

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
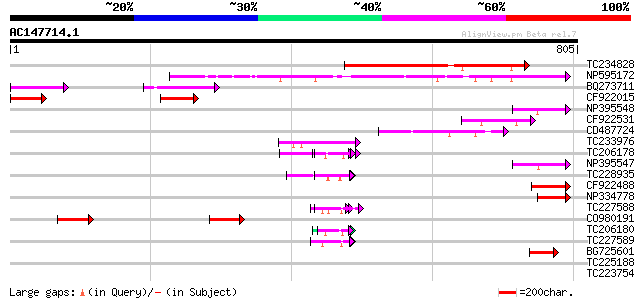
Query= AC147714.1 - phase: 0 /pseudo
(805 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC234828 234 1e-61
NP595172 polyprotein [Glycine max] 146 4e-35
BQ273711 87 2e-17
CF922015 80 5e-15
NP395548 reverse transcriptase [Glycine max] 60 3e-09
CF922531 59 1e-08
CD487724 58 2e-08
TC233976 58 2e-08
TC206178 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, part... 57 4e-08
NP395547 reverse transcriptase [Glycine max] 54 2e-07
TC228935 similar to UP|Q9FHC2 (Q9FHC2) Arabidopsis thaliana geno... 52 8e-07
CF922488 50 3e-06
NP334778 reverse transcriptase [Glycine max] 50 4e-06
TC227588 similar to PIR|T00837|T00837 glycine-rich protein T13L1... 49 1e-05
CO980191 48 2e-05
TC206180 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, part... 48 2e-05
TC227589 46 6e-05
BG725601 similar to PIR|H86337|H86 protein F5M15.26 [imported] -... 45 1e-04
TC225188 similar to UP|Q9FYB7 (Q9FYB7) Splicing factor-like prot... 42 0.001
TC223754 similar to UP|Q86EQ4 (Q86EQ4) Clone ZZD1536 mRNA sequen... 42 0.001
>TC234828
Length = 857
Score = 234 bits (597), Expect = 1e-61
Identities = 120/269 (44%), Positives = 173/269 (63%), Gaps = 7/269 (2%)
Frame = +3
Query: 476 NCNGEGHISSQCTQPKRAPTTGRVFALTGTQTENEDRLIRGTCYINNTPLVAIIDTGATH 535
N G++S+ R RVFA++G++ D LIRG C I + L + D+GATH
Sbjct: 66 NSTRSGNVSNNNISGGRPKVPSRVFAMSGSEAAASDDLIRGKCLIADKLLDVLYDSGATH 245
Query: 536 CFIAFDCVSALGLDLSDMNGEMVVETPAKGSVTTSLVCLKCPLSMFGRDFEMDLVCLPLS 595
FI+ CV LGL +++ +MVV TP VTTS VCLKCP+ + GR F DL+CLPL+
Sbjct: 246 SFISHACVERLGLCATELPYDMVVSTPTSEPVTTSRVCLKCPIIVEGRSFMADLICLPLA 425
Query: 596 GMDVILGMNWLEYNHVLINCFSKSVHFSSVEEESGAEFLSTKQLKQ----LERDGILMFS 651
+DVILGM+WL NH+ ++C K + F G + + ++ LK+ E + + +
Sbjct: 426 HLDVILGMDWLSTNHIFLDCKEKMLVF-------GGDVVPSEPLKEDAANEETEDVRTYM 584
Query: 652 LMATLSIENQAVIDRLPVVCDFPEVFPDEIPDVPPEREVEFSIDLVPGTKPVSMAPYRIS 711
++ ++ +E A + +PVV +FPEVFPD++ ++PPEREVEF ID+VPG PVS+APYR+S
Sbjct: 585 VLFSMYVEEDAEVSCIPVVSEFPEVFPDDVCELPPEREVEFIIDVVPGANPVSIAPYRMS 764
Query: 712 ---ASELKKQLEDLLEKKFVRPSVSPWGA 737
+E+K Q++DLL K+FVRPS SPWGA
Sbjct: 765 PVELAEVKAQVQDLLSKQFVRPSASPWGA 851
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 146 bits (368), Expect = 4e-35
Identities = 153/628 (24%), Positives = 255/628 (40%), Gaps = 59/628 (9%)
Frame = +1
Query: 228 RYDPDGAQTWLKEIERIFRVMQYTEEQKVRFGTHQLAEEADDWWVALLPTLGQEGAVMTW 287
R+D W+ + E+ F + ++ + L ++ W+ L T +W
Sbjct: 310 RFDGKNVMDWIFKAEQFFDYYATPDADRLIIASVHLDQDVVPWYQMLQKT----EPFSSW 477
Query: 288 AVFRREFLRRYFLEDVRGKKEIEFLELKQGNMSVTEYAAKFVELSKFYPHYTAENAEFSR 347
