

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147538.3 - phase: 0
(123 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
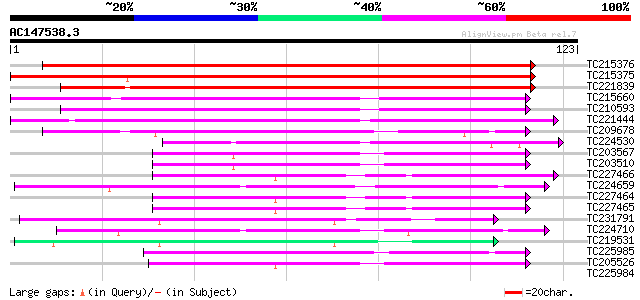
Score E
Sequences producing significant alignments: (bits) Value
TC215376 similar to UP|UGP5_ARATH (Q9C7F7) Uncharacterized GPI-a... 155 6e-39
TC215375 weakly similar to UP|UGP5_ARATH (Q9C7F7) Uncharacterize... 153 2e-38
TC221839 weakly similar to UP|Q6YZD3 (Q6YZD3) Lipid transfer pro... 137 2e-33
TC215660 similar to UP|Q9LJ86 (Q9LJ86) Gb|AAD48513.1 (AT3g22600/... 65 1e-11
TC210593 weakly similar to UP|Q9LJ86 (Q9LJ86) Gb|AAD48513.1 (AT3... 60 2e-10
TC221444 weakly similar to UP|Q8H574 (Q8H574) Protease inhibitor... 60 3e-10
TC209678 weakly similar to GB|AAP12845.1|30017223|BT006196 At2g4... 51 2e-07
TC224530 50 3e-07
TC203567 UP|Q43437 (Q43437) Photosystem II type I chlorophyll a/... 44 2e-05
TC203510 weakly similar to UP|Q9ZVC7 (Q9ZVC7) Expressed protein,... 44 2e-05
TC227466 similar to UP|Q9FMI9 (Q9FMI9) Similarity to nonspecific... 42 5e-05
TC224659 weakly similar to UP|Q6NLF7 (Q6NLF7) At1g62790, partial... 41 1e-04
TC227464 similar to UP|Q8VYI9 (Q8VYI9) AT5g64080/MHJ24_6, partia... 41 1e-04
TC227465 similar to UP|Q6I6X1 (Q6I6X1) Xylogen protein 1, partia... 41 2e-04
TC231791 similar to UP|O22484 (O22484) Lipid transfer protein LP... 40 2e-04
TC224710 similar to UP|Q8ZZP6 (Q8ZZP6) Conserved protein, partia... 40 2e-04
TC219531 40 2e-04
TC225985 weakly similar to UP|Q6Z387 (Q6Z387) Lipid transfer pro... 39 5e-04
TC205526 weakly similar to UP|Q8VYI9 (Q8VYI9) AT5g64080/MHJ24_6,... 39 8e-04
TC225984 weakly similar to UP|Q6Z387 (Q6Z387) Lipid transfer pro... 36 0.004
>TC215376 similar to UP|UGP5_ARATH (Q9C7F7) Uncharacterized GPI-anchored
protein At1g27950 precursor, partial (24%)
Length = 1023
Score = 155 bits (391), Expect = 6e-39
Identities = 70/107 (65%), Positives = 89/107 (82%)
Frame = +2
Query: 8 MFMCLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKAT 67
M M L V+ + G +GA+DLA+KC +V+QKVIPCLDFA GK TPKK+CCDAA SIK +
Sbjct: 113 MGMGLLVVMMACGSASGADDLATKCSAVIQKVIPCLDFAKGKEETPKKQCCDAATSIKES 292
Query: 68 DPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANISDCP 114
+PECLCYII++THKGSP+ KS+GIQE KLLQLP+VC+V A+I++CP
Sbjct: 293 NPECLCYIIEETHKGSPQVKSLGIQEAKLLQLPSVCNVKNASITNCP 433
>TC215375 weakly similar to UP|UGP5_ARATH (Q9C7F7) Uncharacterized
GPI-anchored protein At1g27950 precursor, partial (45%)
Length = 879
Score = 153 bits (387), Expect = 2e-38
Identities = 72/118 (61%), Positives = 94/118 (79%), Gaps = 4/118 (3%)
Frame = +1
Query: 1 MKNQQHQMFMCLCVLALIIGGCNG----AEDLASKCGSVVQKVIPCLDFATGKAPTPKKE 56
+K + FM L +L +++ GC G A+DLA+KC +V+QKVIPCL+FATGK PKKE
