

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147484.9 - phase: 0
(125 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
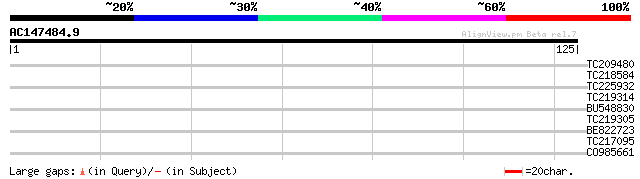
Score E
Sequences producing significant alignments: (bits) Value
TC209480 similar to UP|Q8VXV0 (Q8VXV0) AT3g52570/F22O6_50, parti... 29 0.63
TC218584 28 1.1
TC225932 27 3.1
TC219314 weakly similar to UP|Q7XUE7 (Q7XUE7) OJ991113_30.24 pro... 26 4.1
BU548830 26 4.1
TC219305 weakly similar to UP|Q9LSE7 (Q9LSE7) Emb|CAB45497.1 (AT... 26 4.1
BE822723 25 7.0
TC217095 similar to PDB|1I24_A.0|33357043|1I24_A Chain A, High R... 25 9.1
CO985661 25 9.1
>TC209480 similar to UP|Q8VXV0 (Q8VXV0) AT3g52570/F22O6_50, partial (38%)
Length = 1005
Score = 28.9 bits (63), Expect = 0.63
Identities = 12/52 (23%), Positives = 24/52 (46%), Gaps = 1/52 (1%)
Frame = +3
Query: 27 GKMNTRRGSTEVHEFDGKFRKAPARFKVTS-VIGHVFSVDFPGKYQDWAATD 77
G++N + EV + P++F ++GH+ + +P DW T+
Sbjct: 306 GEVNQENSNDEVRNVLATLQSLPSQFSSRKWLVGHLMGLGYPKALSDWIGTN 461
>TC218584
Length = 637
Score = 28.1 bits (61), Expect = 1.1
Identities = 13/29 (44%), Positives = 17/29 (57%)
Frame = +2
Query: 95 LVDVVIWYCGWIVTVKEKIYVLRVIFVSL 123
LV +V W C ++T+ IYV IF SL
Sbjct: 539 LVQIVWWCCSKVITL*SHIYVTTGIFFSL 625
>TC225932
Length = 2244
Score = 26.6 bits (57), Expect = 3.1
Identities = 12/38 (31%), Positives = 18/38 (46%), Gaps = 3/38 (7%)
Frame = +2
Query: 89 NESNPKLVDVVIW---YCGWIVTVKEKIYVLRVIFVSL 123
N NP L+ +V+W W T++ +IY SL
Sbjct: 812 NNLNPNLLSIVVWNICLICWFTTIRLRIYCTMKYLTSL 925
>TC219314 weakly similar to UP|Q7XUE7 (Q7XUE7) OJ991113_30.24 protein,
partial (5%)
Length = 898
Score = 26.2 bits (56), Expect = 4.1
Identities = 11/34 (32%), Positives = 18/34 (52%), Gaps = 2/34 (5%)
Frame = -1
Query: 86 VIKNESNPKLVDVVIW--YCGWIVTVKEKIYVLR 117
+++ N K + +W CGWI ++ IY LR
Sbjct: 409 LLQYNMNYKTRQIQLWGIICGWIFRIRNSIYKLR 308
>BU548830
Length = 712
Score = 26.2 bits (56), Expect = 4.1
Identities = 11/34 (32%), Positives = 18/34 (52%), Gaps = 2/34 (5%)
Frame = +1
Query: 86 VIKNESNPKLVDVVIW--YCGWIVTVKEKIYVLR 117
+++ N K + +W CGWI ++ IY LR
Sbjct: 430 LLQYNMNYKTRQIQLWGIICGWIFRIRNSIYKLR 531
>TC219305 weakly similar to UP|Q9LSE7 (Q9LSE7) Emb|CAB45497.1
(AT3g25290/MJL12_25), partial (39%)
Length = 627
Score = 26.2 bits (56), Expect = 4.1
Identities = 14/48 (29%), Positives = 21/48 (43%)
Frame = -1
Query: 31 TRRGSTEVHEFDGKFRKAPARFKVTSVIGHVFSVDFPGKYQDWAATDP 78
T R S E+ G+F+ + + +IG S FP + W T P
Sbjct: 219 TMRKSVELQTKRGRFQYSDEHCIESPLIGFQVSCQFPSQQPQWRKTIP 76
>BE822723
Length = 552
Score = 25.4 bits (54), Expect = 7.0
Identities = 6/22 (27%), Positives = 14/22 (63%)
Frame = +2
Query: 87 IKNESNPKLVDVVIWYCGWIVT 108
+ ++ P L +++W+C W+ T
Sbjct: 197 VAHDHRPLLYMLLVWHCAWLAT 262
>TC217095 similar to PDB|1I24_A.0|33357043|1I24_A Chain A, High Resolution
Crystal Structure Of The Wild-Type Protein Sqd1, With
Nad And Udp-Glucose. {Arabidopsis thaliana;} , partial
(84%)
Length = 1346
Score = 25.0 bits (53), Expect = 9.1
Identities = 18/52 (34%), Positives = 25/52 (47%), Gaps = 8/52 (15%)
Frame = +3
Query: 21 ATVLSHGKM-NTRRGSTEVHE-------FDGKFRKAPARFKVTSVIGHVFSV 64
AT L+ G + R T +HE +DG F A RF V + +GH +V
Sbjct: 465 ATDLNQGVVYGVRTDETAMHEELYNRFDYDGIFGTALNRFCVQAAVGHPLTV 620
>CO985661
Length = 553
Score = 25.0 bits (53), Expect = 9.1
Identities = 11/23 (47%), Positives = 15/23 (64%)
Frame = -3
Query: 79 LDLFQAQVIKNESNPKLVDVVIW 101
+ L AQV++ +SN K VVIW
Sbjct: 353 ISLDNAQVVRXDSNAKRNTVVIW 285
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.323 0.137 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,076,652
Number of Sequences: 63676
Number of extensions: 55948
Number of successful extensions: 271
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 270
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 271
length of query: 125
length of database: 12,639,632
effective HSP length: 86
effective length of query: 39
effective length of database: 7,163,496
effective search space: 279376344
effective search space used: 279376344
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 53 (25.0 bits)
Medicago: description of AC147484.9