F R L F +L Q + +V EY +F L +AE
Sbjct: 478 QAFTRA-LELDFGPSAYDCPRATLFKLNQ-SATVNEYYMQFTALVNRVDGLSAE--AILD 645
Query: 348 CIKFENGLRLDIKRAIGYQQLRVFQDLVNSCRIYEE--------------------DTKA 387
C F +GL+ +I R + + R V +++EE +T A
Sbjct: 646 C--FVSGLQEEISRDVKAMEPRTLTKAVALAKLFEEKYTSPPKTKTFSNLARNFTSNTSA 819
Query: 388 HYKVVNERKGKGQQSRPKPYSAPADKGKQKMVDVRRPKKKDAAEI-------VCFNCGEK 440
K + P P K + + KK AEI +C+ C EK
Sbjct: 820 TQKYPPTNQKNDNPKPNLPPLLPTPSTKPFNLRNQNIKKISPAEIQLRREKNLCYFCDEK 999
Query: 441 GHKSNACPEEIKKCVRCGKKGHVVADCNRTDIVCFNCNGEGHISSQCTQPKRAPTTGRVF 500
++ CP ++ + D ++TD T+
Sbjct: 1000FSPAHKCPNRQVMLLQLEE-----TDEDQTDE-----------QVMVTEEANMDDDTHHL 1131
Query: 501 ALTGTQTENEDRLIRGTCYINNTPLVAIIDTGATHCFIAFDCVSALGLDLSDMNGEMVVE 560
+L + N IR T + + ++D G++ FI L L + V+
Sbjct: 1132SLNAMRGSNGVGTIRFTGQVGGIAVKILVDGGSSDNFIQPRVAQVLKLPVEPAPNLRVLV 1311
Query: 561 TPAKGSVTTSL-VCLKCPLSMFGRDFEMDLVCLPLSGMDVILGMNWL--------EYNHV 611
G + ++ + + PL + G++ ++ + L +SG DVILG WL +Y +
Sbjct: 1312--GNGQILSAEGIVQQLPLHIQGQEVKVPVYLLQISGADVILGSTWLATLGPHVADYAAL 1485
Query: 612 LINCFSKSVHFSSVEEESGAEFLSTKQLKQLERDGILMFSLMATLSIEN----QAVIDRL 667
+ F F +++ E +E + QL R L T SIE Q + +
Sbjct: 1486TLKFFQND-KFITLQGEGNSE-ATQAQLHHFRR-------LQNTKSIEECFAIQLIQKEV 1638
Query: 668 P--VVCDFPEVFPDEIP--------------DVPPEREVEFSIDLVPGTKPVSMAPYRI- 710
P + D P E+ +PP+RE + +I L G+ PV + PYR
Sbjct: 1639PEDTLKDLPTNIDPELAILLHTYAQVFAVPASLPPQREQDHAIPLKQGSGPVKVRPYRYP 1818
Query: 711 --SASELKKQLEDLLEKKFVRPSVSPWGASVLLVKKKDGSMRLCIDYRQLNKVTIKNRYP 768
+++K ++++L + ++PS SP+ +LLVKKKDGS R C DYR LN +T+K+ +P
Sbjct: 1819HTQKDQIEKMIQEMLVQGIIQPSNSPFSLPILLVKKKDGSWRFCTDYRALNAITVKDSFP 1998
Query: 769 LPRIDDLMDQLVGARVFSKIDLRSGYHQ 796
+P +D+L+D+L GA+ FSK+DLRSGYHQ
Sbjct: 1999MPTVDELLDELHGAQYFSKLDLRSGYHQ 2082
>BQ273711
Length = 409
Score = 87.4 bits (215), Expect = 2e-17
Identities = 44/109 (40%), Positives = 61/109 (55%)
Frame = -2
Query: 190 LQAVAQAVGQQPNVNAGANAEARMLETFMKKNPPTFKGRYDPDGAQTWLKEIERIFRVMQ 249
+ AV + + Q+ +V AE R L F K +PP F G YDP+GA+ WL E E+IF M
Sbjct: 318 ITAVLETLVQERDVEP---AEYRGLMAFRKNHPPKFSGDYDPEGARLWLAETEKIFEAMG 148
Query: 250 YTEEQKVRFGTHQLAEEADDWWVALLPTLGQEGAVMTWAVFRREFLRRY 298
EE KV + T L EA++WW + P+ G V+ W F+ +FL Y
Sbjct: 147 CLEEHKVLYATFMLQGEAENWWKFVKPSFVAPGGVIPWNAFKDKFLENY 1
Score = 84.7 bits (208), Expect = 1e-16
Identities = 36/82 (43%), Positives = 48/82 (57%)
Frame = -2
Query: 2 FMKKNPPTFKGRYDPDGAQTWLKEIERIFRVMQCTEEQKVRFGTHQLAEAADDWWVALLP 61
F K +PP F G YDP+GA+ WL E E+IF M C EE KV + T L A++WW + P
Sbjct: 246 FRKNHPPKFSGDYDPEGARLWLAETEKIFEAMGCLEEHKVLYATFMLQGEAENWWKFVKP 67
Query: 62 TLGQEGAVVTWAVFRREFLRRY 83
+ G V+ W F+ +FL Y
Sbjct: 66 SFVAPGGVIPWNAFKDKFLENY 1
>CF922015
