

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147496.2 - phase: 0 /pseudo
(550 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
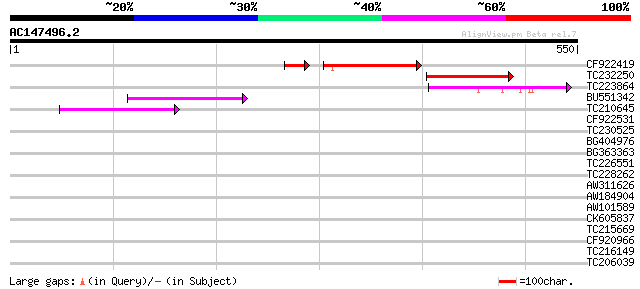
Score E
Sequences producing significant alignments: (bits) Value
CF922419 97 4e-22
TC232250 88 1e-17
TC223864 weakly similar to UP|Q9LIG9 (Q9LIG9) Cytochrome P450-li... 75 1e-13
BU551342 weakly similar to PIR|C86478|C8 protein F15O4.13 [impor... 56 4e-08
TC210645 weakly similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial ... 52 7e-07
CF922531 36 0.051
TC230525 36 0.051
BG404976 35 0.066
BG363363 35 0.087
TC226551 similar to GB|AAP37829.1|30725614|BT008470 At5g12470 {A... 35 0.087
TC228262 similar to UP|Q9MAL4 (Q9MAL4) F27F5.2, partial (4%) 34 0.15
AW311626 34 0.15
AW184904 34 0.15
AW101589 GP|21104742|db OJ1117_G01.13 {Oryza sativa (japonica cu... 34 0.15
CK605837 33 0.25
TC215669 similar to UP|Q9FMU9 (Q9FMU9) Similarity to ATFP3, part... 33 0.33
CF920966 33 0.43
TC216149 weakly similar to UP|MNN4_YEAST (P36044) MNN4 protein, ... 32 0.56
TC206039 32 0.73
>CF922419
Length = 804
Score = 97.1 bits (240), Expect(2) = 4e-22
Identities = 50/98 (51%), Positives = 64/98 (65%), Gaps = 3/98 (3%)
Frame = -2
Query: 305 EEEAEEI---PSGDLFMIRRFLGNQAKEEESNQRETLFHTRCLVQGKVCFLIIDGGSRTN 361
EE +EE+ GDL M+RR LG Q+ + +QRE +F R K C LI+D GS N
Sbjct: 296 EESSEEVYPHEEGDLLMVRRLLGGQSCDLSQSQRENIFLAR*KFLDKTCSLIVDSGSCCN 117
Query: 362 VASTRLVSKMELETKPHPKPYKLQWLNENVEILVDKQV 399
STRLVSK+ L PHPKPY+LQWLNE E++V++QV
Sbjct: 116 CCSTRLVSKLNLTIIPHPKPYELQWLNEQGEMIVNQQV 3
Score = 26.2 bits (56), Expect(2) = 4e-22
Identities = 12/25 (48%), Positives = 17/25 (68%)
Frame = -1
Query: 267 *VFQMSRSRSHSFSLYNKENYDDLG 291
*+ QM+ RSH S+ +KEN+D G
Sbjct: 447 *MLQMTWERSHCLSMPHKENHDHEG 373
>TC232250
Length = 851
Score = 87.8 bits (216), Expect = 1e-17
Identities = 40/84 (47%), Positives = 60/84 (70%)
Frame = +1
Query: 405 IGKYEDDVLCDVVPMEASHLLLGRP*QFDRSVLHDGRTNKYSFMHSGQKISLAPLSPSEV 464
IG Y+D+V CD+VPMEA H+LLGRP QFDR ++ +G TN+ + H G K L P +PS+V
Sbjct: 1 IGTYKDEVNCDIVPMEAGHILLGRPWQFDRKIICNGLTNEITLTHLGTKFVLHPQTPSQV 180
Query: 465 RDDQKKRKEKYEKEKRKIKRKERE 488
DQ K+K ++E++ K+K+++
Sbjct: 181 AKDQLTMKDKRDEEEKLEKQKKKK 252
>TC223864 weakly similar to UP|Q9LIG9 (Q9LIG9) Cytochrome P450-like protein,
partial (5%)
Length = 561
Score = 74.7 bits (182), Expect = 1e-13
Identities = 63/172 (36%), Positives = 89/172 (51%), Gaps = 33/172 (19%)
Frame = +1
Query: 407 KYEDDVLCDVVPMEASHLLLGRP*QFDRSVLHDGRTNKYSFMHSGQK------ISLAPLS 460
++E ++L DVVPMEASH LLGRP* +DR V+H+ TNK+ F+H G+K +S L
Sbjct: 7 QWEYEILXDVVPMEASHFLLGRP**YDRYVVHNDVTNKFPFVHKGKKGDPHTFVSX*GL* 186
Query: 461 PSEVRDDQKKRKEKYE------KEKRKIKRKER-EFA*KKK-------KRRVSRNS---H 503
