

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147517.6 + phase: 0 /pseudo
(1055 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
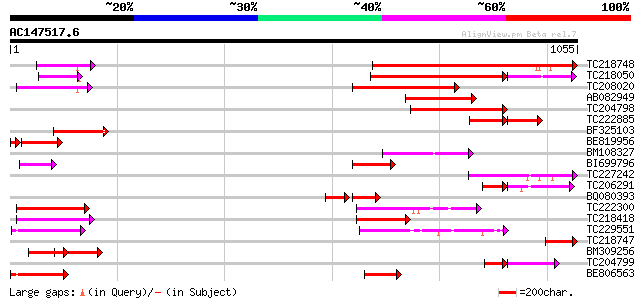
Score E
Sequences producing significant alignments: (bits) Value
TC218748 similar to UP|Q76CU1 (Q76CU1) PDR-type ABC transporter ... 494 e-139
TC218050 weakly similar to UP|Q7PC82 (Q7PC82) PDR14 ABC transpor... 332 7e-98
TC208020 similar to UP|Q76CU2 (Q76CU2) PDR-type ABC transporter ... 321 9e-88
AB082949 224 2e-58
TC204798 similar to UP|Q7PC86 (Q7PC86) PDR7 ABC transporter, par... 218 1e-56
TC222885 weakly similar to UP|Q8GU89 (Q8GU89) PDR-like ABC trans... 111 7e-38
BF325103 150 2e-36
BE819956 similar to GP|20522008|db pleiotropic drug resistance l... 118 3e-31
BM108327 weakly similar to GP|27368815|em PDR-like ABC transport... 121 2e-27
BI699796 similar to PIR|G85167|G85 ABC transporter like protein ... 115 8e-26
TC227242 similar to UP|Q8GU90 (Q8GU90) PDR-like ABC transporter ... 107 3e-23
TC206291 similar to PIR|T02644|T02644 ABC-type transport protein... 67 6e-23
BQ080393 84 2e-22
TC222300 similar to GB|AAP37786.1|30725528|BT008427 At1g51560 {A... 100 4e-21
TC218418 homologue to UP|Q6X4V5 (Q6X4V5) ABC transporter, partia... 98 2e-20
TC229551 weakly similar to UP|Q6X4V5 (Q6X4V5) ABC transporter, p... 97 4e-20
TC218747 similar to UP|Q76CU1 (Q76CU1) PDR-type ABC transporter ... 96 1e-19
BM309256 similar to PIR|T02644|T0 ABC-type transport protein hom... 93 7e-19
TC204799 similar to UP|Q7PC86 (Q7PC86) PDR7 ABC transporter, par... 63 5e-16
BE806563 similar to GP|11994752|db ABC transporter-like protein ... 72 2e-12
>TC218748 similar to UP|Q76CU1 (Q76CU1) PDR-type ABC transporter 2 (Fragment),
partial (37%)
Length = 1384
Score = 494 bits (1271), Expect = e-139
Identities = 271/417 (64%), Positives = 312/417 (73%), Gaps = 36/417 (8%)
Frame = +3
Query: 675 ELVANPSIIFMDEPTSGLDARAAAVVMRTVRNTVDTGRTVVCTIHQPSIDIFESFDELLL 734
ELVANPSIIFMDEPTSGLDARAAA+VMRTVRNTVDTGRTVVCTIHQPSIDIFESFDELLL
Sbjct: 9 ELVANPSIIFMDEPTSGLDARAAAIVMRTVRNTVDTGRTVVCTIHQPSIDIFESFDELLL 188
Query: 735 LKQGGQEIYVGPLGHNSSNLINHFEGIQGVRKIKDGYNPATWMLEVTTSSKERELGIDFA 794
+KQGGQEIYVGPLGH+SS+LIN+FEGIQGV KIKDGYNPATWMLEV+TS+KE ELGIDFA
Sbjct: 189 MKQGGQEIYVGPLGHHSSHLINYFEGIQGVNKIKDGYNPATWMLEVSTSAKEMELGIDFA 368
Query: 795 ELYKNSELYRINKALVKELSAPAPCSKDLYFPSQYSRSFFTQCMACLWKQHWSYWRNPEY 854
E+YKNSELYR NKAL+KELS PAP SKDLYFPSQYS SF TQCMACLWKQHWSYWRNP Y
Sbjct: 369 EVYKNSELYRRNKALIKELSTPAPGSKDLYFPSQYSTSFLTQCMACLWKQHWSYWRNPLY 548
Query: 855 NAIRFLYSTAVAVLLGSMFWDLGSKIEKEQDLFNAMGSMYSAVILIGVMNCNSVQPVVAV 914
AIRFLYSTAVA +LGSMFWDLGSKI+K+QDLFNAMGSMY+AV+LIG+ N N+VQPVVAV
Sbjct: 549 TAIRFLYSTAVAAVLGSMFWDLGSKIDKQQDLFNAMGSMYAAVLLIGIKNANAVQPVVAV 728
Query: 915 ERTVFYRESAAG--NVFYFSICLWAGKYPSPICFCTSCGLWHYSVCD-DW--F*VVYG*S 969
ERTVFYRE AAG + ++ + P + G+ Y++ +W V++
Sbjct: 729 ERTVFYREKAAGMYSALPYAFAQVLIELPYVLVQAVVYGIIIYAMIGFEWTVTKVIWYLF 908
Query: 970 FVVPILLVF--------------HLSLL-----HLLWDDVSG-LDPKQSHF-HYSFL--- 1005
F+ L F H+S + + +W+ SG + P+ F S+L
Sbjct: 909 FMYFTFLTFTYYGMMSVAVTPNQHISSIVSSAFYAVWNLFSGFIVPRPVIFGSLSYLCIE 1088
Query: 1006 -------SILLNMESLLRIHSPKTKNSSVVEVVQLGKSNGMEFVWIGGFTIWRFKKK 1055
IL+ L + T+NSSVVE+VQLGKS MEFVWIGGFTIWR+K K
