

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147714.16 - phase: 0
(322 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
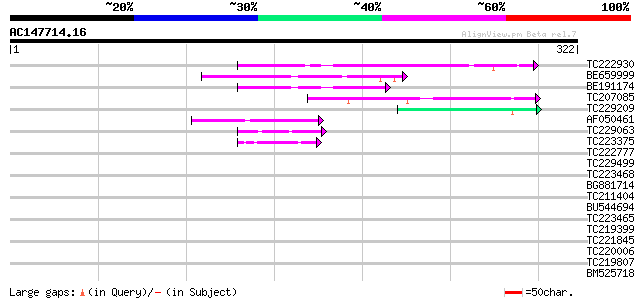
Score E
Sequences producing significant alignments: (bits) Value
TC222930 similar to GB|AAO64916.1|29029074|BT005981 At5g53430 {A... 84 7e-17
BE659999 60 2e-09
BE191174 similar to GP|29029074|gb At5g53430 {Arabidopsis thalia... 58 7e-09
TC207085 similar to UP|Q9SH20 (Q9SH20) F28K19.1, partial (22%) 53 2e-07
TC229209 similar to UP|Q9C5X4 (Q9C5X4) Trithorax-like protein 1,... 46 2e-05
AF050461 44 1e-04
TC229063 similar to GB|AAO64869.1|29028980|BT005934 At2g19260 {A... 44 1e-04
TC223375 weakly similar to UP|Q9Y3Q5 (Q9Y3Q5) PLU-1 protein, par... 42 5e-04
TC222777 weakly similar to GB|AAP68251.1|31711790|BT008812 At2g3... 39 0.002
TC229499 39 0.003
TC223468 similar to GB|AAS92337.1|46402470|BT012421 At5g24330 {A... 39 0.004
BG881714 39 0.004
TC211404 37 0.009
BU544694 weakly similar to GP|13676773|gb PF6 protein {Chlamydom... 37 0.012
TC223465 35 0.046
TC219399 weakly similar to GB|AAP68251.1|31711790|BT008812 At2g3... 35 0.046
TC221845 35 0.046
TC220006 35 0.046
TC219807 similar to UP|Q7ZX31 (Q7ZX31) Ing3-prov protein, partia... 35 0.046
BM525718 similar to PIR|T04592|T045 glycine-rich cell wall struc... 34 0.10
>TC222930 similar to GB|AAO64916.1|29029074|BT005981 At5g53430 {Arabidopsis
thaliana;} , partial (21%)
Length = 865
Score = 84.3 bits (207), Expect = 7e-17
Identities = 56/177 (31%), Positives = 75/177 (41%), Gaps = 6/177 (3%)
Frame = +2
Query: 130 CCVCHSTDANAEDPIVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQCRFKNDIDTDTGP 189
C VC + + I+ C+ C + VH CYG V+ W C C +T
Sbjct: 65 CAVCRWVEDWDYNKIIICNRCQIAVHQECYGARNVQDFT--SWVCKAC------ETPDIK 220
Query: 190 IRCSLCPTKEGAMKQT-TDGKWVHLVCALLVPEVFFVDPEGRE-GIDCSKIPKKRWLEKC 247
C LCP K GA+K T D WVH+ CA PEV F E E + IP +++ C
Sbjct: 221 QECCLCPVKGGALKPTDVDTLWVHVTCAWFRPEVSFASDEKMEPALGILSIPLNSFVKIC 400
Query: 248 YVCGCFDGCALVCSEQKCGLGFHITC----GIKEDLCIEYKEGKKGATVVAGFCKTH 300
+C G C KC FH C G + +L K GK+ T + +C H
Sbjct: 401 VICKEIHGSCTQCC--KCSTYFHAMCASRAGYRMELHCMEKNGKQ-TTRMVSYCAYH 562
>BE659999
Length = 774
Score = 59.7 bits (143), Expect = 2e-09
Identities = 38/125 (30%), Positives = 55/125 (43%), Gaps = 8/125 (6%)
Frame = +3
Query: 110 IEKQSQKQDIEFEVDDDEILCCVCHSTDANAE-DPIVFCDGCNLMVHASCYGNPLVKQIP 168
+E +K + D+ ++ C C D + + ++ C C + VH CYG V
Sbjct: 261 VEAGLEKVVMTCPFDEGQLFCHYCGRGDTGRDSNRLIVCASCKVAVHRKCYG---VHDDI 431
Query: 169 DGDWFCDQCRFKNDIDTDTGPIRCSLCPTKEGAMKQTTDGK---WVHLVCAL----LVPE 221
D W C C+ K D+D P C LCP K GA+K V +CAL + PE
Sbjct: 432 DEAWLCSWCKQKVDVDVSVNP--CVLCPMKGGALKPVNSSAGRCGVSPICALGLLSVDPE 605
Query: 222 VFFVD 226
V+ D
Sbjct: 606 VYIDD 620
>BE191174 similar to GP|29029074|gb At5g53430 {Arabidopsis thaliana}, partial