Length = 172
Score = 79.7 bits (195), Expect = 5e-15
Identities = 34/54 (62%), Positives = 43/54 (78%)
Frame = -3
Query: 214 LETFMKKNPPTFKGRYDPDGAQTWLKEIERIFRVMQYTEEQKVRFGTHQLAEEA 267
L+ F + NPPTFKG YDP+GA+ WL+EIE+IFRVM+ + QKV F TH LA+EA
Sbjct: 164 LDRFQRNNPPTFKGGYDPEGAEAWLREIEKIFRVMECQDHQKVLFATHMLADEA 3
Score = 79.7 bits (195), Expect = 5e-15
Identities = 33/51 (64%), Positives = 41/51 (79%)
Frame = -3
Query: 2 FMKKNPPTFKGRYDPDGAQTWLKEIERIFRVMQCTEEQKVRFGTHQLAEAA 52
F + NPPTFKG YDP+GA+ WL+EIE+IFRVM+C + QKV F TH LA+ A
Sbjct: 155 FQRNNPPTFKGGYDPEGAEAWLREIEKIFRVMECQDHQKVLFATHMLADEA 3
>NP395548 reverse transcriptase [Glycine max]
Length = 762
Score = 60.5 bits (145), Expect = 3e-09
Identities = 35/101 (34%), Positives = 56/101 (54%), Gaps = 19/101 (18%)
Frame = +1
Query: 715 LKKQLEDLLEKKFVRP-SVSPWGASVLLVKKKDG------------------SMRLCIDY 755
++K++ LLE + P S S W + VL+V KK+G S +LCIDY
Sbjct: 1 VRKEVLKLLEVGLIYPISDSAWVSPVLVVSKKEGMTVIRNEKNDLIPTRTVTSWKLCIDY 180
Query: 756 RQLNKVTIKNRYPLPRIDDLMDQLVGARVFSKIDLRSGYHQ 796
R+LN+ T K+ +PLP +D ++++L G + +D GY+Q
Sbjct: 181 RKLNEATRKDHFPLPFMDQMLERLAGHAYYCFLDAYFGYNQ 303
>CF922531
Length = 602
Score = 58.5 bits (140), Expect = 1e-08
Identities = 42/114 (36%), Positives = 63/114 (54%), Gaps = 9/114 (7%)
Frame = -2
Query: 642 LERDGILMFSLMATLSIENQAVIDRLP-----VVCDFPEVFPDEIP-DVPPEREVEFSID 695
L++ L+ S +LS I+ LP ++ +F ++FP EIP +PP R +E ID
Sbjct: 343 LKQSFYLLLSRETSLSTVTIPTIETLPPKVQELLHEFGDIFPKEIPLGLPPLRGIEHQID 164
Query: 696 LVPGTKPVSMAPYRISASELKK---QLEDLLEKKFVRPSVSPWGASVLLVKKKD 746
LVP + YR + E K+ Q+++LLEK +V+ S+S VLLV KKD
Sbjct: 163 LVPRASLPNRPTYRTNPQETKEIESQVKELLEKGWVQESLSLCVVLVLLVPKKD 2
>CD487724
Length = 676
Score = 57.8 bits (138), Expect = 2e-08
Identities = 53/207 (25%), Positives = 92/207 (43%), Gaps = 22/207 (10%)
Frame = +3
Query: 524 PLVAIIDTGATHCFIAFDCVSALGLDLSDMNGEMVVETPAKGSVTTSLVCLKCPLSMFGR 583
PLV ++D G+TH F+ VS LGL V+ T+ +C P+S+
Sbjct: 21 PLVYLVDGGSTHNFVQQPLVSQLGLPCRSTPPLRVMVGNGHHLKCTT-ICEAIPISIQNI 197
Query: 584 DFEMDLVCLPLSGMDVILGMNWLE-YNHVLINCFSKSVHF--------SSVEEESGAEFL 634
+F + L LP+ G +++LG+ WL+ +L++ S S+ F E E+ L
Sbjct: 198 EFLVHLYVLPIVGANIVLGVQWLKTLGPILVDYNSLSMQFFYQHRLVQLKGESEAQLGLL 377
Query: 635 STKQLKQLERDGILMFSLMATLSIEN-------------QAVIDRLPVVCDFPEVFPDEI 681
+ QL++L + + + EN Q ++D+ + P+
Sbjct: 378 NHHQLRRLHQTHEPVTYFHIAILTENTSPTSSPPLPQPIQHLLDQFSALFQ*PQ------ 539
Query: 682 PDVPPEREVEFSIDLVPGTKPVSMAPY 708
+PP RE + I L+P ++PV+M Y
Sbjct: 540 -GLPPARETDHHIHLLP*SEPVNMRLY 617
>TC233976
Length = 763
Score = 57.8 bits (138), Expect = 2e-08
Identities = 36/126 (28%), Positives = 58/126 (45%), Gaps = 10/126 (7%)
Frame = +1
