

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
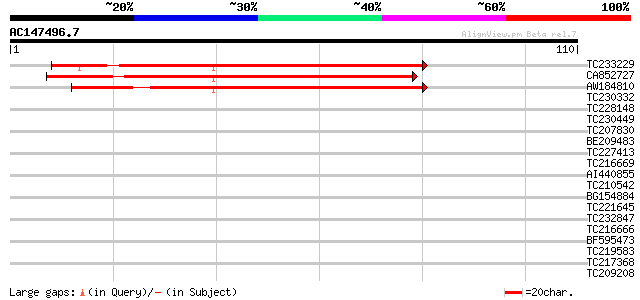
Query= AC147496.7 + phase: 0
(110 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC233229 77 2e-15
CA852727 66 2e-12
AW184810 similar to GP|18390108|gb gb protein {Sorghum bicolor},... 54 1e-08
TC230332 similar to UP|Q9FIL5 (Q9FIL5) Gb|AAB82637.1 (AT5g58960/... 28 0.66
TC228148 similar to UP|Q7YXC8 (Q7YXC8) C. elegans GRD-1 protein ... 26 3.3
TC230449 similar to UP|Q9ZP54 (Q9ZP54) Poly(ADP-ribose) polymera... 25 4.3
TC207830 25 4.3
BE209483 25 4.3
TC227413 similar to UP|Q7PS25 (Q7PS25) ENSANGP00000020753 (Fragm... 25 5.6
TC216669 similar to GB|BAC06968.1|22093674|AP003815 kinase-like ... 25 5.6
AI440855 25 7.3
TC210542 weakly similar to UP|P93806 (P93806) F19P19.1, partial ... 25 7.3
BG154884 25 7.3
TC221645 similar to UP|Q9LSN8 (Q9LSN8) Protein phosphatase 2C-li... 25 7.3
TC232847 UP|Q89DG0 (Q89DG0) Two-component response regulator, pa... 24 9.6
TC216666 similar to GB|BAC06968.1|22093674|AP003815 kinase-like ... 24 9.6
BF595473 homologue to GP|15988117|pdb Chain A Crystal Structure... 24 9.6
TC219583 weakly similar to UP|GRP2_NICSY (P27484) Glycine-rich p... 24 9.6
TC217368 similar to UP|Q93YF4 (Q93YF4) Calcium-dependent protein... 24 9.6
TC209208 similar to PDB|1BOD.0|1431655|1BOD Calbindin D9k Mutant... 24 9.6
>TC233229
Length = 750
Score = 76.6 bits (187), Expect = 2e-15
Identities = 43/75 (57%), Positives = 55/75 (73%), Gaps = 2/75 (2%)
Frame = +1
Query: 9 RNSG-GGGGVKAVKEVVTTAKNVVAAVKEIV-NCTEQEIYDVLRECDMDPNLAVEKLLSQ 66
R SG GGGG KA + ++ +V ++KEIV N E EIY L++C+MDPN AV +LLSQ
Sbjct: 85 RMSGKGGGGQKA--GIPPASRKMVQSLKEIVSNIPEHEIYSTLKDCNMDPNEAVSRLLSQ 258
Query: 67 DTFREVRSKREKRKE 81
DTF EV+SKREK+KE
Sbjct: 259 DTFHEVKSKREKKKE 303
>CA852727
Length = 401
Score = 66.2 bits (160), Expect = 2e-12
Identities = 34/73 (46%), Positives = 51/73 (69%), Gaps = 1/73 (1%)
Frame = +3
Query: 8 NRNSGGGGGVKAVKEVVTTAKNVVAAVKEIV-NCTEQEIYDVLRECDMDPNLAVEKLLSQ 66
+ N+GGG + A + +++ +V ++KEIV N + EIY L++C+MDPN AV +LLSQ
Sbjct: 189 SHNNGGGKALSAT--IPASSRKMVQSLKEIVSNFPDHEIYATLKDCNMDPNEAVSRLLSQ 362
Query: 67 DTFREVRSKREKR 79
D F EV+SKR K+
Sbjct: 363 DPFHEVKSKRXKK 401
>AW184810 similar to GP|18390108|gb gb protein {Sorghum bicolor}, partial
(5%)
Length = 437
Score = 53.5 bits (127), Expect = 1e-08