(8%)
Length = 285
Score = 57.8 bits (138), Expect = 7e-09
Identities = 31/89 (34%), Positives = 42/89 (46%), Gaps = 2/89 (2%)
Frame = +2
Query: 130 CCVCHSTDANAEDPIVFCDGCNLMVHASCYGNPLVKQIPD-GDWFCDQCRFKNDIDTDTG 188
C VC + ++ I+ C+ C + VH CYG K + D W C C +T
Sbjct: 44 CAVCRWVEDWEDNKIIICNRCQIAVHQECYG---AKNVQDFTSWVCRVC------ETPDV 196
Query: 189 PIRCSLCPTKEGAMKQT-TDGKWVHLVCA 216
C LCP K GA+K T + WVH+ CA
Sbjct: 197 ERECCLCPVKGGALKPTDVEMLWVHVTCA 283
>TC207085 similar to UP|Q9SH20 (Q9SH20) F28K19.1, partial (22%)
Length = 1626
Score = 52.8 bits (125), Expect = 2e-07
Identities = 37/145 (25%), Positives = 61/145 (41%), Gaps = 13/145 (8%)
Frame = +3
Query: 170 GDWFCDQCRFKNDIDTDTGPIR--------CSLCPTKEGAMKQTTDGKWVHLVCALLVPE 221
G W+C+ C + + I C+LC GA ++++DG+WVH CA V E
Sbjct: 15 GPWYCELCEDLSSRSSGAAAINFWEKSVVECALCGGTTGAFRKSSDGQWVHAFCAEWVFE 194
Query: 222 VFF----VDP-EGREGIDCSKIPKKRWLEKCYVCGCFDGCALVCSEQKCGLGFHITCGIK 276
F +D EG E + ++ ++ C +C G + C C FH +C +
Sbjct: 195 STFKRGQIDAVEGMETL-------QKGVDICCICHHKHGVCMKCCYGHCQTTFHPSCARR 353
Query: 277 EDLCIEYKEGKKGATVVAGFCKTHS 301
L + + G +C+ HS
Sbjct: 354 AGLYMNVRP-TGGKAQHKAYCEKHS 425
>TC229209 similar to UP|Q9C5X4 (Q9C5X4) Trithorax-like protein 1, partial
(28%)
Length = 1537
Score = 46.2 bits (108), Expect = 2e-05
Identities = 27/94 (28%), Positives = 38/94 (39%), Gaps = 12/94 (12%)
Frame = +2
Query: 221 EVFFVDPEGREGID-CSKIPKKRWLEKCYVCGCFDGCALVCSEQKCGLGFHITCGIKEDL 279
E D + E ID S+I K RW C +CG G + CS C + +H C L
Sbjct: 2 ETCLADVKRMEPIDGLSRISKDRWRLLCSICGVSYGACIQCSNNSCRVAYHPLCARAAGL 181
Query: 280 CIEYK-----------EGKKGATVVAGFCKTHSQ 302
C+E + + + + FCK H Q
Sbjct: 182 CVELENEDRLYLLSVDDDEDQCIRLLSFCKKHRQ 283
>AF050461
Length = 913
Score = 43.9 bits (102), Expect = 1e-04
Identities = 20/76 (26%), Positives = 42/76 (54%), Gaps = 1/76 (1%)
Frame = +1
Query: 104 LPSATPIEKQSQKQDIEFEVDDDEI-LCCVCHSTDANAEDPIVFCDGCNLMVHASCYGNP 162
+ S+ E QS+ + E +V + ++ +C +C DA ED + C C+ +
Sbjct: 682 IKSSGRAEPQSEDESDESDVVEHDVKVCDICG--DAGREDLLAICSRCSDGAEHTYCMRE 855
Query: 163 LVKQIPDGDWFCDQCR 178
+++++P+GDW C++C+
Sbjct: 856 MLEKVPEGDWLCEECK 903
>TC229063 similar to GB|AAO64869.1|29028980|BT005934 At2g19260 {Arabidopsis
thaliana;} , partial (10%)
Length = 1656
Score = 43.5 bits (101), Expect = 1e-04
Identities = 19/51 (37%), Positives = 27/51 (52%)
Frame = +1
Query: 130 CCVCHSTDANAEDPIVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQCRFK 180
C +C D + ++ CD C H SCY NP +K++P +WFC C K
Sbjct: 709 CKICGDLDNSLN--MLLCDHCEDAYHLSCY-NPRLKKLPIDEWFCHSCLIK 852
>TC223375 weakly similar to UP|Q9Y3Q5 (Q9Y3Q5) PLU-1 protein, partial (3%)
Length = 690
Score = 41.6 bits (96), Expect = 5e-04
Identities = 22/48 (45%), Positives = 27/48 (55%)
Frame = +3
Query: 130 CCVCHSTDANAEDPIVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQC 177
