

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147712.6 + phase: 0
(80 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
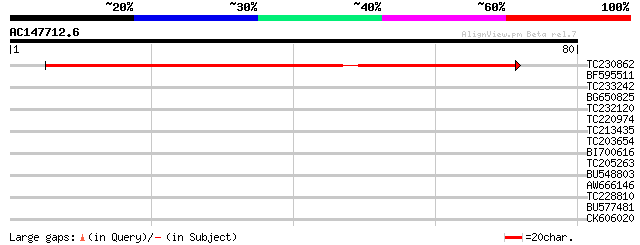
Score E
Sequences producing significant alignments: (bits) Value
TC230862 86 3e-18
BF595511 29 0.38
TC233242 similar to UP|Q9FHK5 (Q9FHK5) Gb|AAF19561.1, partial (11%) 28 1.1
BG650825 similar to PIR|T49915|T49 pre-mRNA splicing factor ATP-... 27 1.9
TC232120 weakly similar to GB|AAQ56803.1|34098841|BT010360 At2g4... 27 1.9
TC220974 similar to UP|MP62_LYTPI (P91753) Mitotic apparatus pro... 27 2.4
TC213435 similar to UP|Q9FKM3 (Q9FKM3) Similarity to AAA-type AT... 27 2.4
TC203654 similar to PIR|T46225|T46225 alpha NAC-like protein - A... 26 3.2
BI700616 26 3.2
TC205263 similar to GB|AAO39931.1|28372898|BT003703 At4g28730 {A... 26 4.2
BU548803 26 4.2
AW666146 25 5.4
TC228810 weakly similar to UP|Q9LIQ9 (Q9LIQ9) Arabidopsis thalia... 25 7.1
BU577481 25 9.3
CK606020 25 9.3
>TC230862
Length = 868
Score = 85.9 bits (211), Expect = 3e-18
Identities = 44/67 (65%), Positives = 55/67 (81%)
Frame = +1
Query: 6 ASEKDLDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEIRVCKNMIKCTVF 65
A E +L+S+RS +KKLEEEL +N +I EL SETGET +Q+L E+EIR+CKNMIKCTV
Sbjct: 427 AIEMELESERSLRKKLEEELRELNCKIDELTSETGETTIQKL--EKEIRICKNMIKCTVC 600
Query: 66 SDRPKEV 72
+DRPKEV
Sbjct: 601 TDRPKEV 621
>BF595511
Length = 420
Score = 29.3 bits (64), Expect = 0.38
Identities = 16/34 (47%), Positives = 24/34 (70%), Gaps = 1/34 (2%)
Frame = +1
Query: 40 GETVVQQLEEE-EEIRVCKNMIKCTVFSDRPKEV 72
G +V ++L+EE EE R ++IKC++ DR KEV
Sbjct: 34 GSSVTEKLQEELEEYR---DIIKCSICQDRAKEV 126
>TC233242 similar to UP|Q9FHK5 (Q9FHK5) Gb|AAF19561.1, partial (11%)
Length = 738
Score = 27.7 bits (60), Expect = 1.1
Identities = 22/74 (29%), Positives = 31/74 (41%)
Frame = +3
Query: 7 SEKDLDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEIRVCKNMIKCTVFS 66
+ K ++ +S K+LEEE+ I E E E + Q E R NM++ V
Sbjct: 291 AHKHVEELKSKAKQLEEEVERQRTVILEGAEEKREAIRQLCFSLEHYRDGYNMLRQHVMG 470
Query: 67 DRPKEVKTGQLIQL 80
R V LI L
Sbjct: 471 HRRVPVLAA*LINL 512
>BG650825 similar to PIR|T49915|T49 pre-mRNA splicing factor ATP-dependent
RNA helicase-like protein - Arabidopsis thaliana,
partial (4%)
Length = 392
Score = 26.9 bits (58), Expect = 1.9
Identities = 17/54 (31%), Positives = 30/54 (55%), Gaps = 8/54 (14%)
Frame = +1
Query: 7 SEKDLDS------KRSSQKK--LEEELMNVNNQIAELNSETGETVVQQLEEEEE 52
S KD D+ KR Q+K +EEE+ N+NN AE+ E + +++ + ++
Sbjct: 37 SVKDSDTSLLEHKKRQKQEKTAMEEEMENLNNVQAEVEKERKQKEKEKMAKHQQ 198
>TC232120 weakly similar to GB|AAQ56803.1|34098841|BT010360 At2g44950
{Arabidopsis thaliana;} , partial (7%)
Length = 715
Score = 26.9 bits (58), Expect = 1.9
Identities = 15/32 (46%), Positives = 19/32 (58%)
Frame = +2
Query: 41 ETVVQQLEEEEEIRVCKNMIKCTVFSDRPKEV 72
E + Q+LEE EI IKC++ DR KEV
Sbjct: 62 EKLQQELEEYREI------IKCSICQDRAKEV 139
>TC220974 similar to UP|MP62_LYTPI (P91753) Mitotic apparatus protein p62,
partial (7%)
Length = 436
Score = 26.6 bits (57), Expect = 2.4
Identities = 17/48 (35%), Positives = 22/48 (45%)
Frame = +1
Query: 5 SASEKDLDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEE 52
S E+ + SSQK EEE V++ E E E EEEE+
Sbjct: 127 SEEEEPQQQQPSSQKNKEEEDEEVSSGEEEEEDEEEEEAASSEEEEED 270
>TC213435 similar to UP|Q9FKM3 (Q9FKM3) Similarity to AAA-type ATPase,
partial (19%)
Length = 524
Score = 26.6 bits (57), Expect = 2.4
Identities = 19/53 (35%), Positives = 30/53 (55%), Gaps = 1/53 (1%)
Frame = +1
Query: 1 MSVASASEKDLDSKRSSQKKLEEELMNVNNQIAELNSETGE-TVVQQLEEEEE 52
M+ A SE + ++R +K +EE L + + AE+N + G V +EEEEE