S + +K+ +EK KEK +RKER E+ *KKK K R +N+ H
Sbjct: 187 GSNKNESEKRTREKRREKQN**KEKA*KERKERKEWR*KKKRNCKKR*KSRREKNTELVH 366
Query: 504 EQ----------TAPIFTTLQKVALLTNTQNFPSCTKFLLQECEDVFPKKVP 545
E+ +FT + + + F + LL+E DVFPK P
Sbjct: 367 ERKRGEESDAS*ATYVFTNVS*LWFVFCC*FFA*GVEELLKEFGDVFPKDTP 522
>BU551342 weakly similar to PIR|C86478|C8 protein F15O4.13 [imported] -
Arabidopsis thaliana, partial (5%)
Length = 467
Score = 56.2 bits (134), Expect = 4e-08
Identities = 44/116 (37%), Positives = 62/116 (52%)
Frame = -2
Query: 115 LG*DEENHEEKVCAFLLP*RLAQQIAKTYSRFEKC*RIFQGDGDCEN*S*CGGRQ*SSHG 174
LG E E+K+C+F L R + Q KT++R KC* + QGDG +*S*C G +G
Sbjct: 445 LGXXEXVDEKKICSFSLSKRAS*QATKTHTR**KC**VLQGDGGFHD*S*CYGGLRGYYG 266
Query: 175 *ISSWSKS*H**HS*TSSLC*NG*LGTSSYQSRTTTQKKSPS*KKHFFF*LFKLEG 230
+W+K * + C G L TSS+ + TTT++K ++ F L LEG
Sbjct: 265 SFPTWAK**CKGCCGVAKSCGIGELSTSSF*AGTTTEEKECHEEEFI*FSLS*LEG 98
>TC210645 weakly similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial (3%)
Length = 832
Score = 52.0 bits (123), Expect = 7e-07
Identities = 45/116 (38%), Positives = 64/116 (54%)
Frame = +3
Query: 49 PIFCWKE*PGSIPRTGD*N*TNF*LP*LLYC*KGAGCFH*VQGVCFSMVGSIDQRKKEVW 108
PIF ++* S+ D +* L L *K G H V + +VG +R+ ++
Sbjct: 396 PIFHGEK*SKSLFGVRDED*GTDCLSQLYKR*KDEGGSHGVH*LSSYLVGPTIKREVKI* 575
Query: 109 RTTD*YLG*DEENHEEKVCAFLLP*RLAQQIAKTYSRFEKC*RIFQGDGDCEN*S* 164
RT D LG DE+ +E+KVC F L *R +QQ +K ++R +KC I Q DG +*S*
Sbjct: 576 RTLDE*LGGDEKINEKKVCFFSLS*RPSQQASKAHTRKQKCR*ILQRDGGFLD*S* 743
>CF922531
Length = 602
Score = 35.8 bits (81), Expect = 0.051
Identities = 34/128 (26%), Positives = 53/128 (40%), Gaps = 31/128 (24%)
Frame = -2
Query: 449 HSGQKISLAPLSPSEVRDDQ----KKRKEKYEKEKRKIKRKEREFA*KKK---------- 494
H K L P +PS+V DQ KR E+ + EK+K K+ + + K K
Sbjct: 586 HLSTKXVLHPQTPSQVAKDQLTMKDKRDEEEKLEKQKKKKDSKALSSKAKGKEKKEKDSS 407
Query: 495 KRRVSRNSHEQTAPIF-----------------TTLQKVALLTNTQNFPSCTKFLLQECE 537
K+ + + +H T T+L V + T + P + LL E
Sbjct: 406 KKIIKKENHFATKGDIKIALLLKQSFYLLLSRETSLSTVTIPT-IETLPPKVQELLHEFG 230
Query: 538 DVFPKKVP 545
D+FPK++P
Sbjct: 229 DIFPKEIP 206
>TC230525
Length = 551
Score = 35.8 bits (81), Expect = 0.051
Identities = 25/84 (29%), Positives = 38/84 (44%)
Frame = -3
Query: 455 SLAPLSPSEVRDDQKKRKEKYEKEKRKIKRKEREFA*KKKKRRVSRNSHEQTAPIFTTLQ 514
+L PLSP EV DDQ + +EK E+EK + +R K ++HE+ F
Sbjct: 258 TLKPLSPREVCDDQIRMREKREQEKENSEAPQRNMK-KSDTPEEKSDTHERKFNYFAEAS 82
Query: 515 KVALLTNTQNFPSCTKFLLQECED 538
+ + P+ LQ C+D
Sbjct: 81 EARKV-----LPAHEPLNLQYCKD 25
>BG404976
Length = 305
Score = 35.4 bits (80), Expect = 0.066
Identities = 21/61 (34%), Positives = 31/61 (50%)
Frame = -2
Query: 436 VLHDGRTNKYSFMHSGQKISLAPLSPSEVRDDQKKRKEKYEKEKRKIKRKEREFA*KKKK 495