C VC TD + +D IV CDGC+ H C P IP G+WFC +C
Sbjct: 3 CRVC-LTDQD-DDRIVLCDGCDHAYHIYCMKPPRT-SIPRGNWFCRKC 137
>TC222777 weakly similar to GB|AAP68251.1|31711790|BT008812 At2g36720
{Arabidopsis thaliana;} , partial (26%)
Length = 1071
Score = 39.3 bits (90), Expect = 0.002
Identities = 17/52 (32%), Positives = 26/52 (49%), Gaps = 7/52 (13%)
Frame = +1
Query: 130 CCVCHSTDANAED----PIVFCDGCNLMVHASCYGN---PLVKQIPDGDWFC 174
C +C S+D + I+ CD C H C + +K++P+GDWFC
Sbjct: 73 CVLCRSSDFSRSGFGPRTIIICDQCEKEYHVGCLRDHKMAYLKELPEGDWFC 228
>TC229499
Length = 915
Score = 38.9 bits (89), Expect = 0.003
Identities = 18/48 (37%), Positives = 23/48 (47%), Gaps = 1/48 (2%)
Frame = +1
Query: 131 CVCHSTDANAED-PIVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQC 177
C+C + +D IV CD C+ H C P IP G WFC +C
Sbjct: 346 CICQVCLTDKDDNKIVLCDACDHAYHVYCM-KPPQNSIPKGKWFCIKC 486
>TC223468 similar to GB|AAS92337.1|46402470|BT012421 At5g24330 {Arabidopsis
thaliana;} , partial (33%)
Length = 507
Score = 38.5 bits (88), Expect = 0.004
Identities = 15/53 (28%), Positives = 26/53 (48%)
Frame = +2
Query: 125 DDEILCCVCHSTDANAEDPIVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQC 177
+D++ C C ++ ++ CD C+ H C P++ +P G WFC C
Sbjct: 80 NDDVSCEECGG--GHSPSKLILCDKCDRGYHLFCL-RPILPSVPKGSWFCPSC 229
>BG881714
Length = 182
Score = 38.5 bits (88), Expect = 0.004
Identities = 18/51 (35%), Positives = 24/51 (46%)
Frame = +1
Query: 132 VCHSTDANAEDPIVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQCRFKND 182
VCH+ E ++ CD C+ H C G L + DGDWFC C +
Sbjct: 4 VCHA--GTDEHLLLLCDLCDTASHTYCVG--LGYTVSDGDWFCHDCAISRE 144
>TC211404
Length = 1050
Score = 37.4 bits (85), Expect = 0.009
Identities = 25/78 (32%), Positives = 38/78 (48%)
Frame = +1
Query: 113 QSQKQDIEFEVDDDEILCCVCHSTDANAEDPIVFCDGCNLMVHASCYGNPLVKQIPDGDW 172
Q K D + D D+ LCC+C++ +A+A+ I C H SCYG + + + +
Sbjct: 562 QDDKVDSVGDTDLDDSLCCICYACEADAQ--IAPCS------HRSCYG-CITRHLLN--- 705
Query: 173 FCDQCRFKNDIDTDTGPI 190
C +C F N TD I
Sbjct: 706 -CQRCFFCNATVTDVSKI 756
>BU544694 weakly similar to GP|13676773|gb PF6 protein {Chlamydomonas
reinhardtii}, partial (0%)
Length = 500
Score = 37.0 bits (84), Expect = 0.012
Identities = 20/66 (30%), Positives = 31/66 (46%)
Frame = +2
Query: 19 IYQQQQQQEEQDLSHCSLLPTKKRKESRNSSLFHTPPSSPTPPPSTYSLPTKKRITALQP 78
++Q Q Q S + T+ R ++L PSSPT PP T + P K + ++P
Sbjct: 158 LHQHQNQSTSTTTSTTTNQSTRTRTNI*CANLXXISPSSPTKPPPTTNHPQPKPPSPIKP 337
Query: 79 HLHHHN 84
HH +
Sbjct: 338 XXHHRH 355
>TC223465
Length = 661
Score = 35.0 bits (79), Expect = 0.046
Identities = 24/96 (25%), Positives = 43/96 (44%), Gaps = 4/96 (4%)
Frame = +3
Query: 87 PNDAVPLIDLNV--EYSPSLPSATPIEKQSQKQDIEFEVDDDEILCCVCHSTDANAEDPI 144
P+D LNV + S L +++ + + + + D C C + P
Sbjct: 21 PDDGSQFEGLNVSGDLSGYLENSSTLFXEECGDEPTLSLQDTGFSCMQCSQMEPA---PD 191