Query: 382 EEDTKAHYKVVNERKGKGQQ----SRPKPYSAPADK----GKQKMVDVRRPKKKDAAEI- 432
+++ A YK N GK ++ SR KPYSAP + G Q+ + +
Sbjct: 25 QQEKVAFYKNANASHGKEKKPMTHSRAKPYSAPLEYENHYGGQRTSGGHHLAGGSSQLVN 204
Query: 433 -VCFNCGEKGHKSNACPEEIKKCVRCGKKGHVVADCNRTDIVCFNCNGEGHISSQCTQPK 491
V G G + A +C +CG+ GH +C ++ CFN G+GH+S+ C P+
Sbjct: 205 RVSQPAGRGGSGAPAIVTTPLRCRKCGRLGHNAHECTYREVTCFNYQGKGHLSTNCPHPR 384
Query: 492 RAPTTG 497
+ +G
Sbjct: 385 KEKRSG 402
>TC206178 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, partial (69%)
Length = 1138
Score = 56.6 bits (135), Expect = 4e-08
Identities = 25/65 (38%), Positives = 38/65 (58%), Gaps = 6/65 (9%)
Frame = +2
Query: 433 VCFNCGEKGHKSNACPEEIKKCVRCGKKGHVVADCNRTDI------VCFNCNGEGHISSQ 486
+C+NC E GH +++CP E C CGK GH +C+ + +C NC +GHI+++
Sbjct: 455 LCWNCKEPGHMASSCPNE-GICHTCGKAGHRARECSAPPMPPGDLRLCNNCYKQGHIAAE 631
Query: 487 CTQPK 491
CT K
Sbjct: 632 CTNEK 646
Score = 55.1 bits (131), Expect = 1e-07
Identities = 29/118 (24%), Positives = 55/118 (46%), Gaps = 3/118 (2%)
Frame = +2
Query: 384 DTKAHYKVVNERKGKGQQSRPKPYSAP--ADKGKQKMVDVRRPKKKD-AAEIVCFNCGEK 440
+ K +K+ ++ + + + P +D+ + RR ++ + + +C NC
Sbjct: 185 ERKKKWKMSSDSRSRSRSRSRSPMDRKIRSDRFSYRDAPYRRDSRRGFSRDNLCKNCKRP 364
Query: 441 GHKSNACPEEIKKCVRCGKKGHVVADCNRTDIVCFNCNGEGHISSQCTQPKRAPTTGR 498
GH + CP + C CG GH+ ++C T +C+NC GH++S C T G+
Sbjct: 365 GHYARECPN-VAICHNCGLPGHIASECT-TKSLCWNCKEPGHMASSCPNEGICHTCGK 532
Score = 52.4 bits (124), Expect = 8e-07
Identities = 21/63 (33%), Positives = 34/63 (53%), Gaps = 6/63 (9%)
Frame = +2
Query: 431 EIVCFNCGEKGHKSNAC------PEEIKKCVRCGKKGHVVADCNRTDIVCFNCNGEGHIS 484
E +C CG+ GH++ C P +++ C C K+GH+ A+C + C NC GH++
Sbjct: 506 EGICHTCGKAGHRARECSAPPMPPGDLRLCNNCYKQGHIAAECT-NEKACNNCRKTGHLA 682
Query: 485 SQC 487
C
Sbjct: 683 RDC 691
Score = 41.2 bits (95), Expect = 0.002
Identities = 24/85 (28%), Positives = 30/85 (35%), Gaps = 25/85 (29%)
Frame = +2
Query: 431 EIVCFNCGEKGHKSNACPEEIKKCVRCGKKGHVVADCNRT-------------------- 470
E C NC + GH + CP + C C GHV C +
Sbjct: 641 EKACNNCRKTGHLARDCPND-PICNLCNVSGHVARQCPKANVLGDRSGGGGGGGGARGGG 817
Query: 471 -----DIVCFNCNGEGHISSQCTQP 490
D+VC NC GH+S C P
Sbjct: 818 GGGYRDVVCRNCQQLGHMSRDCMGP 892
Score = 39.3 bits (90), Expect = 0.007
Identities = 14/37 (37%), Positives = 21/37 (55%)
Frame = +2
Query: 431 EIVCFNCGEKGHKSNACPEEIKKCVRCGKKGHVVADC 467
++VC NC + GH S C + C CG +GH+ +C
Sbjct: 833 DVVCRNCQQLGHMSRDCMGPLMICHNCGGRGHLAYEC 943
Score = 38.9 bits (89), Expect = 0.009
Identities = 13/34 (38%), Positives = 20/34 (58%)
Frame = +2
Query: 454 CVRCGKKGHVVADCNRTDIVCFNCNGEGHISSQC 487
C C + GH+ DC ++C NC G GH++ +C
Sbjct: 842 CRNCQQLGHMSRDCMGPLMICHNCGGRGHLAYEC 943