Identities = 28/70 (40%), Positives = 45/70 (64%), Gaps = 1/70 (1%)
Frame = +2
Query: 13 GGGGVKAVKEVVTTAKNVVAAVKEIV-NCTEQEIYDVLRECDMDPNLAVEKLLSQDTFRE 71
GG G + V + + ++KEIV N ++ +IY L+E +MDPN +KLL+QD F E
Sbjct: 224 GGTGTHLLSARV---RKTIQSIKEIVGNHSDADIYVALKETNMDPNETTQKLLNQDPFHE 394
Query: 72 VRSKREKRKE 81
V+ +R+++KE
Sbjct: 395 VKRRRDRKKE 424
>TC230332 similar to UP|Q9FIL5 (Q9FIL5) Gb|AAB82637.1 (AT5g58960/k19m22_160),
partial (36%)
Length = 827
Score = 28.1 bits (61), Expect = 0.66
Identities = 23/77 (29%), Positives = 38/77 (48%)
Frame = +3
Query: 9 RNSGGGGGVKAVKEVVTTAKNVVAAVKEIVNCTEQEIYDVLRECDMDPNLAVEKLLSQDT 68
R SGGGGG K + ++ VA+V+E+V E + ++ +E + V+ L
Sbjct: 300 RRSGGGGGRKKGRRRGGGGRDGVASVREVVAPYEAVVEELKKE------VKVKDL----- 446
Query: 69 FREVRSKREKRKEVILL 85
EV++ REK + L
Sbjct: 447 --EVKNLREKLDSAVAL 491
>TC228148 similar to UP|Q7YXC8 (Q7YXC8) C. elegans GRD-1 protein
(Corresponding sequence R08B4.1b), partial (3%)
Length = 1180
Score = 25.8 bits (55), Expect = 3.3
Identities = 11/27 (40%), Positives = 15/27 (54%)
Frame = +3
Query: 2 GSESVVNRNSGGGGGVKAVKEVVTTAK 28
G S ++ GGG GVK K++V K
Sbjct: 531 GGSSSSSKGGGGGNGVKEQKKMVNNGK 611
>TC230449 similar to UP|Q9ZP54 (Q9ZP54) Poly(ADP-ribose) polymerase ,
partial (23%)
Length = 925
Score = 25.4 bits (54), Expect = 4.3
Identities = 19/72 (26%), Positives = 36/72 (49%)
Frame = -3
Query: 39 NCTEQEIYDVLRECDMDPNLAVEKLLSQDTFREVRSKREKRKEVILLRYVNIDMRLNLSL 98
N E ++Y VL + V+KLLS D E + +++ ++ ++ LNLS
Sbjct: 770 NSRESKMYYVL--------VMVKKLLSVDDTPEQETSL*S-SSLVMKPHLQ*ELHLNLSS 618
Query: 99 MMNNVCMYFDFR 110
++NN+ + + R
Sbjct: 617 IINNILIVHELR 582
>TC207830
Length = 956
Score = 25.4 bits (54), Expect = 4.3
Identities = 10/29 (34%), Positives = 18/29 (61%)
Frame = +2
Query: 62 KLLSQDTFREVRSKREKRKEVILLRYVNI 90
+L D+F V ++R+E +LLR V++
Sbjct: 287 ELFGMDSFMSVEDDSKQRQETVLLRNVSV 373
>BE209483
Length = 447
Score = 25.4 bits (54), Expect = 4.3
Identities = 18/72 (25%), Positives = 29/72 (40%)
Frame = +1
Query: 9 RNSGGGGGVKAVKEVVTTAKNVVAAVKEIVNCTEQEIYDVLRECDMDPNLAVEKLLSQDT 68
RN G K E + N + +CT + ++C M P KLL +
Sbjct: 97 RNGYDEKGSKRASESIVANGNDSMGHGQKPSCTVEV---QTKQCSMAPAFDARKLLIEKA 267
Query: 69 FREVRSKREKRK 80
+E+R K E+ +
Sbjct: 268 RKEIRKKLEEMR 303
>TC227413 similar to UP|Q7PS25 (Q7PS25) ENSANGP00000020753 (Fragment),
partial (38%)
Length = 1408