Sbjct: 7 IKKMKRVEFMGLGLLVVVMMGCCGSATAADDLATKCSAVIQKVIPCLNFATGKEEMPKKE 186
Query: 57 CCDAANSIKATDPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANISDCP 114
CCDAA +IK ++PECLCYIIQ+THKGSP+ KS+GIQE KLLQLP+VC+V A+I++CP
Sbjct: 187 CCDAATAIKESNPECLCYIIQETHKGSPQVKSLGIQEAKLLQLPSVCNVKNASITNCP 360
>TC221839 weakly similar to UP|Q6YZD3 (Q6YZD3) Lipid transfer protein-like,
partial (37%)
Length = 476
Score = 137 bits (344), Expect = 2e-33
Identities = 61/103 (59%), Positives = 81/103 (78%)
Frame = +2
Query: 12 LCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKATDPEC 71
LC+ L +G GA DLA+KC +VQKV+PCL+FATG+A P K+CC+A + IK +DPEC
Sbjct: 29 LCLFLLAVGESEGA-DLAAKCNLLVQKVLPCLNFATGQAAVPTKDCCEATSEIKKSDPEC 205
Query: 72 LCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANISDCP 114
LC+ IQQTHKGSPE K+MGIQE +LLQLP+ C++ A+ ++CP
Sbjct: 206 LCFAIQQTHKGSPEVKNMGIQEARLLQLPSACNLKNASTTNCP 334
>TC215660 similar to UP|Q9LJ86 (Q9LJ86) Gb|AAD48513.1 (AT3g22600/F16J14_17),
partial (48%)
Length = 797
Score = 64.7 bits (156), Expect = 1e-11
Identities = 34/113 (30%), Positives = 54/113 (47%)
Frame = +3
Query: 1 MKNQQHQMFMCLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDA 60
M + + M + L V+A++ G A S C +V+ + PCL++ TG + TP CC
Sbjct: 75 MAHTKVDMGLVLVVMAMLCAGA--AAQSQSSCTNVLVSLSPCLNYITGNSSTPSSGCCSQ 248
Query: 61 ANSIKATDPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANISDC 113
S+ + P+CLC ++ G S + I + + L LP C V S C
Sbjct: 249 LASVVRSQPQCLCQVL----SGGGSSLGININQTQALALPVACKVQTPPTSQC 395
>TC210593 weakly similar to UP|Q9LJ86 (Q9LJ86) Gb|AAD48513.1
(AT3g22600/F16J14_17), partial (56%)
Length = 728
Score = 60.5 bits (145), Expect = 2e-10
Identities = 29/102 (28%), Positives = 48/102 (46%)
Frame = +2
Query: 12 LCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKATDPEC 71
LC++A+I+ S C + + + PCL++ G + TP CC +SI + P+C
Sbjct: 125 LCLVAVIVATMWSQNAAQSGCTNTLTSLSPCLNYIMGSSSTPSASCCSQLSSIVQSSPQC 304
Query: 72 LCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANISDC 113
LC ++ G + + I + L LP C V +S C
Sbjct: 305 LCSVL----NGGGSTFGITINQTLALSLPGACEVQTPPVSQC 418
>TC221444 weakly similar to UP|Q8H574 (Q8H574) Protease inhibitor-like
protein, partial (32%)
Length = 657
Score = 60.1 bits (144), Expect = 3e-10
Identities = 33/119 (27%), Positives = 57/119 (47%)
Frame = +1
Query: 1 MKNQQHQMFMCLCVLALIIGGCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDA 60
M +++ +M + + L + + G A+ S C +V + PCLD+ T A P CC
Sbjct: 79 MASRRIEMLLSMS-LVMALWGVTLAQSDQSSCTNVFISLSPCLDYVTENASIPSSSCCSQ 255
Query: 61 ANSIKATDPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANISDCPSKSKT 119
+ + P CLC ++ S + S I + + L LPT C+V +IS C + + +
Sbjct: 256 LAFVVRSQPLCLCEVV--NGGASSIAASFNINQTRALALPTSCNVQTPSISSCSASASS 426
>TC209678 weakly similar to GB|AAP12845.1|30017223|BT006196 At2g44300