Sbjct: 1089 ICTFQA*KILIYFILLYFPSNNITENSSVVEMVQLGKSCSMEFVWIGGFTIWRYKAK 1259
Score = 45.4 bits (106), Expect = 1e-04
Identities = 36/118 (30%), Positives = 63/118 (52%), Gaps = 8/118 (6%)
Frame = +3
Query: 50 TGEMLVGPTKALFMDEISTGLDSSTTFQIVNSMRQYVHILKGTVVISLLQPPPETYNLFD 109
T E++ P+ +FMDE ++GLD+ ++ ++R V + TVV ++ QP + + FD
Sbjct: 3 TSELVANPS-IIFMDEPTSGLDARAAAIVMRTVRNTVDTGR-TVVCTIHQPSIDIFESFD 176
Query: 110 DIILLSD-SHIIYQGP----REHVLEFFESIG--FKCPNRKGVADFLQEV-TSRKDQE 159
+++L+ IY GP H++ +FE I K + A ++ EV TS K+ E
Sbjct: 177 ELLLMKQGGQEIYVGPLGHHSSHLINYFEGIQGVNKIKDGYNPATWMLEVSTSAKEME 350
>TC218050 weakly similar to UP|Q7PC82 (Q7PC82) PDR14 ABC transporter, partial
(28%)
Length = 1436
Score = 332 bits (852), Expect(2) = 7e-98
Identities = 160/255 (62%), Positives = 197/255 (76%)
Frame = +2
Query: 672 VAVELVANPSIIFMDEPTSGLDARAAAVVMRTVRNTVDTGRTVVCTIHQPSIDIFESFDE 731
+AVELV+NPSIIFMDEPTSGLDARAAAVVMR V+N V TGRT VCTIHQPSIDIFE+FDE
Sbjct: 2 IAVELVSNPSIIFMDEPTSGLDARAAAVVMRAVKNVVATGRTTVCTIHQPSIDIFETFDE 181
Query: 732 LLLLKQGGQEIYVGPLGHNSSNLINHFEGIQGVRKIKDGYNPATWMLEVTTSSKERELGI 791
L+L+K GG+ IY G LGH+SS LI +F+ I GV KIKD YNPATWMLE T++S E EL I
Sbjct: 182 LILMKSGGRIIYSGMLGHHSSRLIEYFQNIPGVPKIKDNYNPATWMLEATSASVEAELKI 361
Query: 792 DFAELYKNSELYRINKALVKELSAPAPCSKDLYFPSQYSRSFFTQCMACLWKQHWSYWRN 851
DFA++YK S L R LV+ELS P P +KDL+F +++ ++ Q MACLWKQH SYWR+
Sbjct: 362 DFAQIYKESHLCRDTLELVRELSEPPPGTKDLHFSTRFPQNSLGQFMACLWKQHLSYWRS 541
Query: 852 PEYNAIRFLYSTAVAVLLGSMFWDLGSKIEKEQDLFNAMGSMYSAVILIGVMNCNSVQPV 911
PEYN RF++ A++ G++FW G+KI +QDLFN +GSMY AVI +G+ C+++ P
Sbjct: 542 PEYNLTRFIFMIVCAIMFGAVFWQKGNKINNQQDLFNVLGSMYIAVIFLGLNYCSTILPY 721
Query: 912 VAVERTVFYRESAAG 926
VA ER V YRE AG
Sbjct: 722 VATERAVLYREKFAG 766
Score = 45.1 bits (105), Expect(2) = 7e-98
Identities = 33/129 (25%), Positives = 65/129 (49%), Gaps = 1/129 (0%)
Frame = +1
Query: 927 NVFYFSICLWAGKYPSPICFCTSCGLWHYSVCDDWF*VVYG*SFVVPILLVFHLSLLHLL 986
NV + S+ + G + + I + Y + +D F +V S +V + + ++ +L +
Sbjct: 766 NVLFHSLFICTGGH*NTIYLGAINFICGYHISNDRFPLVSSESVLVLLHHILYIFILCIS 945
Query: 987 WDDVSGLDPKQSHFHYSFLSILLN-MESLLRIHSPKTKNSSVVEVVQLGKSNGMEFVWIG 1045
W+D +D +S + + + L+ ++SL I +T+NS +V +V L S + W
Sbjct: 946 WND-GNVDELKSRYSFCIVDCSLHHIQSLFWISHARTENS*MVGLVLLDLSYCLVLEWPP 1122
Query: 1046 GFTIWRFKK 1054
FTIWR ++
Sbjct: 1123 NFTIWRHRE 1149
Score = 43.5 bits (101), Expect = 5e-04
Identities = 27/87 (31%), Positives = 49/87 (56%), Gaps = 5/87 (5%)
Frame = +2
Query: 54 LVGPTKALFMDEISTGLDSSTTFQIVNSMRQYVHILKGTVVISLLQPPPETYNLFDDIIL 113
LV +FMDE ++GLD+ ++ +++ V + T V ++ QP + + FD++IL
Sbjct: 14 LVSNPSIIFMDEPTSGLDARAAAVVMRAVKNVVATGR-TTVCTIHQPSIDIFETFDELIL 190
Query: 114 L-SDSHIIYQGPREH----VLEFFESI 135
+ S IIY G H ++E+F++I
Sbjct: 191 MKSGGRIIYSGMLGHHSSRLIEYFQNI 271
>TC208020 similar to UP|Q76CU2 (Q76CU2) PDR-type ABC transporter 1, partial
(14%)
Length = 765
Score = 321 bits (823), Expect = 9e-88
Identities = 158/199 (79%), Positives = 179/199 (89%)
Frame = +2
Query: 638 QKVMELVELKPLRNALVGLPGVCGLSMEQRKRLTVAVELVANPSIIFMDEPTSGLDARAA 697
++VMELVEL PLRN+LVGLPGV GLS EQRKRLT+AVELVANPSIIFMDEPTSGLDARAA
Sbjct: 167 EEVMELVELNPLRNSLVGLPGVSGLSTEQRKRLTIAVELVANPSIIFMDEPTSGLDARAA 346