Score = 25.0 bits (53), Expect = 5.6
Identities = 11/30 (36%), Positives = 17/30 (56%)
Frame = +1
Query: 31 VAAVKEIVNCTEQEIYDVLRECDMDPNLAV 60
+A V ++ E E YD++R D+D L V
Sbjct: 592 IATVGRAMSSYEDEAYDIMRSLDVDYVLVV 681
>TC216669 similar to GB|BAC06968.1|22093674|AP003815 kinase-like protein
{Oryza sativa (japonica cultivar-group);} , partial
(16%)
Length = 1336
Score = 25.0 bits (53), Expect = 5.6
Identities = 13/36 (36%), Positives = 19/36 (52%)
Frame = +2
Query: 46 YDVLRECDMDPNLAVEKLLSQDTFREVRSKREKRKE 81
+ V +E P L + KLL + R+ + EKRKE
Sbjct: 752 FSVKKELGSKPKLKLLKLLQE*GLRKSQGSGEKRKE 859
>AI440855
Length = 361
Score = 24.6 bits (52), Expect = 7.3
Identities = 12/31 (38%), Positives = 17/31 (54%)
Frame = -3
Query: 7 VNRNSGGGGGVKAVKEVVTTAKNVVAAVKEI 37
V+ N G GVK V ++ AKN+V E+
Sbjct: 203 VDANHAAGIGVKLVNKIRLLAKNLVVLSHEV 111
>TC210542 weakly similar to UP|P93806 (P93806) F19P19.1, partial (27%)
Length = 688
Score = 24.6 bits (52), Expect = 7.3
Identities = 12/34 (35%), Positives = 19/34 (55%)
Frame = +2
Query: 7 VNRNSGGGGGVKAVKEVVTTAKNVVAAVKEIVNC 40
++ +SGGGGG + V A VAA+ ++ C
Sbjct: 353 LSSSSGGGGGHPHTERTVALAVGGVAALGFLIVC 454
>BG154884
Length = 475
Score = 24.6 bits (52), Expect = 7.3
Identities = 12/33 (36%), Positives = 18/33 (54%)
Frame = +2
Query: 75 KREKRKEVILLRYVNIDMRLNLSLMMNNVCMYF 107
+R K K +L Y + NL MMNN C+++
Sbjct: 152 ERSKEKVNFILLYWLC*RKFNLLNMMNNGCLHY 250
>TC221645 similar to UP|Q9LSN8 (Q9LSN8) Protein phosphatase 2C-like protein,
partial (29%)
Length = 680
Score = 24.6 bits (52), Expect = 7.3
Identities = 14/39 (35%), Positives = 21/39 (52%)
Frame = -3
Query: 4 ESVVNRNSGGGGGVKAVKEVVTTAKNVVAAVKEIVNCTE 42
E+V+ +SGGGGG+ VVT + + A + C E
Sbjct: 327 EAVLAFSSGGGGGISG---VVTVPERLPTARDKAHQCME 220
>TC232847 UP|Q89DG0 (Q89DG0) Two-component response regulator, partial
(69%)
Length = 315
Score = 24.3 bits (51), Expect = 9.6
Identities = 10/14 (71%), Positives = 11/14 (78%)
Frame = +3
Query: 46 YDVLRECDMDPNLA 59
YD+LRE DPNLA
Sbjct: 81 YDLLREVRADPNLA 122
>TC216666 similar to GB|BAC06968.1|22093674|AP003815 kinase-like protein
{Oryza sativa (japonica cultivar-group);} , partial
(14%)
Length = 667
Score = 24.3 bits (51), Expect = 9.6
Identities = 14/39 (35%), Positives = 21/39 (52%)
Frame = +3
Query: 46 YDVLRECDMDPNLAVEKLLSQDTFREVRSKREKRKEVIL 84
+ V +E P L + KLL + R+ + EKRKE +L