>NP395547 reverse transcriptase [Glycine max]
Length = 762
Score = 54.3 bits (129), Expect = 2e-07
Identities = 34/101 (33%), Positives = 52/101 (50%), Gaps = 19/101 (18%)
Frame = +1
Query: 715 LKKQLEDLLEKKFVRP-SVSPWGASVLLVKKKDGSM------------------RLCIDY 755
++K++ LLE + P S S W + V +V KK G R+CIDY
Sbjct: 1 VRKEVFKLLEAGLIYPISDSSWVSPVQVVPKKGGMTVVKNDRNELIPTRRVTRWRMCIDY 180
Query: 756 RQLNKVTIKNRYPLPRIDDLMDQLVGARVFSKIDLRSGYHQ 796
R+LN+ T K+ YPLP +D ++ +L + +D SGY+Q
Sbjct: 181 RKLNEATRKDHYPLPFMDQMLKRLARQSFYRFLDGYSGYNQ 303
>TC228935 similar to UP|Q9FHC2 (Q9FHC2) Arabidopsis thaliana genomic DNA,
chromosome 5, TAC clone:K24M7, partial (36%)
Length = 1224
Score = 52.4 bits (124), Expect = 8e-07
Identities = 23/70 (32%), Positives = 31/70 (43%), Gaps = 13/70 (18%)
Frame = +1
Query: 433 VCFNCGEKGHKSNACPE------EIKKCVRCGKKGHVVADC-------NRTDIVCFNCNG 479
+C C +GH++ CPE + K C CG+ GH + C CF CN
Sbjct: 343 ICLRCRRRGHRAKNCPEVLDGAKDAKYCYNCGENGHALTQCLHPLQEGGTKFAECFVCNQ 522
Query: 480 EGHISSQCTQ 489
GH+S C Q
Sbjct: 523 RGHLSKNCPQ 552
Score = 48.9 bits (115), Expect = 9e-06
Identities = 31/109 (28%), Positives = 46/109 (41%), Gaps = 11/109 (10%)
Frame = +1
Query: 393 NERKGKGQQSRPKPYSAPADKGKQKMVDVRRPKKKDAAEIVCFNCGEKGHKSNACPEEI- 451
+++K K + S K G +K +R P K CF C H + CPE+
Sbjct: 154 DKKKKKNKNSAFKRKRPEPKPGSRKRHLLRVPGMKPGES--CFICKAMDHIAKLCPEKAE 327
Query: 452 ----KKCVRCGKKGHVVADCN------RTDIVCFNCNGEGHISSQCTQP 490
K C+RC ++GH +C + C+NC GH +QC P
Sbjct: 328 WEKNKICLRCRRRGHRAKNCPEVLDGAKDAKYCYNCGENGHALTQCLHP 474
>CF922488
Length = 741
Score = 50.4 bits (119), Expect = 3e-06
Identities = 24/55 (43%), Positives = 36/55 (64%)
Frame = +3
Query: 742 VKKKDGSMRLCIDYRQLNKVTIKNRYPLPRIDDLMDQLVGARVFSKIDLRSGYHQ 796
V K+DG + +C+DYR LN + K+++PLP I+ L+D FS +D SGY+Q
Sbjct: 3 VLKEDGKV*MCVDYRDLN*ASPKDKFPLPHINVLVDNTTSFSQFSFMDGFSGYNQ 167
>NP334778 reverse transcriptase [Glycine max]
Length = 431
Score = 50.1 bits (118), Expect = 4e-06
Identities = 22/47 (46%), Positives = 32/47 (67%)
Frame = +3
Query: 750 RLCIDYRQLNKVTIKNRYPLPRIDDLMDQLVGARVFSKIDLRSGYHQ 796
R+C+DYR LN+ + K+ +PLP ID LM + +FS +D SGY+Q
Sbjct: 3 RMCVDYRDLNRASPKDNFPLPHIDILMANMASFALFSFMDGFSGYNQ 143
>TC227588 similar to PIR|T00837|T00837 glycine-rich protein T13L16.11 -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(10%)
Length = 1300
Score = 48.5 bits (114), Expect = 1e-05
Identities = 25/71 (35%), Positives = 36/71 (50%), Gaps = 2/71 (2%)
Frame = +1
Query: 434 CFNCGEKGHKSNACP--EEIKKCVRCGKKGHVVADCNRTDIVCFNCNGEGHISSQCTQPK 491
CFNCGE+GH + C + K C CG GH C++ CF C GH + C P+
Sbjct: 61 CFNCGEEGHAAVNCSAVKRKKPCYVCGCLGHNARQCSKVQ-DCFICKKGGHRAKDC--PE 231
Query: 492 RAPTTGRVFAL 502
+ +T + A+
