

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147712.16 + phase: 0
(236 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
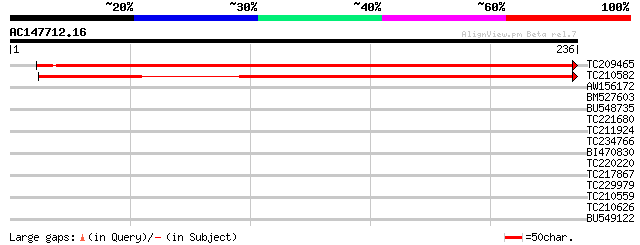
Score E
Sequences producing significant alignments: (bits) Value
TC209465 similar to UP|Q9LTU6 (Q9LTU6) Gb|AAF50915.1, partial (97%) 385 e-108
TC210582 similar to UP|Q9LTU6 (Q9LTU6) Gb|AAF50915.1, partial (64%) 289 9e-79
AW156172 30 0.73
BM527603 similar to GP|10946337|gb| CONSTANS-like protein {Ipomo... 30 0.96
BU548735 28 2.8
TC221680 similar to GB|AAO64901.1|29029044|BT005966 At4g37460 {A... 28 2.8
TC211924 similar to UP|Q9V3T8 (Q9V3T8) SR family splicing factor... 28 2.8
TC234766 homologue to UP|Q7Y137 (Q7Y137) MADS-box protein PTM5, ... 28 2.8
BI470830 28 3.6
TC220220 similar to UP|P74283 (P74283) Hydrogenase component, pa... 28 3.6
TC217867 similar to UP|Q9ASX3 (Q9ASX3) AT5g46190/MCL19_25 (Fragm... 28 4.7
TC229979 similar to UP|Q9FIN8 (Q9FIN8) Arabidopsis thaliana geno... 28 4.7
TC210559 similar to UP|Q84MI1 (Q84MI1) MADS-box protein (Fragmen... 27 8.1
TC210626 weakly similar to UP|Q7AKE8 (Q7AKE8) Thioesterase II, p... 27 8.1
BU549122 27 8.1
>TC209465 similar to UP|Q9LTU6 (Q9LTU6) Gb|AAF50915.1, partial (97%)
Length = 1000
Score = 385 bits (989), Expect = e-108
Identities = 191/225 (84%), Positives = 207/225 (91%)
Frame = +1
Query: 12 FSCSLNSCTMASSSMEVDNVLSEQKVLPPKPKFEPLKPHEMPGAAVQFRKVSVPPHRYTP 71
F+C + S +SSMEV+ V SE K+LPPKPKFEPLKPHEM VQFRKV+VPPHRYTP
Sbjct: 100 FTCRMES-NQEASSMEVETVPSEAKLLPPKPKFEPLKPHEMSDGQVQFRKVNVPPHRYTP 276
Query: 72 LKKVWMDIYTPVFEQMKIDIRMNLKGRKVELKTRHDTPDISNLQKCADFVHAFMLGFDVI 131
LKK WMDIY+P++EQMKID+RMNLK R+VELKTR DTPDISNLQKCADFVHAFMLGFDVI
Sbjct: 277 LKKAWMDIYSPIYEQMKIDVRMNLKARRVELKTRPDTPDISNLQKCADFVHAFMLGFDVI 456
Query: 132 DAIAILRLDELYVESFEIKDVKTLRGDHLSRAIGRLSGKGGKTKFTIENASKTRIVIADT 191
DAIA+LRLDELY+ESFEIKDVKTLRGDHLSRAIGRLSGKGGKTKF IENASKTRIVIADT
Sbjct: 457 DAIALLRLDELYIESFEIKDVKTLRGDHLSRAIGRLSGKGGKTKFAIENASKTRIVIADT 636
Query: 192 KIHILGSFANIKIARDSLCYLIMGSPAGKVYSKLRAVTSRMAERF 236
KIHILGSFANIKIARDSLC LI+GSPAGKVYSKLRAVT+R+AERF
Sbjct: 637 KIHILGSFANIKIARDSLCSLILGSPAGKVYSKLRAVTARLAERF 771
>TC210582 similar to UP|Q9LTU6 (Q9LTU6) Gb|AAF50915.1, partial (64%)
Length = 798
Score = 289 bits (739), Expect = 9e-79
Identities = 155/224 (69%), Positives = 167/224 (74%)
Frame = +2
Query: 13 SCSLNSCTMASSSMEVDNVLSEQKVLPPKPKFEPLKPHEMPGAAVQFRKVSVPPHRYTPL 72
+C A+SSMEV+ V SE K+LPPKPKFEPLKPHEM
Sbjct: 65 TCCRMESNQAASSMEVETVPSEAKLLPPKPKFEPLKPHEMSDG----------------- 193
Query: 73 KKVWMDIYTPVFEQMKIDIRMNLKGRKVELKTRHDTPDISNLQKCADFVHAFMLGFDVID 132
+ R+VELKTR DTPDISNLQKCADFVHAFMLGFDVID
Sbjct: 194 -----------------------QARRVELKTRPDTPDISNLQKCADFVHAFMLGFDVID 304