Query: 698 AVVMRTVRNTVDTGRTVVCTIHQPSIDIFESFDELLLLKQGGQEIYVGPLGHNSSNLINH 757
A+VMRTVRNTVDTGRTVVCTIHQPSIDIFE+FDEL L+K+GGQEIYVGPLG +S++LI +
Sbjct: 347 AIVMRTVRNTVDTGRTVVCTIHQPSIDIFEAFDELFLMKRGGQEIYVGPLGRHSTHLIKY 526
Query: 758 FEGIQGVRKIKDGYNPATWMLEVTTSSKERELGIDFAELYKNSELYRINKALVKELSAPA 817
FE I GV KIKDGYNPATWMLEVTTS++E LG+DF +LYKNS+LYR NK L++EL PA
Sbjct: 527 FESIGGVSKIKDGYNPATWMLEVTTSAQELSLGVDFTDLYKNSDLYRRNKQLIQELGQPA 706
Query: 818 PCSKDLYFPSQYSRSFFTQ 836
P KDLYFP+QYS+SF Q
Sbjct: 707 PG*KDLYFPTQYSQSFLVQ 763
Score = 61.6 bits (148), Expect = 2e-09
Identities = 44/149 (29%), Positives = 79/149 (52%), Gaps = 7/149 (4%)
Frame = +2
Query: 13 LITDYVLRVLGLEICADTVVGNAMIRAISGGQKKRLTTGEMLVGPTKALFMDEISTGLDS 72
+ + V+ ++ L +++VG + +S Q+KRLT LV +FMDE ++GLD+
Sbjct: 158 MFIEEVMELVELNPLRNSLVGLPGVSGLSTEQRKRLTIAVELVANPSIIFMDEPTSGLDA 337
Query: 73 STTFQIVNSMRQYVHILKGTVVISLLQPPPETYNLFDDIILLS-DSHIIYQGP----REH 127
++ ++R V + TVV ++ QP + + FD++ L+ IY GP H
Sbjct: 338 RAAAIVMRTVRNTVDTGR-TVVCTIHQPSIDIFEAFDELFLMKRGGQEIYVGPLGRHSTH 514
Query: 128 VLEFFESIG--FKCPNRKGVADFLQEVTS 154
++++FESIG K + A ++ EVT+
Sbjct: 515 LIKYFESIGGVSKIKDGYNPATWMLEVTT 601
>AB082949
Length = 403
Score = 224 bits (571), Expect = 2e-58
Identities = 104/133 (78%), Positives = 120/133 (90%)
Frame = +3
Query: 736 KQGGQEIYVGPLGHNSSNLINHFEGIQGVRKIKDGYNPATWMLEVTTSSKERELGIDFAE 795
++GGQEIYVGPLGH+S +LI++FEGI+GVR I+DGYNPATWMLEVTTS+KE ELGIDFAE
Sbjct: 3 QRGGQEIYVGPLGHHSYHLISYFEGIKGVRTIEDGYNPATWMLEVTTSAKEMELGIDFAE 182
Query: 796 LYKNSELYRINKALVKELSAPAPCSKDLYFPSQYSRSFFTQCMACLWKQHWSYWRNPEYN 855
LYKNS+LYR NK L++ELS PAP SKDLYF S+YSRSF TQCMACLWKQHWSYWRN EY
Sbjct: 183 LYKNSDLYRRNKELIEELSTPAPGSKDLYFSSKYSRSFITQCMACLWKQHWSYWRNNEYT 362
Query: 856 AIRFLYSTAVAVL 868
A+RFL++ AVA L
Sbjct: 363 ALRFLFTIAVASL 401
>TC204798 similar to UP|Q7PC86 (Q7PC86) PDR7 ABC transporter, partial (20%)
Length = 1048
Score = 218 bits (554), Expect = 1e-56
Identities = 100/181 (55%), Positives = 134/181 (73%)
Frame = +3
Query: 746 PLGHNSSNLINHFEGIQGVRKIKDGYNPATWMLEVTTSSKERELGIDFAELYKNSELYRI 805
PLG NS + +FE I GV KIK+ YNPATWMLEV++ + E LG+DFAE YK S L++
Sbjct: 18 PLGRNSHKITEYFEAIPGVPKIKEMYNPATWMLEVSSVAAEVRLGMDFAEYYKTSSLFQR 197
Query: 806 NKALVKELSAPAPCSKDLYFPSQYSRSFFTQCMACLWKQHWSYWRNPEYNAIRFLYSTAV 865
NKALVKELS P P + DLYFP++YS+S Q +C WKQ +YWR+P+YN +R+ ++ A
Sbjct: 198 NKALVKELSTPPPGATDLYFPTKYSQSTLGQFKSCFWKQWLTYWRSPDYNLVRYFFTLAC 377
Query: 866 AVLLGSMFWDLGSKIEKEQDLFNAMGSMYSAVILIGVMNCNSVQPVVAVERTVFYRESAA 925
A+++G++FW +G E DL +G+MY+AVI +G+ NC +VQP+VAVERTVFYRE AA
Sbjct: 378 ALMIGTVFWRIGKNRESSADLTMIIGAMYAAVIFVGINNCQTVQPIVAVERTVFYRERAA 557
Query: 926 G 926
G
Sbjct: 558 G 560
Score = 38.1 bits (87), Expect = 0.021
Identities = 25/75 (33%), Positives = 39/75 (51%)
Frame = +2
Query: 915 ERTVFYRESAAGNVFYFSICLWAGKYPSPICFCTSCGLWHYSVCDDWF*VVYG*SFVVPI 974
+ +V R+S NV F++CL G S IC +C L +SVC *+ G +V +
Sbjct: 527 KNSVLSRKSCR-NVCSFALCLSTGLL*STICVFPNCLLLTHSVCYGEL*MESGEVLLVLL 703
Query: 975 LLVFHLSLLHLLWDD 989
+ +LH+LW+D
Sbjct: 704 CQLLLFLVLHILWND 748
>TC222885 weakly similar to UP|Q8GU89 (Q8GU89) PDR-like ABC transporter (PDR4
ABC transporter), partial (9%)
Length = 484
Score = 111 bits (277), Expect(2) = 7e-38
Identities = 51/71 (71%), Positives = 65/71 (90%)
Frame = +2