Sbjct: 232 KHTSTSKSIAI 264
Score = 47.8 bits (112), Expect = 2e-05
Identities = 27/75 (36%), Positives = 34/75 (45%), Gaps = 14/75 (18%)
Frame = +1
Query: 428 DAAEIVCFNCGEKGH-----KSNACPEEIKKCVRCGKKGHVVADCNR---------TDIV 473
D EI C+ C GH +A P EI C +CG+ GH C+R T
Sbjct: 328 DLKEIQCYVCKRLGHLCCVNTDDATPGEIS-CYKCGQLGHTGLACSRLRDEITSGATPSS 504
Query: 474 CFNCNGEGHISSQCT 488
CF C EGH + +CT
Sbjct: 505 CFKCGEEGHFARECT 549
Score = 42.7 bits (99), Expect = 6e-04
Identities = 20/64 (31%), Positives = 30/64 (46%), Gaps = 14/64 (21%)
Frame = +1
Query: 434 CFNCGEKGHKSNACPEE-------IKKCVRCGKKGHVVADCNR-------TDIVCFNCNG 479
CF C + GH++ CPE+ I C++CG GH + C +I C+ C
Sbjct: 184 CFICKKGGHRAKDCPEKHTSTSKSIAICLKCGNSGHDIFSCRNDYSQDDLKEIQCYVCKR 363
Query: 480 EGHI 483
GH+
Sbjct: 364 LGHL 375
Score = 32.3 bits (72), Expect = 0.85
Identities = 18/68 (26%), Positives = 28/68 (40%)
Frame = +1
Query: 434 CFNCGEKGHKSNACPEEIKKCVRCGKKGHVVADCNRTDIVCFNCNGEGHISSQCTQPKRA 493
CF CGE+GH + C IK R + H ++ + + + KR+
Sbjct: 505 CFKCGEEGHFARECTSSIKSGKRNWESSHTKDKRSQKENDYMGNRSASNDMVGARRKKRS 684
Query: 494 PTTGRVFA 501
PT R F+
Sbjct: 685 PTEERGFS 708
>CO980191
Length = 802
Score = 47.8 bits (112), Expect = 2e-05
Identities = 24/50 (48%), Positives = 30/50 (60%)
Frame = +1
Query: 69 VVTWAVFRREFLRRYFPEDVRGKKEIEFLELKQGNMSVTEYAAQFVELSK 118
V+T VF+ FL ++ ED+ K+ EFLELK GNM V Y F EL K
Sbjct: 361 VIT*EVFKEIFLEKHLHEDI*NHKKTEFLELKHGNMIVANYLVNFDELPK 510
Score = 46.6 bits (109), Expect = 4e-05
Identities = 24/50 (48%), Positives = 30/50 (60%)
Frame = +1
Query: 284 VMTWAVFRREFLRRYFLEDVRGKKEIEFLELKQGNMSVTEYAAKFVELSK 333
V+T VF+ FL ++ ED+ K+ EFLELK GNM V Y F EL K
Sbjct: 361 VIT*EVFKEIFLEKHLHEDI*NHKKTEFLELKHGNMIVANYLVNFDELPK 510
>TC206180 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, partial (42%)
Length = 655
Score = 47.8 bits (112), Expect = 2e-05
Identities = 19/57 (33%), Positives = 31/57 (54%), Gaps = 6/57 (10%)
Frame = +3
Query: 437 CGEKGHKSNAC------PEEIKKCVRCGKKGHVVADCNRTDIVCFNCNGEGHISSQC 487
CG+ GH++ C P +++ C C K+GH+ A+C + C NC GH++ C
Sbjct: 9 CGKAGHRARECSAPPMPPGDLRLCNNCYKQGHIAAECT-NEKACNNCRKTGHLARDC 176
Score = 42.4 bits (98), Expect = 8e-04
Identities = 24/82 (29%), Positives = 30/82 (36%), Gaps = 22/82 (26%)
Frame = +3
Query: 431 EIVCFNCGEKGHKSNACPEEIKKCVRCGKKGHVVADCNRT-------------------- 470
E C NC + GH + CP + C C GHV C +
Sbjct: 126 EKACNNCRKTGHLARDCPND-PICNLCNVSGHVARQCPKANVLGDXXGGGGGARGGGGGG 302
Query: 471 --DIVCFNCNGEGHISSQCTQP 490
D+VC NC GH+S C P
Sbjct: 303 YRDVVCRNCQQLGHMSRDCMGP 368
Score = 39.7 bits (91), Expect = 0.005
Identities = 19/77 (24%), Positives = 29/77 (36%), Gaps = 22/77 (28%)
Frame = +3