Sbjct: 166 MTPADISEVLIKNRRKIEKAVEELLETLKLR-AEMNEKNGVLRVTNGVEEEEE 321
>TC203654 similar to PIR|T46225|T46225 alpha NAC-like protein - Arabidopsis
thaliana {Arabidopsis thaliana;} , partial (76%)
Length = 1139
Score = 26.2 bits (56), Expect = 3.2
Identities = 13/48 (27%), Positives = 25/48 (52%)
Frame = +3
Query: 6 ASEKDLDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEI 53
A +DL S+ +Q + + +V + +A+ + + Q EEEEE+
Sbjct: 486 AKIEDLSSQLQTQAAQQFRMPDVGSVLAKQDQDAAAAAAQPEEEEEEV 629
>BI700616
Length = 421
Score = 26.2 bits (56), Expect = 3.2
Identities = 13/40 (32%), Positives = 25/40 (62%)
Frame = +1
Query: 11 LDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEE 50
+D RSS KLE +L+++ I S+T ++++ ++EE
Sbjct: 37 IDKTRSSVDKLESDLISLRQCI----SDTTSSILEMIDEE 144
>TC205263 similar to GB|AAO39931.1|28372898|BT003703 At4g28730 {Arabidopsis
thaliana;} , partial (64%)
Length = 966
Score = 25.8 bits (55), Expect = 4.2
Identities = 17/41 (41%), Positives = 24/41 (58%), Gaps = 1/41 (2%)
Frame = -2
Query: 13 SKRSSQKKLEEELMNVNNQIAE-LNSETGETVVQQLEEEEE 52
S R K+ EEE N ++ E L E G+++V Q E+EEE
Sbjct: 224 SSRRDPKEDEEEA-RTNTEVGETLMREMGKSLVIQQEDEEE 105
>BU548803
Length = 715
Score = 25.8 bits (55), Expect = 4.2
Identities = 18/55 (32%), Positives = 26/55 (46%), Gaps = 3/55 (5%)
Frame = -2
Query: 5 SASEKD---LDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEIRVC 56
SAS+ D D++ ++ EEE + N+ E E GE + EEEE C
Sbjct: 417 SASDSDDEVCDAEEGVDEQQEEENESEKNRGEEEEKECGEEDIATGAEEEEEEEC 253
>AW666146
Length = 428
Score = 25.4 bits (54), Expect = 5.4
Identities = 16/57 (28%), Positives = 29/57 (50%), Gaps = 2/57 (3%)
Frame = +3
Query: 1 MSVASA--SEKDLDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEIRV 55
+S+ S+ SEK+ S R+S +K N+N+Q + S + Q + E +R+
Sbjct: 27 LSIGSSDMSEKNDQSNRNSSEKSSNNSNNINDQNNKQQSNNMALLRVQEQAREHLRI 197
>TC228810 weakly similar to UP|Q9LIQ9 (Q9LIQ9) Arabidopsis thaliana genomic
DNA, chromosome 3, BAC clone:F14O13, partial (30%)
Length = 1277
Score = 25.0 bits (53), Expect = 7.1
Identities = 13/52 (25%), Positives = 28/52 (53%), Gaps = 9/52 (17%)
Frame = +3
Query: 15 RSSQKKLEEELMNVNNQIAELNSETG---------ETVVQQLEEEEEIRVCK 57
RSS+ KLE++L N +I+ + ++ ++ L+EE+++ + K
Sbjct: 30 RSSELKLEKQLENAKEEISSYRKKMSSLDKDRHDLQSTIEALQEEKKMLLSK 185
>BU577481
Length = 409
Score = 24.6 bits (52), Expect = 9.3
Identities = 14/39 (35%), Positives = 23/39 (58%), Gaps = 4/39 (10%)
Frame = -2
Query: 29 NNQIAELN----SETGETVVQQLEEEEEIRVCKNMIKCT 63
N +A+L S+T + + ++E + I+VCKNM K T
Sbjct: 357 NVDVADLKF*CLSDTSKWLA*KIELNKIIKVCKNMWKST 241
>CK606020
Length = 340
Score = 24.6 bits (52), Expect = 9.3
Identities = 17/53 (32%), Positives = 26/53 (48%), Gaps = 2/53 (3%)
Frame = +1
Query: 3 VASASEKD--LDSKRSSQKKLEEELMNVNNQIAELNSETGETVVQQLEEEEEI 53
VA+ E D LD + +++ EEE + E E E V++ EEEE+
Sbjct: 163 VANDDEIDPSLDEEEEEEEEEEEEEEEEEEEEIEEEEEEEEEEVEEEVEEEEV 321
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.304 0.122 0.303
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,097,397
Number of Sequences: 63676
Number of extensions: 14921
Number of successful extensions: 82
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 79
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 81
length of query: 80
length of database: 12,639,632
effective HSP length: 56
effective length of query: 24
effective length of database: 9,073,776
effective search space: 217770624
effective search space used: 217770624
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 43 (21.9 bits)
S2: 52 (24.6 bits)
Medicago: description of AC147712.6