Query: 856 AIRFLYSTAVAVLLGSMFWDLGSKIEKEQDLFNAMGSMYSAVILIGVMNCNSVQPVVAVE 915
A+RFL++ AVA+L GS++W+LGSKI+K+QDLFNAMGSMY+AV+L+G+ N NS QP+VAVE
Sbjct: 80 ALRFLFTIAVALLFGSIYWNLGSKIKKQQDLFNAMGSMYAAVLLLGIKNSNSAQPLVAVE 259
Query: 916 RTVFYRESAAG 926
RTVFYRE AAG
Sbjct: 260 RTVFYREKAAG 292
Score = 65.9 bits (159), Expect(2) = 7e-38
Identities = 35/64 (54%), Positives = 42/64 (64%)
Frame = +1
Query: 927 NVFYFSICLWAGKYPSPICFCTSCGLWHYSVCDDWF*VVYG*SFVVPILLVFHLSLLHLL 986
NVF F IC +G + C T+CGL YSV +DWF*VV * F+V +L V HL LL+LL
Sbjct: 292 NVFSFGICFCSGCSRTTSCSSTNCGL*CYSVRNDWF*VVCH*IFLVSLLHVLHLPLLYLL 471
Query: 987 WDDV 990
W DV
Sbjct: 472 WHDV 483
>BF325103
Length = 393
Score = 150 bits (380), Expect = 2e-36
Identities = 69/103 (66%), Positives = 88/103 (84%)
Frame = -2
Query: 82 MRQYVHILKGTVVISLLQPPPETYNLFDDIILLSDSHIIYQGPREHVLEFFESIGFKCPN 141
+RQ VH++ T++ISLLQP PET++LFDDIILLS+ HIIYQGPRE+VL+FFES+GFKCP
Sbjct: 389 LRQLVHVMDVTMIISLLQPAPETFDLFDDIILLSEGHIIYQGPRENVLDFFESVGFKCPE 210
Query: 142 RKGVADFLQEVTSRKDQEQYCEHKDRPYRFITA*TVFLTHFKH 184
RKG+ADFLQEVTSRKDQEQY +D+PYR+++ F+ HF +
Sbjct: 209 RKGIADFLQEVTSRKDQEQYWFARDKPYRYVSV-PEFVAHFNN 84
Score = 33.5 bits (75), Expect = 0.51
Identities = 24/76 (31%), Positives = 44/76 (57%)
Frame = -2
Query: 713 TVVCTIHQPSIDIFESFDELLLLKQGGQEIYVGPLGHNSSNLINHFEGIQGVRKIKDGYN 772
T++ ++ QP+ + F+ FD+++LL + G IY GP N+++ FE + K +
Sbjct: 359 TMIISLLQPAPETFDLFDDIILLSE-GHIIYQGP----RENVLDFFESVG--FKCPERKG 201
Query: 773 PATWMLEVTTSSKERE 788
A ++ EV TS K++E
Sbjct: 200 IADFLQEV-TSRKDQE 156
>BE819956 similar to GP|20522008|db pleiotropic drug resistance like protein
{Nicotiana tabacum}, partial (6%)
Length = 673
Score = 118 bits (295), Expect(2) = 3e-31
Identities = 58/75 (77%), Positives = 67/75 (89%)
Frame = -1
Query: 23 GLEICADTVVGNAMIRAISGGQKKRLTTGEMLVGPTKALFMDEISTGLDSSTTFQIVNSM 82
GLE+CADT+VGNAM+R ISGGQ+KR+TT EMLVG A FMDEISTGLDSSTT+QI+NS+
Sbjct: 445 GLEVCADTIVGNAMLRGISGGQRKRVTTWEMLVGLANA*FMDEISTGLDSSTTYQILNSL 266
Query: 83 RQYVHILKGTVVISL 97
+Q VHILKGT VISL
Sbjct: 265 KQCVHILKGTTVISL 221
Score = 37.0 bits (84), Expect(2) = 3e-31
Identities = 16/19 (84%), Positives = 19/19 (99%)
Frame = -2
Query: 2 KAVATEGQKENLITDYVLR 20
KA+ATEG+KENL+TDYVLR
Sbjct: 501 KALATEGEKENLMTDYVLR 445
Score = 34.7 bits (78), Expect = 0.23
Identities = 23/72 (31%), Positives = 39/72 (53%), Gaps = 2/72 (2%)
Frame = -1
Query: 646 LKPLRNALVGLPGVCGLSMEQRKRLTVAVELVANPSIIFMDEPTSGLDARAAAVVMRTVR 705
L+ + +VG + G+S QRKR+T LV + FMDE ++GLD+ ++ +++
Sbjct: 442 LEVCADTIVGNAMLRGISGGQRKRVTTWEMLVGLANA*FMDEISTGLDSSTTYQILNSLK 263
Query: 706 NTVD--TGRTVV 715
V G TV+
Sbjct: 262 QCVHILKGTTVI 227
>BM108327 weakly similar to GP|27368815|em PDR-like ABC transporter {Oryza
sativa (japonica cultivar-group)}, partial (5%)
Length = 539
Score = 121 bits (303), Expect = 2e-27
Identities = 73/171 (42%), Positives = 95/171 (54%), Gaps = 3/171 (1%)
Frame = +1
Query: 695 RAAAVVMRTVRNTVDTGRTVVCTIHQPSIDIFESFDELLLLKQGGQEIYVGPLGHNSSNL 754
RAA +VMR V+ TGRTV+CTIHQPSI IFE FD LLLL++GG Y G LG +S +
Sbjct: 1 RAAQIVMRGVQAIARTGRTVLCTIHQPSITIFELFDALLLLQRGGYTAYFGDLGKDSVVM 180
Query: 755 INHFEGIQGVRKIKDGYNPATWMLEVTTSSKERELGIDFAELYKNSELYRINKALVKELS 814
+ +F I G +I+ YNPAT+ML+V + R D++ Y S L + N LS