Query: 433 VCFNCGEKGHKSNACPE----------------------EIKKCVRCGKKGHVVADCNRT 470
+C C GH + CP+ C C + GH+ DC
Sbjct: 189 ICNLCNVSGHVARQCPKANVLGDXXGGGGGARGGGGGGYRDVVCRNCQQLGHMSRDCMGP 368
Query: 471 DIVCFNCNGEGHISSQC 487
++C NC G GH++ +C
Sbjct: 369 LMICHNCGGRGHLAYEC 419
Score = 39.3 bits (90), Expect = 0.007
Identities = 14/37 (37%), Positives = 21/37 (55%)
Frame = +3
Query: 431 EIVCFNCGEKGHKSNACPEEIKKCVRCGKKGHVVADC 467
++VC NC + GH S C + C CG +GH+ +C
Sbjct: 309 DVVCRNCQQLGHMSRDCMGPLMICHNCGGRGHLAYEC 419
Score = 36.2 bits (82), Expect = 0.059
Identities = 15/41 (36%), Positives = 23/41 (55%), Gaps = 6/41 (14%)
Frame = +3
Query: 457 CGKKGHVVADCNRTDI------VCFNCNGEGHISSQCTQPK 491
CGK GH +C+ + +C NC +GHI+++CT K
Sbjct: 9 CGKAGHRARECSAPPMPPGDLRLCNNCYKQGHIAAECTNEK 131
>TC227589
Length = 547
Score = 46.2 bits (108), Expect = 6e-05
Identities = 23/64 (35%), Positives = 30/64 (45%), Gaps = 2/64 (3%)
Frame = +2
Query: 428 DAAEIVCFNCGEKGHKSNAC--PEEIKKCVRCGKKGHVVADCNRTDIVCFNCNGEGHISS 485
D++ CFNCGE GH + C + K C CG GH C + CF C GH +
Sbjct: 29 DSSWGACFNCGEDGHAAVNCSAAKRKKPCYVCGGLGHNARQCTKAQ-DCFICKKGGHRAK 205
Query: 486 QCTQ 489
C +
Sbjct: 206 DCLE 217
Score = 44.3 bits (103), Expect = 2e-04
Identities = 25/77 (32%), Positives = 33/77 (42%), Gaps = 14/77 (18%)
Frame = +2
Query: 428 DAAEIVCFNCGEKGH-----KSNACPEEIKKCVRCGKKGHVVADCNR---------TDIV 473
D EI C+ C GH +A P EI C +CG+ GH C++ T
Sbjct: 314 DLKEIQCYVCKRVGHLCCVNTDDATPGEIS-CYKCGQLGHTGLACSKLPDEITSAATPSS 490
Query: 474 CFNCNGEGHISSQCTQP 490
C C GH + +CT P
Sbjct: 491 CCKCGEAGHFAQECTSP 541
Score = 39.7 bits (91), Expect = 0.005
Identities = 22/89 (24%), Positives = 39/89 (43%), Gaps = 12/89 (13%)
Frame = +2
Query: 413 KGKQKMVD-VRRPKKKDAAEIVCFNCGEKGHKSNAC-----PEEIK--KCVRCGKKGHVV 464
KG + D + + + + +C CG GH +C P+++K +C C + GH+
Sbjct: 185 KGGHRAKDCLEKHTSRSKSVAICLKCGNSGHDMFSCRNDYSPDDLKEIQCYVCKRVGHLC 364
Query: 465 A----DCNRTDIVCFNCNGEGHISSQCTQ 489
D +I C+ C GH C++
Sbjct: 365 CVNTDDATPGEISCYKCGQLGHTGLACSK 451
Score = 38.1 bits (87), Expect = 0.015
Identities = 18/64 (28%), Positives = 29/64 (45%), Gaps = 14/64 (21%)
Frame = +2
Query: 434 CFNCGEKGHKSNACPEE-------IKKCVRCGKKGHVVADCNR-------TDIVCFNCNG 479
CF C + GH++ C E+ + C++CG GH + C +I C+ C
Sbjct: 170 CFICKKGGHRAKDCLEKHTSRSKSVAICLKCGNSGHDMFSCRNDYSPDDLKEIQCYVCKR 349
Query: 480 EGHI 483
GH+
Sbjct: 350 VGHL 361
>BG725601 similar to PIR|H86337|H86 protein F5M15.26 [imported] - Arabidopsis
thaliana, partial (1%)
Length = 285
Score = 45.1 bits (105), Expect = 1e-04
Identities = 21/42 (50%), Positives = 27/42 (64%)
Frame = -3
Query: 738 SVLLVKKKDGSMRLCIDYRQLNKVTIKNRYPLPRIDDLMDQL 779
SV++VKK +G R+C DY LN K+ YPLP ID + D L