{Arabidopsis thaliana;} , partial (49%)
Length = 764
Score = 50.8 bits (120), Expect = 2e-07
Identities = 32/111 (28%), Positives = 56/111 (49%), Gaps = 5/111 (4%)
Frame = +2
Query: 8 MFMCLCVLALIIGGCNGAEDLAS---KCGSVVQKVIPCLDFATGKAPTPKKECCDAANSI 64
M M L V+ L++G DL+ +C + + C+ + G+A TP +CC ++
Sbjct: 74 MLMLLVVVLLVVGFAKS--DLSKDREECADKLIDLASCVPYVGGEAKTPTIDCCTGLKAV 247
Query: 65 KATDPECLCYIIQQTHKGSPESKSMGIQEDKLL--QLPTVCHVNGANISDC 113
+CLC +I+ + ++GI+ + L QLP+ CH + ANI+ C
Sbjct: 248 LDRSKKCLCILIKDR-----DDPNLGIKINATLAIQLPSACH-SPANITQC 382
>TC224530
Length = 556
Score = 50.1 bits (118), Expect = 3e-07
Identities = 27/90 (30%), Positives = 51/90 (56%), Gaps = 3/90 (3%)
Frame = +3
Query: 34 SVVQKVIPCLDFATGKAPTPKKECCDAANSIKATDPECLCYIIQQTHKGSPESKSMGIQE 93
S + K+ PCL++ G P CC+ S+ +D ECLC ++ +++G+ +++ GI
Sbjct: 114 SCLNKLSPCLNYLNG-TEDPPDSCCEPLKSVIESDAECLCSLV--SNRGTRQAEQAGINI 284
Query: 94 DKLLQLPTVC--HVNGAN-ISDCPSKSKTD 120
++ QLP C HVN + +++ P + +D
Sbjct: 285 NEAQQLPGRCGQHVNPLSCLTNSPGPTNSD 374
>TC203567 UP|Q43437 (Q43437) Photosystem II type I chlorophyll a/b-binding
protein precursor, partial (52%)
Length = 1448
Score = 43.5 bits (101), Expect = 2e-05
Identities = 25/84 (29%), Positives = 39/84 (45%), Gaps = 2/84 (2%)
Frame = -2
Query: 32 CGSVVQKVIPCLDFAT--GKAPTPKKECCDAANSIKATDPECLCYIIQQTHKGSPESKSM 89
C + + CL + K P K CC + ++P CLC ++ G P+S +
Sbjct: 766 CLMALANMSDCLTYVEDGSKLSKPDKGCCPELAGLVDSNPICLCEML-----GKPDSIGI 602
Query: 90 GIQEDKLLQLPTVCHVNGANISDC 113
I +K L+LP+VC V +S C
Sbjct: 601 KIDLNKALKLPSVCGVTTPPVSTC 530
>TC203510 weakly similar to UP|Q9ZVC7 (Q9ZVC7) Expressed protein, partial
(24%)
Length = 975
Score = 43.5 bits (101), Expect = 2e-05
Identities = 25/84 (29%), Positives = 39/84 (45%), Gaps = 2/84 (2%)
Frame = +1
Query: 32 CGSVVQKVIPCLDFAT--GKAPTPKKECCDAANSIKATDPECLCYIIQQTHKGSPESKSM 89
C + + CL + K P K CC + ++P CLC ++ G P+S +
Sbjct: 262 CLMALTNMSDCLTYVEDGSKLAKPDKGCCPELAGLIDSNPICLCELL-----GKPDSIGI 426
Query: 90 GIQEDKLLQLPTVCHVNGANISDC 113
I +K L+LP+VC V +S C
Sbjct: 427 KIDLNKALKLPSVCGVTTPPVSTC 498
>TC227466 similar to UP|Q9FMI9 (Q9FMI9) Similarity to nonspecific
lipid-transfer protein, partial (51%)
Length = 674
Score = 42.4 bits (98), Expect = 5e-05
Identities = 26/90 (28%), Positives = 40/90 (43%), Gaps = 2/90 (2%)
Frame = +3
Query: 32 CGSVVQKVIPCLDFATGKAPTPKKE--CCDAANSIKATDPECLCYIIQQTHKGSPESKSM 89
C ++V + CL F T + K E CC S+ T P CLC + K S + +
Sbjct: 165 CTNLVLTMADCLSFVTNGSTVTKPEGTCCSGLKSVLKTAPACLC----EAFKSSAQF-GV 329
Query: 90 GIQEDKLLQLPTVCHVNGANISDCPSKSKT 119
+ K LP C V+ + ++C +S T
Sbjct: 330 VLNVTKATSLPAACKVSAPSATNCGCQSIT 419
>TC224659 weakly similar to UP|Q6NLF7 (Q6NLF7) At1g62790, partial (59%)
Length = 706