Query: 133 AIAILRLDELYVESFEIKDVKTLRGDHLSRAIGRLSGKGGKTKFTIENASKTRIVIADTK 192
AIA+LRLDELY+ESFEIKDVKTLRGDHLSRAIGRLSGKGGKTKF IENASKTRIVIADTK
Sbjct: 305 AIALLRLDELYIESFEIKDVKTLRGDHLSRAIGRLSGKGGKTKFAIENASKTRIVIADTK 484
Query: 193 IHILGSFANIKIARDSLCYLIMGSPAGKVYSKLRAVTSRMAERF 236
IHILGSFANIKIARDS+C LI+GSPAGKVYSKLRAVT+R+AERF
Sbjct: 485 IHILGSFANIKIARDSICSLILGSPAGKVYSKLRAVTARLAERF 616
>AW156172
Length = 306
Score = 30.4 bits (67), Expect = 0.73
Identities = 16/46 (34%), Positives = 23/46 (49%)
Frame = -3
Query: 38 LPPKPKFEPLKPHEMPGAAVQFRKVSVPPHRYTPLKKVWMDIYTPV 83
LPP F P K + + G V F PP LKKV ++ ++P+
Sbjct: 142 LPPPQGFFPPKKNFLGGGGVGFSPWGPPPFFLKKLKKVGINFFSPL 5
>BM527603 similar to GP|10946337|gb| CONSTANS-like protein {Ipomoea nil},
partial (9%)
Length = 427
Score = 30.0 bits (66), Expect = 0.96
Identities = 21/48 (43%), Positives = 23/48 (47%)
Frame = +1
Query: 28 VDNVLSEQKVLPPKPKFEPLKPHEMPGAAVQFRKVSVPPHRYTPLKKV 75
V N L E VLPP PKF PL P V K +PP PL K+
Sbjct: 292 VTNGLLELSVLPPLPKFPPLHP-------VPIYKKPLPP--LPPLPKL 408
>BU548735
Length = 659
Score = 28.5 bits (62), Expect = 2.8
Identities = 14/40 (35%), Positives = 21/40 (52%)
Frame = -3
Query: 4 LLCFFNQLFSCSLNSCTMASSSMEVDNVLSEQKVLPPKPK 43
L FFN+ +SCS +C++ SS VL + + K K
Sbjct: 375 LKTFFNKDYSCSKEACSVESSKRSATEVLGTTEDISIKEK 256
>TC221680 similar to GB|AAO64901.1|29029044|BT005966 At4g37460 {Arabidopsis
thaliana;} , partial (36%)
Length = 1001
Score = 28.5 bits (62), Expect = 2.8
Identities = 11/29 (37%), Positives = 18/29 (61%)
Frame = -3
Query: 75 VWMDIYTPVFEQMKIDIRMNLKGRKVELK 103
+WM PVF + I+I +N+K K+ +K
Sbjct: 759 LWMQFLAPVFLKKSINIPVNIKPAKITIK 673
>TC211924 similar to UP|Q9V3T8 (Q9V3T8) SR family splicing factor SC35
(CG5442-PB) (LD32469p), partial (9%)
Length = 510
Score = 28.5 bits (62), Expect = 2.8
Identities = 16/62 (25%), Positives = 29/62 (45%)
Frame = +1
Query: 5 LCFFNQLFSCSLNSCTMASSSMEVDNVLSEQKVLPPKPKFEPLKPHEMPGAAVQFRKVSV 64
L N++FS NS + ASSS+ + ++ ++ PP P P ++ ++
Sbjct: 196 LLSLNRIFSPPTNSSSTASSSLSISSLTNQTLPKPPTL*LRSKNPTRNPNPNIRRA*PNL 375
Query: 65 PP 66
PP
Sbjct: 376 PP 381
>TC234766 homologue to UP|Q7Y137 (Q7Y137) MADS-box protein PTM5, partial
(33%)
Length = 490
Score = 28.5 bits (62), Expect = 2.8
Identities = 18/66 (27%), Positives = 32/66 (48%), Gaps = 1/66 (1%)
Frame = +2
Query: 172 GKTKFT-IENASKTRIVIADTKIHILGSFANIKIARDSLCYLIMGSPAGKVYSKLRAVTS 230
GKT+ IENA+ ++ + + +L + + D+ LI+ SP GK+Y +
Sbjct: 248 GKTQLRRIENATSRQVTFSKRRNGLLKKAFELSVLCDAEVALIIFSPRGKLYEFASSSMQ 427
Query: 231 RMAERF 236
ER+
Sbjct: 428 DTIERY 445
>BI470830
Length = 428
Score = 28.1 bits (61), Expect = 3.6
Identities = 12/33 (36%), Positives = 18/33 (54%)
Frame = -2