+L+ + Y + +L PLS EV DDQ + +EK E+EK + +R KKK
Sbjct: 187 LLNRNKEEPYKRIAKSSSFTLKPLSHREVCDDQIRMREKREQEKENSEAPQRNM--KKKP 14
Query: 496 R 496
R
Sbjct: 13 R 11
>BG363363
Length = 401
Score = 35.0 bits (79), Expect = 0.087
Identities = 15/38 (39%), Positives = 31/38 (81%)
Frame = +1
Query: 468 QKKRKEKYEKEKRKIKRKEREFA*KKKKRRVSRNSHEQ 505
QKK+K+K +K+K+K K+K+++ KKKK+++ +N+ ++
Sbjct: 115 QKKKKKKKKKKKKKKKKKKKK---KKKKKKLKKNNKKK 219
Score = 31.6 bits (70), Expect = 0.96
Identities = 15/46 (32%), Positives = 27/46 (58%)
Frame = +1
Query: 468 QKKRKEKYEKEKRKIKRKEREFA*KKKKRRVSRNSHEQTAPIFTTL 513
+KK+K+K +K+K+K K+K++ KKK + + P F+ L
Sbjct: 136 KKKKKKKKKKKKKKKKKKKKLKKNNKKKNKKKKKKXXLXFPFFSPL 273
Score = 31.2 bits (69), Expect = 1.3
Identities = 15/29 (51%), Positives = 24/29 (82%)
Frame = +3
Query: 468 QKKRKEKYEKEKRKIKRKEREFA*KKKKR 496
+KK+K+K +K+K+KIK+K + *KKKK+
Sbjct: 153 KKKKKKKKKKKKKKIKKK**KKK*KKKKK 239
>TC226551 similar to GB|AAP37829.1|30725614|BT008470 At5g12470 {Arabidopsis
thaliana;} , partial (75%)
Length = 1883
Score = 35.0 bits (79), Expect = 0.087
Identities = 15/34 (44%), Positives = 27/34 (79%)
Frame = +1
Query: 468 QKKRKEKYEKEKRKIKRKEREFA*KKKKRRVSRN 501
+KK+K+K +K+K+K KRK+++ KKKK++ +N
Sbjct: 1552 KKKKKKKKKKKKKKKKRKKKKKKKKKKKKKKKKN 1653
Score = 33.9 bits (76), Expect = 0.19
Identities = 16/42 (38%), Positives = 28/42 (66%)
Frame = +2
Query: 468 QKKRKEKYEKEKRKIKRKEREFA*KKKKRRVSRNSHEQTAPI 509
+KK+K+K +K+K+K K K+++ KKKK++ + API
Sbjct: 1553 KKKKKKKKKKKKKKKKEKKKKKKKKKKKKKKKKTRGGARAPI 1678
>TC228262 similar to UP|Q9MAL4 (Q9MAL4) F27F5.2, partial (4%)
Length = 657
Score = 34.3 bits (77), Expect = 0.15
Identities = 15/42 (35%), Positives = 27/42 (63%)
Frame = +3
Query: 465 RDDQKKRKEKYEKEKRKIKRKEREFA*KKKKRRVSRNSHEQT 506
R+++K +KEK +EKRK K K+ + ++K + RN ++T
Sbjct: 45 REEEKAKKEKEREEKRKEKEKKEKDREREKDKSKERNKKDET 170
>AW311626
Length = 385
Score = 34.3 bits (77), Expect = 0.15
Identities = 14/31 (45%), Positives = 26/31 (83%)
Frame = +1
Query: 467 DQKKRKEKYEKEKRKIKRKEREFA*KKKKRR 497
D+KK+K+K +K+K+K K+K+++ KKKK++
Sbjct: 40 DEKKKKKKKKKKKKKKKKKKKKKKKKKKKKK 132
>AW184904
Length = 365
Score = 34.3 bits (77), Expect = 0.15
Identities = 16/43 (37%), Positives = 28/43 (64%)
Frame = +2
Query: 468 QKKRKEKYEKEKRKIKRKEREFA*KKKKRRVSRNSHEQTAPIF 510
+KKRK+K +K+K+K K+K+++ KKKK++ + P F
Sbjct: 11 KKKRKKKKKKKKKKKKKKKKKKKKKKKKKKKKKGGAVLGVPTF 139
Score = 30.0 bits (66), Expect = 2.8
Identities = 13/30 (43%), Positives = 24/30 (79%)
Frame = +3
Query: 468 QKKRKEKYEKEKRKIKRKEREFA*KKKKRR 497
+KKR +K +K+K+K K+K+++ KKKK++
Sbjct: 9 KKKRGKKKKKKKKKKKKKKKKKKKKKKKKK 98
>AW101589 GP|21104742|db OJ1117_G01.13 {Oryza sativa (japonica
cultivar-group)}, partial (24%)
Length = 235
