV 804
W L EF +N +Y++S+ + PF+ LYG C P + E+ T ++
Sbjct: 39 WPKRLSWAEFWFNTNYNNSLKLTPFKVLYG--CDPPHL-LKGAIISSTAEEVNVMTNDRD 209
Query: 805 QMIQEKM--KASQSRQKSYHDKRRKDLEFQEGDHVFLRVTP 843
QM+ + A Q Y D+ R+ + GD V+L++ P
Sbjct: 210 QMLHDLKGNLAKAQNQMKYADRSRRSIPLNVGDWVYLKLQP 332
>TC234722 similar to UP|Q6WAY9 (Q6WAY9) Pol (Fragment), partial (32%)
Length = 482
Score = 46.2 bits (108), Expect = 7e-05
Identities = 20/63 (31%), Positives = 37/63 (57%)
Frame = -3
Query: 715 YHPQTDGQSERTIQSLEDLLRICVLEQGGTWDSHLPLIEFTYNNSYHSSIGMAPFEALYG 774
YHPQT+GQ+E + + ++ +L V+ W L + Y ++ + IG++PF+ +YG
Sbjct: 234 YHPQTNGQAEVSNKEIKRVLENIVVSSRKDWALKLDDAFWAYRIAFKTPIGLSPFQLVYG 55
Query: 775 RRC 777
+ C
Sbjct: 54 KAC 46
>CO982196
Length = 812
Score = 44.7 bits (104), Expect = 2e-04
Identities = 30/85 (35%), Positives = 44/85 (51%), Gaps = 1/85 (1%)
Frame = +1
Query: 614 LLDVPXWKWDSISM-IL*RVAKYX*RDXRIGGILXR*RKSAHFLXXNISFPVSQXXEIYI 672
LL +P W ISM + + K +D I ++ R K AHF + + + E++I
Sbjct: 559 LLPIPTKVWTDISMDFIGGLPKAQGKD-NILVVVDRLTKYAHFFALSHPYTAKEVAELFI 735
Query: 673 KEIVKCMGVPSSIVSDRDPRFTSRF 697
KE+V+ G P+SIVSD F S F
Sbjct: 736 KELVRLHGFPASIVSDXXRLFMSLF 810
>BE801213 weakly similar to GP|6691193|gb| F7F22.17 {Arabidopsis thaliana},
partial (3%)
Length = 416
Score = 30.4 bits (67), Expect(2) = 0.032
Identities = 16/55 (29%), Positives = 30/55 (54%), Gaps = 3/55 (5%)
Frame = +3
Query: 673 KEIVKCMGVPSSIVSDRDPRFTSRFWKSLQEAL---GSKLRLSSAYHPQTDGQSE 724
K I G+P ++SD F ++ L + L + ++ ++YHPQT+GQ++
Sbjct: 36 KNIFSRFGMPRILISDGGSHF---YYSQLNKVLKHDSVRHKVETSYHPQTNGQAK 191
Score = 25.8 bits (55), Expect(2) = 0.032
Identities = 10/37 (27%), Positives = 18/37 (48%)
Frame = +2
Query: 745 WDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCRTPL 781
W S L + + + IG+ PF+ +Y + C P+
Sbjct: 254 WSSKLEDALWACKTAKKTPIGLTPFQMVYRKACHLPV 364
>TC224482 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 669
Score = 36.2 bits (82), Expect = 0.072
Identities = 35/137 (25%), Positives = 52/137 (37%), Gaps = 15/137 (10%)
Frame = +1
Query: 745 WDSHLPLIEFTYNNSYHSSIGMAPFEALYGRRCRTPLCWFE--------SGERVVLGPEI 796
W LP Y S +S G PF +YG P FE E + E
Sbjct: 61 WHEMLPFALHGYRTSVRTSTGATPFSLVYGMEAVLP---FEVEVPSLRILAESGLKESEW 231
Query: 797 VQQTTEKVQMIQEKM-------KASQSRQKSYHDKRRKDLEFQEGDHVFLRVTPMTGVXR 849
Q +++ +I+ K + Q R KS DK+ +F EGD V +++ R
Sbjct: 232 AQTRYDQLNLIEGKRLTAMSHGRLYQQRMKSAFDKKVCLRKFHEGDLVLKKMSHAVKDHR 411
Query: 850 ALKSKKLTPKFIGPYQI 866
K P + GP+ +
Sbjct: 412 G----KWAPNYEGPFVV 450
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.350 0.155 0.524
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 39,369,192
Number of Sequences: 63676
Number of extensions: 511741
Number of successful extensions: 4189
Number of sequences better than 10.0: 47
Number of HSP's better than 10.0 without gapping: 3070
Number of HSP's successfully gapped in prelim test: 126
Number of HSP's that attempted gapping in prelim test: 1072
Number of HSP's gapped (non-prelim): 3258
length of query: 971
length of database: 12,639,632
effective HSP length: 106
effective length of query: 865
effective length of database: 5,889,976
effective search space: 5094829240
effective search space used: 5094829240
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.9 bits)
S2: 64 (29.3 bits)
Medicago: description of AC147714.2