Sbjct: 127 SVVMVKKPNGKWRICTDYIDLN*ACPKDAYPLPNIDHMTDGL 2
>TC225188 similar to UP|Q9FYB7 (Q9FYB7) Splicing factor-like protein, partial
(62%)
Length = 1131
Score = 41.6 bits (96), Expect = 0.001
Identities = 22/67 (32%), Positives = 32/67 (46%), Gaps = 2/67 (2%)
Frame = +1
Query: 434 CFNCGEKGHKSNACP--EEIKKCVRCGKKGHVVADCNRTDIVCFNCNGEGHISSQCTQPK 491
CFNCG GH + C + KC RCG++GH+ +C + + G S+
Sbjct: 364 CFNCGIDGHWARDCKAGDWKNKCYRCGERGHIEKNCKNSP---KKLSQRGRRLSRSPVRS 534
Query: 492 RAPTTGR 498
R+P GR
Sbjct: 535 RSPRRGR 555
Score = 37.7 bits (86), Expect = 0.020
Identities = 18/49 (36%), Positives = 21/49 (42%), Gaps = 3/49 (6%)
Frame = +1
Query: 453 KCVRCGKKGHVVADCNRTD--IVCFNCNGEGHISSQC-TQPKRAPTTGR 498
+C CG GH DC D C+ C GHI C PK+ GR
Sbjct: 361 RCFNCGIDGHWARDCKAGDWKNKCYRCGERGHIEKNCKNSPKKLSQRGR 507
>TC223754 similar to UP|Q86EQ4 (Q86EQ4) Clone ZZD1536 mRNA sequence, partial
(22%)
Length = 742
Score = 41.6 bits (96), Expect = 0.001
Identities = 22/80 (27%), Positives = 28/80 (34%), Gaps = 24/80 (30%)
Frame = +2
Query: 434 CFNCGEKGHKSNACPEEIKK---------CVRCGKKGHVVADCNRT-------------- 470
CF CG GH + C C RCG+ GH+ DC
Sbjct: 230 CFRCGGFGHMARDCATGKGNIGGGGSGGGCFRCGEVGHLARDCGMEGGRFGGGGGSGGGG 409
Query: 471 -DIVCFNCNGEGHISSQCTQ 489
CFNC GH + +C +
Sbjct: 410 GKSTCFNCGKPGHFARECVE 469
Score = 41.6 bits (96), Expect = 0.001
Identities = 17/46 (36%), Positives = 24/46 (51%), Gaps = 10/46 (21%)
Frame = +2
Query: 454 CVRCGKKGHVVADCNRTDIV----------CFNCNGEGHISSQCTQ 489
C +CG+ GH+ DCNR+ CFNC G GH++ C +
Sbjct: 44 CYQCGEFGHLARDCNRSSNSGGGGGGSGGGCFNCGGFGHLARDCVR 181
Score = 40.8 bits (94), Expect = 0.002
Identities = 23/80 (28%), Positives = 28/80 (34%), Gaps = 22/80 (27%)
Frame = +2
Query: 434 CFNCGEKGHKSNACPEEIKK----------CVRCGKKGHVVADCNRTDI----------- 472
C+ CGE GH + C C CG GH+ DC R
Sbjct: 44 CYQCGEFGHLARDCNRSSNSGGGGGGSGGGCFNCGGFGHLARDCVRGGGGSVGIGGGGGG 223
Query: 473 -VCFNCNGEGHISSQCTQPK 491
CF C G GH++ C K
Sbjct: 224 GSCFRCGGFGHMARDCATGK 283
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.322 0.138 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 33,441,372
Number of Sequences: 63676
Number of extensions: 445013
Number of successful extensions: 2251
Number of sequences better than 10.0: 71
Number of HSP's better than 10.0 without gapping: 1875
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2186
length of query: 805
length of database: 12,639,632
effective HSP length: 105
effective length of query: 700
effective length of database: 5,953,652
effective search space: 4167556400
effective search space used: 4167556400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 63 (28.9 bits)
Medicago: description of AC147714.1