Query: 145 VFCDGCNLMVHASCYGNPLVKQIPD--GDWFCDQCR 178
+FC+ C +++H+ C +P V ++P G W C CR
Sbjct: 192 LFCEICGILIHSQC--SPWV-EVPSRLGSWRCGNCR 290
>TC219399 weakly similar to GB|AAP68251.1|31711790|BT008812 At2g36720
{Arabidopsis thaliana;} , partial (29%)
Length = 1270
Score = 35.0 bits (79), Expect = 0.046
Identities = 15/60 (25%), Positives = 27/60 (45%), Gaps = 7/60 (11%)
Frame = +1
Query: 122 EVDDDEILCCVCHSTDANAED----PIVFCDGCNLMVHASCYGN---PLVKQIPDGDWFC 174
+++ D C +C D + I+ CD C H C + +K++P+G+W C
Sbjct: 178 DIEADLSSCALCRGVDFSRSGFGPRTIILCDQCEKEYHVGCLRDHKMAYLKELPEGNWLC 357
>TC221845
Length = 976
Score = 35.0 bits (79), Expect = 0.046
Identities = 19/60 (31%), Positives = 29/60 (47%), Gaps = 2/60 (3%)
Frame = +3
Query: 121 FEVDDDEILCCVCHSTDANAEDPIVFCDGCNLMVHASCYGNPLVKQIPD--GDWFCDQCR 178
F + D C C + P +FC+ C +++H+ C +P V +IP G W C CR
Sbjct: 390 FSLQDTGFSCMKCSQMEPA---PDLFCEICGILIHSQC--SPWV-EIPSRLGSWRCGNCR 551
>TC220006
Length = 806
Score = 35.0 bits (79), Expect = 0.046
Identities = 19/41 (46%), Positives = 26/41 (63%), Gaps = 1/41 (2%)
Frame = -2
Query: 23 QQQQEEQDLSHCSLLPTKKR-KESRNSSLFHTPPSSPTPPP 62
Q QQ + L+ SLL +K+ ++RNSS FH+ S P PPP
Sbjct: 232 QLQQHQHHLTRLSLLAIEKKLNQNRNSSCFHS--SDPVPPP 116
>TC219807 similar to UP|Q7ZX31 (Q7ZX31) Ing3-prov protein, partial (14%)
Length = 562
Score = 35.0 bits (79), Expect = 0.046
Identities = 17/59 (28%), Positives = 26/59 (43%)
Frame = +2
Query: 119 IEFEVDDDEILCCVCHSTDANAEDPIVFCDGCNLMVHASCYGNPLVKQIPDGDWFCDQC 177
+ VD +E C C+ A +V CD N + +G +K+ P G W+C C
Sbjct: 8 VRLPVDPNEPTYCFCNQVSYGA---MVACDNPNCKIEWFHFGCVGLKEQPKGKWYCSNC 175
>BM525718 similar to PIR|T04592|T045 glycine-rich cell wall structural
protein homolog F23E13.120 - Arabidopsis thaliana,
partial (11%)
Length = 326
Score = 33.9 bits (76), Expect = 0.10
Identities = 23/76 (30%), Positives = 30/76 (39%), Gaps = 7/76 (9%)
Frame = +2
Query: 24 QQQEEQDLSHCSLLPTKKRKESRNSSLFHTPPSS-------PTPPPSTYSLPTKKRITAL 76
+ + Q H +P S +SS HTPPSS P PPP R +
Sbjct: 71 ESRRVQTQHHKYSMPMVPSSSSSSSSSSHTPPSS*PHHLLHPLPPPHPGPSHLPLRPPSR 250
Query: 77 QPHLHHHNNIPNDAVP 92
P L H + PN + P
Sbjct: 251 APGLRRHQSPPNRSPP 298
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.137 0.445
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,699,110
Number of Sequences: 63676
Number of extensions: 423399
Number of successful extensions: 6088
Number of sequences better than 10.0: 236
Number of HSP's better than 10.0 without gapping: 4791
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 5717
length of query: 322
length of database: 12,639,632
effective HSP length: 97
effective length of query: 225
effective length of database: 6,463,060
effective search space: 1454188500
effective search space used: 1454188500
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC147714.16