Score = 41.2 bits (95), Expect = 1e-04
Identities = 32/119 (26%), Positives = 52/119 (42%), Gaps = 3/119 (2%)
Frame = +1
Query: 2 KNQQHQMFMCLCVLALIIG---GCNGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECC 58
K +Q +M C+L L++ + AE + Q++IPCLD+ G P CC
Sbjct: 79 KTKQEKMGGRKCLLTLVLSLVLMMSMAEAQSGTSADCAQELIPCLDYLNGTI-NPPSSCC 255
Query: 59 DAANSIKATDPECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVNGANISDCPSKS 117
D + CLC I + +S+ + D+ L L C V +++S C + S
Sbjct: 256 DPLKRTVQNELACLCNI----YFSPGLLQSVNVTVDEALGLSRRCGVT-SDLSSCKNGS 417
>TC227464 similar to UP|Q8VYI9 (Q8VYI9) AT5g64080/MHJ24_6, partial (53%)
Length = 829
Score = 41.2 bits (95), Expect = 1e-04
Identities = 24/84 (28%), Positives = 37/84 (43%), Gaps = 2/84 (2%)
Frame = +2
Query: 32 CGSVVQKVIPCLDFATGKAPTPKKE--CCDAANSIKATDPECLCYIIQQTHKGSPESKSM 89
C ++V + CL F T + K E CC S+ T P CLC + K S + +
Sbjct: 128 CSNLVLTMADCLSFVTNGSTVTKPEGTCCSGLKSVLKTAPACLC----EAFKSSAQF-GV 292
Query: 90 GIQEDKLLQLPTVCHVNGANISDC 113
+ K LP C V+ + ++C
Sbjct: 293 VLNVTKATTLPAACKVSAPSATNC 364
>TC227465 similar to UP|Q6I6X1 (Q6I6X1) Xylogen protein 1, partial (53%)
Length = 488
Score = 40.8 bits (94), Expect = 2e-04
Identities = 24/84 (28%), Positives = 37/84 (43%), Gaps = 2/84 (2%)
Frame = +2
Query: 32 CGSVVQKVIPCLDFATGKAPTPKKE--CCDAANSIKATDPECLCYIIQQTHKGSPESKSM 89
C ++V + CL F T + K E CC S+ T P CLC + K S + +
Sbjct: 158 CTNLVLTMADCLSFVTNGSTVTKPEGTCCSGLKSVLKTAPACLC----EAFKSSAQF-GV 322
Query: 90 GIQEDKLLQLPTVCHVNGANISDC 113
+ K LP C V+ + ++C
Sbjct: 323 VLNVTKATSLPAACKVSAPSATNC 394
>TC231791 similar to UP|O22484 (O22484) Lipid transfer protein LPT III,
partial (16%)
Length = 463
Score = 40.4 bits (93), Expect = 2e-04
Identities = 28/114 (24%), Positives = 50/114 (43%), Gaps = 10/114 (8%)
Frame = +3
Query: 3 NQQHQMFMCLCVLALIIGGCNGAEDLASK---CGSVVQKVIPCLDFATGKAPTPKKECCD 59
++ Q+F+ L +++ A D A K C + + C+++ TG +P CC+
Sbjct: 54 DKMKQVFVAFLALQVVLVLTIMAADPAGKGYDCEKAKRSLKSCMEYLTGNVDSPSAACCN 233
Query: 60 AANSIKATDP-------ECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVN 106
+KA+ P EC C I++ P K +D+ + LP C V+
Sbjct: 234 GVKELKASAPTKDEKIAECQC--IEEALTPIPNFK-----QDRAIALPKECGVD 374
>TC224710 similar to UP|Q8ZZP6 (Q8ZZP6) Conserved protein, partial (5%)
Length = 787
Score = 40.4 bits (93), Expect = 2e-04
Identities = 31/111 (27%), Positives = 49/111 (43%), Gaps = 4/111 (3%)
Frame = +3
Query: 11 CLCVLALIIGGC---NGAEDLASKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKAT 67
C C+++L++ + AE + + Q++IPC++F G TP CCD
Sbjct: 102 CKCLVSLVLALVLMRSLAEAQSGSSTTCAQELIPCVNFLNGTT-TPPSSCCDPLKQTVEN 278
Query: 68 DPECLCYIIQQTHKGSPE-SKSMGIQEDKLLQLPTVCHVNGANISDCPSKS 117
+CLC I SP +S + D+ L L C V I+ C + S
Sbjct: 279 QLDCLCNIF-----FSPGLLQSFNVSVDQALALSRRCGVTN-GITSCTNGS 413