Sbjct: 181 LEYFASIPGTEEIRPQYNPATYMLDVIGAGIGRN-SKDYSVEYAKSALCKANNERTLALS 357
Query: 815 APAP---CSKDLYFPSQYSRSFFTQCMACLWKQHWSYWRNPEYNAIRFLYS 862
P+P L F + F+ Q KQ +YWRNP YN +R S
Sbjct: 358 QPSPEFIPFSTLKF-DPIAXGFWNQFXELTXKQRLTYWRNPXYNVMRLFPS 507
>BI699796 similar to PIR|G85167|G85 ABC transporter like protein [imported] -
Arabidopsis thaliana, partial (13%)
Length = 421
Score = 115 bits (289), Expect = 8e-26
Identities = 56/79 (70%), Positives = 68/79 (85%)
Frame = +1
Query: 639 KVMELVELKPLRNALVGLPGVCGLSMEQRKRLTVAVELVANPSIIFMDEPTSGLDARAAA 698
+V+ +EL ++++LVG+P + GLS EQRKRLT+AVELVANPSIIFMDEPT+GLDARAAA
Sbjct: 184 EVIHTIELDGIKDSLVGMPNISGLSTEQRKRLTIAVELVANPSIIFMDEPTTGLDARAAA 363
Query: 699 VVMRTVRNTVDTGRTVVCT 717
VVMR V+N V TGRTV CT
Sbjct: 364 VVMRAVKNVVGTGRTVACT 420
Score = 42.7 bits (99), Expect = 8e-04
Identities = 23/69 (33%), Positives = 38/69 (54%)
Frame = +1
Query: 18 VLRVLGLEICADTVVGNAMIRAISGGQKKRLTTGEMLVGPTKALFMDEISTGLDSSTTFQ 77
V+ + L+ D++VG I +S Q+KRLT LV +FMDE +TGLD+
Sbjct: 187 VIHTIELDGIKDSLVGMPNISGLSTEQRKRLTIAVELVANPSIIFMDEPTTGLDARAAAV 366
Query: 78 IVNSMRQYV 86
++ +++ V
Sbjct: 367 VMRAVKNVV 393
>TC227242 similar to UP|Q8GU90 (Q8GU90) PDR-like ABC transporter (PDR3 ABC
transporter), partial (17%)
Length = 1126
Score = 107 bits (267), Expect = 3e-23
Identities = 75/236 (31%), Positives = 119/236 (49%), Gaps = 34/236 (14%)
Frame = +1
Query: 854 YNAIRFLYSTAVAVLLGSMFWDLGSKIEKEQDLFNAMGSMYSAVILIGVMNCNSVQPVVA 913
Y A+RF ++T +A++ G+MFWDLGS+ DL NA+GS Y+AV+ +G+ N +SVQPVVA
Sbjct: 4 YTAVRFFFTTFIALMFGTMFWDLGSRRTTRGDLLNALGSTYTAVLFLGIQNASSVQPVVA 183
Query: 914 VERTVFYRESAAG--NVFYFSICLWAGKYPSPICFCTSCGLWHYSVCD-DW--------- 961
VERTV YRE AAG + ++ + P + GL Y++ DW
Sbjct: 184 VERTVLYREKAAGMYSALPYAFAQVLVEIPYIFAQAVTYGLIVYAMIGFDWTAEKFFWYL 363
Query: 962 --------F*VVYG*SFVVPILLVFHLSLL-----HLLWDDVSGLDPKQSHFHYSFLSIL 1008
+ YG V + H++ + + +W+ SG + + L I
Sbjct: 364 FFSFFSLLYFTFYG-MMAVGVTPNHHVAAIVAAAFYAIWNLFSGFIVVRPVSCCTTL*IN 540
Query: 1009 ------LNMESLLRIHSP---KTKNSSVVEVVQLGKSNGMEFVWIGGFTIWRFKKK 1055
N + ++ P T+N+ VVE+V LG S+G++ +WI ++WR+ K
Sbjct: 541 S*MVHGFNGCMYIYVYCPFPFSTENARVVEMVLLGMSSGLDLIWIDCISVWRYNGK 708
>TC206291 similar to PIR|T02644|T02644 ABC-type transport protein homolog
F12C20.5 - Arabidopsis thaliana {Arabidopsis thaliana;}
, partial (16%)
Length = 1292
Score = 67.0 bits (162), Expect(2) = 6e-23
Identities = 32/46 (69%), Positives = 38/46 (82%)
Frame = +1
Query: 881 EKEQDLFNAMGSMYSAVILIGVMNCNSVQPVVAVERTVFYRESAAG 926
E +QDLFNAMGSMYSA++ IG+ N +VQPVV+VER V YRE AAG
Sbjct: 253 ETQQDLFNAMGSMYSAILFIGITNGTAVQPVVSVERFVSYRERAAG 390
Score = 60.1 bits (144), Expect(2) = 6e-23
Identities = 44/129 (34%), Positives = 65/129 (50%), Gaps = 5/129 (3%)
Frame = +3
Query: 927 NVFYFSICLWAGKYPSPICFCTSCG-----LWHYSVCDDWF*VVYG*SFVVPILLVFHLS 981
+VF F IC+ +G P+C T L+H +C D+ *V +V IL VF+
Sbjct: 390 DVFSFIICICSGCD*VPLCVRTGYYILFNILFHGFLCLDFR*V-----HLVLILHVFYDV 554
Query: 982 LLHLLWDDVSGLDPKQSHFHYSFLSILLNMESLLRIHSPKTKNSSVVEVVQLGKSNGMEF 1041
+L+LLW+D +G K + SI +E ++ +N +VE+V LGK ME
Sbjct: 555 ILYLLWNDDNGWHTKS*RCCHHSCSIRYVVEPFQWLYDSSQENPYLVEMVLLGKPCSMES 734
Query: 1042 VWIGGFTIW 1050
VW FT+W
Sbjct: 735 VWSANFTVW 761
>BQ080393
Length = 347
Score = 84.3 bits (207), Expect(2) = 2e-22