Sbjct: 129 FSVKKELGSKPKLKLLKLLLE*GPRKSQGSGEKRKEKLL 245
>BF595473 homologue to GP|15988117|pdb Chain A Crystal Structure Of Soybean
Proglycinin A1ab1b Homotrimer, partial (7%)
Length = 412
Score = 24.3 bits (51), Expect = 9.6
Identities = 12/46 (26%), Positives = 27/46 (58%), Gaps = 4/46 (8%)
Frame = -2
Query: 60 VEKLLSQDTFR----EVRSKREKRKEVILLRYVNIDMRLNLSLMMN 101
V ++S +TF E+ K++K ++L +++ + L+LSL ++
Sbjct: 168 VLSIMSHETFAGIKGEIPKKKKKNSRLVLSLSLSLSLSLSLSLSLS 31
>TC219583 weakly similar to UP|GRP2_NICSY (P27484) Glycine-rich protein 2,
partial (45%)
Length = 995
Score = 24.3 bits (51), Expect = 9.6
Identities = 8/23 (34%), Positives = 17/23 (73%)
Frame = +3
Query: 83 ILLRYVNIDMRLNLSLMMNNVCM 105
++L +N+ +RLN L++ N+C+
Sbjct: 849 LMLDLINLLIRLNAVLILQNICI 917
>TC217368 similar to UP|Q93YF4 (Q93YF4) Calcium-dependent protein kinase 2,
partial (66%)
Length = 1349
Score = 24.3 bits (51), Expect = 9.6
Identities = 11/31 (35%), Positives = 19/31 (60%), Gaps = 1/31 (3%)
Frame = +1
Query: 26 TAKNVVAAVKEI-VNCTEQEIYDVLRECDMD 55
T + + A +K + N E EIYD+++ D+D
Sbjct: 736 TFEELKAGLKRVGANLKESEIYDLMQAADVD 828
>TC209208 similar to PDB|1BOD.0|1431655|1BOD Calbindin D9k Mutant With Ala 14
Deleted, Ala 15 Replaced By Asp, Pro 20 Replaced By Gly,
Asn 21 Deleted, And Pro 43 Replaced By Met
(A14del,A15d,P20g,N21del,P43m) (Nmr, 24 Structures).
{Bos taurus;} , partial (22%)
Length = 784
Score = 24.3 bits (51), Expect = 9.6
Identities = 15/53 (28%), Positives = 27/53 (50%)
Frame = +3
Query: 41 TEQEIYDVLRECDMDPNLAVEKLLSQDTFREVRSKREKRKEVILLRYVNIDMR 93
TE +I ++LR+ D + + + K + F+E SK + LLR + + R
Sbjct: 183 TENQIREILRKADSNGDGRLSKEELKKAFKEFGSKMPGWRACRLLRKADTNHR 341
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.135 0.369
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,525,686
Number of Sequences: 63676
Number of extensions: 31049
Number of successful extensions: 275
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 275
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 275
length of query: 110
length of database: 12,639,632
effective HSP length: 86
effective length of query: 24
effective length of database: 7,163,496
effective search space: 171923904
effective search space used: 171923904
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 51 (24.3 bits)
Medicago: description of AC147496.7