Query: 1 MLHLLCFFNQLFSCSLNSCTMASSSMEVDNVLS 33
+LH+LCF N SC+ NS + + N +S
Sbjct: 307 ILHVLCFVNSNLSCAKNSSFNRGLILAISNSIS 209
>TC220220 similar to UP|P74283 (P74283) Hydrogenase component, partial (19%)
Length = 751
Score = 28.1 bits (61), Expect = 3.6
Identities = 13/35 (37%), Positives = 16/35 (45%)
Frame = +2
Query: 40 PKPKFEPLKPHEMPGAAVQFRKVSVPPHRYTPLKK 74
P P ++P PG R S PPHR L+K
Sbjct: 152 PSPPTPGIRPSIFPGGGSPTRSFSPPPHRRNLLQK 256
>TC217867 similar to UP|Q9ASX3 (Q9ASX3) AT5g46190/MCL19_25 (Fragment),
partial (35%)
Length = 1429
Score = 27.7 bits (60), Expect = 4.7
Identities = 14/32 (43%), Positives = 19/32 (58%)
Frame = +3
Query: 161 SRAIGRLSGKGGKTKFTIENASKTRIVIADTK 192
S IGR+ GKGG T ++ AS I + D+K
Sbjct: 18 SDKIGRVIGKGGSTIKSMRQASGAHIEVDDSK 113
>TC229979 similar to UP|Q9FIN8 (Q9FIN8) Arabidopsis thaliana genomic DNA,
chromosome 5, TAC clone:K16H17 (At5g24340), partial
(18%)
Length = 535
Score = 27.7 bits (60), Expect = 4.7
Identities = 13/23 (56%), Positives = 15/23 (64%), Gaps = 2/23 (8%)
Frame = -1
Query: 2 LHLLCFFNQLFSCS--LNSCTMA 22
L LC FN LFSC LN C++A
Sbjct: 151 LKALCSFNSLFSCQWLLNKCSIA 83
>TC210559 similar to UP|Q84MI1 (Q84MI1) MADS-box protein (Fragment), partial
(64%)
Length = 710
Score = 26.9 bits (58), Expect = 8.1
Identities = 15/59 (25%), Positives = 28/59 (47%)
Frame = +1
Query: 178 IENASKTRIVIADTKIHILGSFANIKIARDSLCYLIMGSPAGKVYSKLRAVTSRMAERF 236
IENA+ ++ + + +L + + D+ LI+ SP GK+Y + ER+
Sbjct: 370 IENATSRQVTFSKRRNGLLKKAFELSVLCDAEVALIIFSPRGKLYEFASSSMQDTIERY 546
>TC210626 weakly similar to UP|Q7AKE8 (Q7AKE8) Thioesterase II, partial (7%)
Length = 773
Score = 26.9 bits (58), Expect = 8.1
Identities = 12/26 (46%), Positives = 16/26 (61%)
Frame = +1
Query: 1 MLHLLCFFNQLFSCSLNSCTMASSSM 26
+LHL CF + L+ CS +C S SM
Sbjct: 589 LLHLFCFNSLLWICSKYNCLHRSLSM 666
>BU549122
Length = 670
Score = 26.9 bits (58), Expect = 8.1
Identities = 11/27 (40%), Positives = 16/27 (58%)
Frame = +1
Query: 39 PPKPKFEPLKPHEMPGAAVQFRKVSVP 65
PP PK P PH P + ++F ++VP
Sbjct: 478 PPLPKIHP--PHSPPHSQIRFASLAVP 552
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.324 0.138 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,144,875
Number of Sequences: 63676
Number of extensions: 138819
Number of successful extensions: 877
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 868
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 877
length of query: 236
length of database: 12,639,632
effective HSP length: 94
effective length of query: 142
effective length of database: 6,654,088
effective search space: 944880496
effective search space used: 944880496
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 57 (26.6 bits)
Medicago: description of AC147712.16