Identities = 42/52 (80%), Positives = 48/52 (91%)
Frame = +2
Query: 639 KVMELVELKPLRNALVGLPGVCGLSMEQRKRLTVAVELVANPSIIFMDEPTS 690
+VM+LVEL L++A+VGLPGV GLS EQRKRLT+AVELVANPSIIFMDEPTS
Sbjct: 191 QVMDLVELDNLKDAIVGLPGVTGLSTEQRKRLTIAVELVANPSIIFMDEPTS 346
Score = 40.8 bits (94), Expect(2) = 2e-22
Identities = 21/45 (46%), Positives = 28/45 (61%)
Frame = +1
Query: 588 NFRLSKKARNLCKNFWIL*AN*YPFSLCYCL*IFALSNMAPVVPR 632
N + ++ RNLCK+ W+L* N*Y F+ Y IFAL + PV R
Sbjct: 22 NLWVPQEPRNLCKSLWLL*TN*YSFTPSYY*RIFALLCLPPVTKR 156
Score = 33.5 bits (75), Expect = 0.51
Identities = 18/50 (36%), Positives = 27/50 (54%)
Frame = +2
Query: 16 DYVLRVLGLEICADTVVGNAMIRAISGGQKKRLTTGEMLVGPTKALFMDE 65
D V+ ++ L+ D +VG + +S Q+KRLT LV +FMDE
Sbjct: 188 DQVMDLVELDNLKDAIVGLPGVTGLSTEQRKRLTIAVELVANPSIIFMDE 337
>TC222300 similar to GB|AAP37786.1|30725528|BT008427 At1g51560 {Arabidopsis
thaliana;} , partial (37%)
Length = 801
Score = 100 bits (248), Expect = 4e-21
Identities = 50/134 (37%), Positives = 81/134 (60%)
Frame = +1
Query: 14 ITDYVLRVLGLEICADTVVGNAMIRAISGGQKKRLTTGEMLVGPTKALFMDEISTGLDSS 73
ITD + +GL+ C D ++G +R ISGG+KKRL+ ++ + LF+DE ++GLDS+
Sbjct: 4 ITDATIIEMGLQDCDDRLIGKWHLRGISGGEKKRLSIAL*ILTRPRLLFLDEPTSGLDSA 183
Query: 74 TTFQIVNSMRQYVHILKGTVVISLLQPPPETYNLFDDIILLSDSHIIYQGPREHVLEFFE 133
+ F +V ++R + TV+ S+ QP E + L+DD+ LLS +Y G + +EFF+
Sbjct: 184 SAFFVVQTLRNVARD-ERTVIYSIHQPDSEVFALYDDLFLLSGGETVYFGEAKSAIEFFD 360
Query: 134 SIGFKCPNRKGVAD 147
GF CP ++ D
Sbjct: 361 EAGFPCPRKRNPYD 402
Score = 93.2 bits (230), Expect = 5e-19
Identities = 71/259 (27%), Positives = 128/259 (49%), Gaps = 27/259 (10%)
Frame = +1
Query: 646 LKPLRNALVGLPGVCGLSMEQRKRLTVAVELVANPSIIFMDEPTSGLDARAAAVVMRTVR 705
L+ + L+G + G+S ++KRL++A+ ++ P ++F+DEPTSGLD+ +A V++T+R
Sbjct: 34 LQDCDDRLIGKWHLRGISGGEKKRLSIAL*ILTRPRLLFLDEPTSGLDSASAFFVVQTLR 213
Query: 706 NTVDTGRTVVCTIHQPSIDIFESFDELLLLKQGGQEIYVGPLGH-------------NSS 752
N RTV+ +IHQP ++F +D+L LL GG+ +Y G
Sbjct: 214 NVARDERTVIYSIHQPDSEVFALYDDLFLL-SGGETVYFGEAKSAIEFFDEAGFPCPRKR 390
Query: 753 NLINHF------------EGIQGVRKIKDGYNPATWMLEVTTSSKERELGIDFAELYKNS 800
N +HF I+G ++I D N A + + T+ E+ E YK S
Sbjct: 391 NPYDHFLRCINYDFDIVTATIKGSQRIHDVPNSADPFMNLATA----EIKATLVEKYKRS 558
Query: 801 ELYRINKALVKELSAPAPCSKDLYFPSQY--SRSFFTQCMACLWKQHWSYWRNPEYNAIR 858
R K ++ELS + L+ P+Q+ S++ Q + + + R+ Y +R
Sbjct: 559 TYARRAKNRIQELST----DEGLHPPTQHGSQASWWKQLLTLTKRSFVNMCRDVGYYWLR 726
Query: 859 FLYSTAVAVLLGSMFWDLG 877
+ V++ + + ++D+G
Sbjct: 727 IIVYIIVSICV*TDYFDVG 783
>TC218418 homologue to UP|Q6X4V5 (Q6X4V5) ABC transporter, partial (49%)
Length = 1220
Score = 97.8 bits (242), Expect = 2e-20
Identities = 52/146 (35%), Positives = 85/146 (57%), Gaps = 1/146 (0%)
Frame = +1
Query: 14 ITDYVLRVLGLEICADTVVGNAMIRAISGGQKKRLTTGEMLVGPTKALFMDEISTGLDSS 73
+ + + +GL+ CADTV+GN +R ISGG+K+R++ ++ + LF+DE ++GLDS+
Sbjct: 592 LVESTIVAMGLQDCADTVIGNWHLRGISGGEKRRVSIALEILMRPRLLFLDEPTSGLDSA 771
Query: 74 TTFQIVNSMRQYVHILKGTVVISLLQPPPETYNLFDDIILLSDSHIIYQGPREHVLEFFE 133
+ F + ++R + TV+ S+ QP E + LFD + LLS +Y G EFF
Sbjct: 772 SAFFVTQTLRALARDGR-TVIASIHQPSSEVFELFDQLYLLSSGKTVYFGQASEAYEFFA 948
Query: 134 SIGFKCPNRKGVAD-FLQEVTSRKDQ 158
GF CP + +D FL+ + S D+
Sbjct: 949 QAGFPCPALRNPSDHFLRCINSDFDK 1026