>TC219531
Length = 576
Score = 40.4 bits (93), Expect = 2e-04
Identities = 30/118 (25%), Positives = 47/118 (39%), Gaps = 13/118 (11%)
Frame = +1
Query: 2 KNQQHQM---FMCLCVLALIIGGCNGAEDLASK---CGSVVQKVIPCLDFATGKAPTPKK 55
+N H+M F+ L +++ A + A K C V PC+ + T K TP
Sbjct: 94 RNYLHKMKKVFVAFLTLQVVLVLTIIAAEPAGKGYNCEKAKSSVTPCIKYLTSKVDTPSA 273
Query: 56 ECCDAANSIKATDP-------ECLCYIIQQTHKGSPESKSMGIQEDKLLQLPTVCHVN 106
CC+ +K++ P C C TH ++ED+ LP C V+
Sbjct: 274 ACCNGVKEVKSSAPTKDEKIAACQCLKEVTTH-------IPNLKEDRATALPKQCGVD 426
>TC225985 weakly similar to UP|Q6Z387 (Q6Z387) Lipid transfer protein-like,
partial (40%)
Length = 1027
Score = 39.3 bits (90), Expect = 5e-04
Identities = 22/84 (26%), Positives = 34/84 (40%)
Frame = +3
Query: 30 SKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKATDPECLCYIIQQTHKGSPESKSM 89
++C + + CL + G+A P +CC + CLC +I+ S +
Sbjct: 165 AECTDKLLGLAGCLSYVGGEAKVPTMDCCSGIKEVINKSKRCLCILIKDR---DDPSLGL 335
Query: 90 GIQEDKLLQLPTVCHVNGANISDC 113
I L LP VC NI+ C
Sbjct: 336 KINVTLALNLPDVCE-TPTNITQC 404
>TC205526 weakly similar to UP|Q8VYI9 (Q8VYI9) AT5g64080/MHJ24_6, partial
(59%)
Length = 942
Score = 38.5 bits (88), Expect = 8e-04
Identities = 20/85 (23%), Positives = 38/85 (44%), Gaps = 2/85 (2%)
Frame = +3
Query: 31 KCGSVVQKVIPCLDFATGKAPTPKKE--CCDAANSIKATDPECLCYIIQQTHKGSPESKS 88
+C ++V + CL F + + K + CC + ++ T P+CLC S
Sbjct: 186 ECSNLVLTLSDCLTFVSNGSTVTKPQGTCCSSLKTVLNTAPKCLCEAF-----NSSAQLG 350
Query: 89 MGIQEDKLLQLPTVCHVNGANISDC 113
+ I K + LP C ++ + ++C
Sbjct: 351 LAINVTKAVTLPAACKLSTPSAANC 425
>TC225984 weakly similar to UP|Q6Z387 (Q6Z387) Lipid transfer protein-like,
partial (28%)
Length = 1140
Score = 36.2 bits (82), Expect = 0.004
Identities = 20/84 (23%), Positives = 33/84 (38%)
Frame = +2
Query: 30 SKCGSVVQKVIPCLDFATGKAPTPKKECCDAANSIKATDPECLCYIIQQTHKGSPESKSM 89
++C + + CL + G+A P +CC + CLC +I+ + +
Sbjct: 233 AECTDKLLGLAGCLPYVGGEAKVPAMDCCSGIREVIDKSKRCLCILIKDR---DDPNLGL 403
Query: 90 GIQEDKLLQLPTVCHVNGANISDC 113
I L LP C NI+ C
Sbjct: 404 KINVTLALSLPDACQ-TPTNITQC 472
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.134 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,695,487
Number of Sequences: 63676
Number of extensions: 92202
Number of successful extensions: 499
Number of sequences better than 10.0: 70
Number of HSP's better than 10.0 without gapping: 493
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 494
length of query: 123
length of database: 12,639,632
effective HSP length: 85
effective length of query: 38
effective length of database: 7,227,172
effective search space: 274632536
effective search space used: 274632536
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 53 (25.0 bits)
Medicago: description of AC147538.3