Score = 34.3 bits (77), Expect = 0.15
Identities = 16/43 (37%), Positives = 29/43 (67%)
Frame = +3
Query: 468 QKKRKEKYEKEKRKIKRKEREFA*KKKKRRVSRNSHEQTAPIF 510
+KK+K+K +K+K+K K+K+++ KKKK++ + PIF
Sbjct: 66 KKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKGGALLKDPIF 194
>CK605837
Length = 498
Score = 33.5 bits (75), Expect = 0.25
Identities = 14/38 (36%), Positives = 28/38 (72%)
Frame = +1
Query: 468 QKKRKEKYEKEKRKIKRKEREFA*KKKKRRVSRNSHEQ 505
QKK+K+K +K+K+K K+K+++ KKKK++ + ++
Sbjct: 61 QKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKK 174
>TC215669 similar to UP|Q9FMU9 (Q9FMU9) Similarity to ATFP3, partial (54%)
Length = 1305
Score = 33.1 bits (74), Expect = 0.33
Identities = 22/62 (35%), Positives = 30/62 (47%), Gaps = 8/62 (12%)
Frame = +3
Query: 469 KKRKEKYEKEKRKIKRKEREFA*KKKKRRVSR--------NSHEQTAPIFTTLQKVALLT 520
+KRK K+ K+KR+ KRK R K+KK + R H+Q T L + L
Sbjct: 807 QKRKRKWRKQKRRRKRKRRRVVVKEKKTKRRRKKKPK*RKQQHQQLLKTPTRLFQRLRLM 986
Query: 521 NT 522
NT
Sbjct: 987 NT 992
>CF920966
Length = 614
Score = 32.7 bits (73), Expect = 0.43
Identities = 19/54 (35%), Positives = 30/54 (55%)
Frame = -3
Query: 452 QKISLAPLSPSEVRDDQKKRKEKYEKEKRKIKRKEREFA*KKKKRRVSRNSHEQ 505
Q +S P P + + KKR++ + KRK KRKER+F K+K ++ + Q
Sbjct: 468 QALSQNPD*P*TLPSEAKKRRKGNFQSKRKQKRKERKFPIKEKAKKEGKEIPNQ 307
>TC216149 weakly similar to UP|MNN4_YEAST (P36044) MNN4 protein, partial (4%)
Length = 2040
Score = 32.3 bits (72), Expect = 0.56
Identities = 17/48 (35%), Positives = 31/48 (64%), Gaps = 2/48 (4%)
Frame = +1
Query: 460 SPSEVRDDQKKRKEKYEKE--KRKIKRKEREFA*KKKKRRVSRNSHEQ 505
SP R +KKR+++ +++ K K KR++R+ KKK+RR S+ ++
Sbjct: 1234 SPRRKRSQRKKRRKRSQRKRVKPKRKRRKRKSLRKKKRRRKSQRKKKR 1377
>TC206039
Length = 834
Score = 32.0 bits (71), Expect = 0.73
Identities = 14/30 (46%), Positives = 26/30 (86%)
Frame = -2
Query: 468 QKKRKEKYEKEKRKIKRKEREFA*KKKKRR 497
+KK+K+K +K+K+KIK+K+++ KKKK++
Sbjct: 224 KKKKKKKKKKKKKKIKKKKKK---KKKKKK 144
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.358 0.160 0.569
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 22,630,514
Number of Sequences: 63676
Number of extensions: 299227
Number of successful extensions: 3600
Number of sequences better than 10.0: 73
Number of HSP's better than 10.0 without gapping: 2045
Number of HSP's successfully gapped in prelim test: 159
Number of HSP's that attempted gapping in prelim test: 1438
Number of HSP's gapped (non-prelim): 2316
length of query: 550
length of database: 12,639,632
effective HSP length: 102
effective length of query: 448
effective length of database: 6,144,680
effective search space: 2752816640
effective search space used: 2752816640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC147496.2