Score = 84.3 bits (207), Expect = 2e-16
Identities = 40/100 (40%), Positives = 69/100 (69%)
Frame = +1
Query: 646 LKPLRNALVGLPGVCGLSMEQRKRLTVAVELVANPSIIFMDEPTSGLDARAAAVVMRTVR 705
L+ + ++G + G+S +++R+++A+E++ P ++F+DEPTSGLD+ +A V +T+R
Sbjct: 622 LQDCADTVIGNWHLRGISGGEKRRVSIALEILMRPRLLFLDEPTSGLDSASAFFVTQTLR 801
Query: 706 NTVDTGRTVVCTIHQPSIDIFESFDELLLLKQGGQEIYVG 745
GRTV+ +IHQPS ++FE FD+L LL G+ +Y G
Sbjct: 802 ALARDGRTVIASIHQPSSEVFELFDQLYLL-SSGKTVYFG 918
>TC229551 weakly similar to UP|Q6X4V5 (Q6X4V5) ABC transporter, partial (34%)
Length = 1436
Score = 97.1 bits (240), Expect = 4e-20
Identities = 80/294 (27%), Positives = 138/294 (46%), Gaps = 17/294 (5%)
Frame = +3
Query: 651 NALVGLPGVCGLSMEQRKRLTVAVELVANPSIIFMDEPTSGLDARAAAVVMRTVRNTVDT 710
N VG GLS QR+RL++ +E++ +P ++F+DEPTSGLD+ A+ VM + N +
Sbjct: 414 NTRVGGWNCKGLSGGQRRRLSICIEILTHPKLLFLDEPTSGLDSAASYYVMSGIANLIQR 593
Query: 711 G---RTVVCTIHQPSIDIFESFDELLLLKQGGQEIYVGPLGHNSSNLINHFEGIQGVRKI 767
RT+V ++HQPS ++F+ F +L LL G+ +Y GP ++ N F G
Sbjct: 594 DGIQRTIVASVHQPSSEVFQLFHDLFLL-SSGETVYFGP-----ASDANQFFASNGF-PC 752
Query: 768 KDGYNPATWMLEVTTS--SKERELGIDFAEL-------YKNSEL-YRINKALVKELSAPA 817
YNP+ L + +++ + GI E YK+SE + + K +
Sbjct: 753 PPLYNPSDHYLRIINKDFNQDADEGITTEEATKILVNSYKSSEFSNHVQSEIAKSETDFG 932
Query: 818 PCSKDLYFPSQYSRSFFTQCMACLWKQHWSYWRNPEYNAIRFLYSTAVAVLLGSMFWDLG 877
C K + +F TQC+ + + +R+ + R + +++ +GS+F+ G
Sbjct: 933 ACGK----KKKIHAAFITQCLILIRRASLQIYRDTNNHRARLVVFIFISLSVGSIFYHSG 1100
Query: 878 S----KIEKEQDLFNAMGSMYSAVILIGVMNCNSVQPVVAVERTVFYRESAAGN 927
I L S+ + + L+G + P++ E VF RE G+
Sbjct: 1101GPDLRSIMDRGSLLCFFLSVVTFMTLVG-----GISPLIE-EMNVFKRERLNGH 1244
Score = 88.2 bits (217), Expect = 2e-17
Identities = 48/139 (34%), Positives = 80/139 (57%), Gaps = 2/139 (1%)
Frame = +3
Query: 4 VATEGQKENLITDYVLRVLGLEICADTVVGNAMIRAISGGQKKRLTTGEMLVGPTKALFM 63
++ E +KE D LR +GL+ +T VG + +SGGQ++RL+ ++ K LF+
Sbjct: 345 MSVEEKKER--ADMTLREMGLQDAINTRVGGWNCKGLSGGQRRRLSICIEILTHPKLLFL 518
Query: 64 DEISTGLDSSTTFQIVNSMRQYVHI--LKGTVVISLLQPPPETYNLFDDIILLSDSHIIY 121
DE ++GLDS+ ++ +++ + + ++ T+V S+ QP E + LF D+ LLS +Y
Sbjct: 519 DEPTSGLDSAASYYVMSGIANLIQRDGIQRTIVASVHQPSSEVFQLFHDLFLLSSGETVY 698
Query: 122 QGPREHVLEFFESIGFKCP 140
GP +FF S GF CP
Sbjct: 699 FGPASDANQFFASNGFPCP 755
>TC218747 similar to UP|Q76CU1 (Q76CU1) PDR-type ABC transporter 2 (Fragment),
partial (9%)
Length = 617
Score = 95.5 bits (236), Expect = 1e-19
Identities = 45/58 (77%), Positives = 51/58 (87%)
Frame = +1
Query: 998 SHFHYSFLSILLNMESLLRIHSPKTKNSSVVEVVQLGKSNGMEFVWIGGFTIWRFKKK 1055
+HF YSFL IL ++ESLLRIHSP T+NSSV+EVVQLGKS MEFVWIGGFTIWR+K K
Sbjct: 1 THFFYSFLCILCSVESLLRIHSPTTENSSVLEVVQLGKSCSMEFVWIGGFTIWRYKAK 174
>BM309256 similar to PIR|T02644|T0 ABC-type transport protein homolog
F12C20.5 - Arabidopsis thaliana, partial (9%)
Length = 441
Score = 92.8 bits (229), Expect = 7e-19
Identities = 42/73 (57%), Positives = 54/73 (73%)
Frame = +1
Query: 36 MIRAISGGQKKRLTTGEMLVGPTKALFMDEISTGLDSSTTFQIVNSMRQYVHILKGTVVI 95
++ + GG KRLTTGE+L+GP + LFMDEISTGLD STT+QI+ ++ L GT ++
Sbjct: 4 VVIVLPGGLHKRLTTGELLIGPARVLFMDEISTGLDRSTTYQIIRYLKHSTRALDGTTIV 183
Query: 96 SLLQPPPETYNLF 108
SLLQP PETY LF
Sbjct: 184 SLLQPAPETYELF 222
Score = 88.6 bits (218), Expect = 1e-17
Identities = 44/89 (49%), Positives = 57/89 (63%)
Frame = +2
Query: 84 QYVHILKGTVVISLLQPPPETYNLFDDIILLSDSHIIYQGPREHVLEFFESIGFKCPNRK 143
Q VH++ G + N F +ILL + I+YQGPRE ++FF+ +GF CP RK
Sbjct: 152 QPVHLM-GPQLYPFFNQLQRPMNCFLYVILLCEGQIVYQGPREAAVDFFKQMGFSCPERK 328
Query: 144 GVADFLQEVTSRKDQEQYCEHKDRPYRFI 172
VADFLQEVTS+KDQEQ DRPYR++
Sbjct: 329 NVADFLQEVTSKKDQEQDWSVPDRPYRYV 415
Score = 34.7 bits (78), Expect = 0.23
Identities = 19/63 (30%), Positives = 35/63 (55%), Gaps = 1/63 (1%)
Frame = +1
Query: 668 KRLTVAVELVANPSIIFMDEPTSGLDARAAAVVMRTVR-NTVDTGRTVVCTIHQPSIDIF 726
KRLT L+ ++FMDE ++GLD ++R ++ +T T + ++ QP+ + +
Sbjct: 34 KRLTTGELLIGPARVLFMDEISTGLDRSTTYQIIRYLKHSTRALDGTTIVSLLQPAPETY 213
Query: 727 ESF 729
E F
Sbjct: 214 ELF 222
>TC204799 similar to UP|Q7PC86 (Q7PC86) PDR7 ABC transporter, partial (10%)
Length = 555
Score = 62.8 bits (151), Expect(2) = 5e-16
Identities = 28/44 (63%), Positives = 36/44 (81%)
Frame = +2
Query: 883 EQDLFNAMGSMYSAVILIGVMNCNSVQPVVAVERTVFYRESAAG 926
+ DL +G+MY+AVI +G+ NC +VQP+VAVERTVFYRE AAG
Sbjct: 5 QADLTMIIGAMYAAVIFVGINNCQTVQPIVAVERTVFYRERAAG 136
Score = 40.8 bits (94), Expect(2) = 5e-16
Identities = 31/96 (32%), Positives = 45/96 (46%)
Frame = +1
Query: 927 NVFYFSICLWAGKYPSPICFCTSCGLWHYSVCDDWF*VVYG*SFVVPILLVFHLSLLHLL 986
NV F++CL G + IC +C L +SVC+ *+ G +V + + +LH+L
Sbjct: 136 NVCSFALCLSTGLLRNTICVFPNCLLLTHSVCNGEL*MESGEVLLVLLCQLLLFLVLHIL 315
Query: 987 WDDVSGLDPKQSHFHYSFLSILLNMESLLRIHSPKT 1022
W+D K SIL + SLL PKT
Sbjct: 316 WNDDCVHHTKPPSGINLCSSILWTL*SLLWFLYPKT 423
>BE806563 similar to GP|11994752|db ABC transporter-like protein {Arabidopsis
thaliana}, partial (20%)
Length = 428
Score = 71.6 bits (174), Expect = 2e-12
Identities = 31/69 (44%), Positives = 48/69 (68%)
Frame = +1
Query: 661 GLSMEQRKRLTVAVELVANPSIIFMDEPTSGLDARAAAVVMRTVRNTVDTGRTVVCTIHQ 720
G+S +RKR+++ E++ NPS++F+DEPTSGLD+ A +++ + GRTVV TIHQ
Sbjct: 220 GISGGERKRVSIGQEMLVNPSLLFVDEPTSGLDSTTAQLIVSVLHGLARAGRTVVATIHQ 399
Query: 721 PSIDIFESF 729
PS ++ F
Sbjct: 400 PSSRLYRMF 426
Score = 63.2 bits (152), Expect = 6e-10
Identities = 44/110 (40%), Positives = 69/110 (62%), Gaps = 3/110 (2%)
Frame = +1
Query: 2 KAVATEGQKENLITDYVLRVLGLEICADTVVGN--AMIRAISGGQKKRLTTG-EMLVGPT 58
K+++ E +KE+ + V+ LGL C ++ VG A+ R ISGG++KR++ G EMLV P+
Sbjct: 109 KSLSREEKKEH--AEMVIAELGLTRCRNSPVGGCMALFRGISGGERKRVSIGQEMLVNPS 282
Query: 59 KALFMDEISTGLDSSTTFQIVNSMRQYVHILKGTVVISLLQPPPETYNLF 108
LF+DE ++GLD STT Q++ S+ + TVV ++ QP Y +F
Sbjct: 283 -LLFVDEPTSGLD-STTAQLIVSVLHGLARAGRTVVATIHQPSSRLYRMF 426
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.341 0.149 0.502
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 51,880,759
Number of Sequences: 63676
Number of extensions: 851740
Number of successful extensions: 8867
Number of sequences better than 10.0: 106
Number of HSP's better than 10.0 without gapping: 8562
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 8836
length of query: 1055
length of database: 12,639,632
effective HSP length: 107
effective length of query: 948
effective length of database: 5,826,300
effective search space: 5523332400
effective search space used: 5523332400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 64 (29.3 bits)
Medicago: description of AC147517.6


