

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147484.3 - phase: 0 /pseudo
(526 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
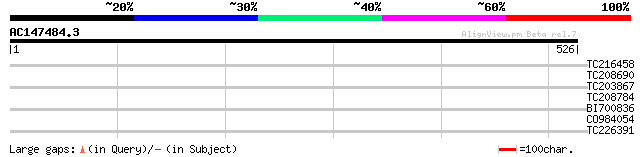
Score E
Sequences producing significant alignments: (bits) Value
TC216458 similar to UP|Q9WUG9 (Q9WUG9) ARL-6 interacting protein... 32 0.69
TC208690 similar to GB|AAD49977.1|5734712|F24J5 ESTs gb|T41781 a... 30 2.0
TC203867 similar to UP|FK22_ARATH (Q38936) FK506-binding protein... 30 3.4
TC208784 similar to UP|Q9FYC8 (Q9FYC8) Non-phototropic hypocotyl... 29 5.9
BI700836 29 5.9
CO984054 25 6.6
TC226391 homologue to GB|AAN31107.1|23506191|AY149953 At5g61670/... 28 7.7
>TC216458 similar to UP|Q9WUG9 (Q9WUG9) ARL-6 interacting protein-5
(Fragment), partial (14%)
Length = 1207
Score = 32.0 bits (71), Expect = 0.69
Identities = 21/78 (26%), Positives = 38/78 (47%), Gaps = 5/78 (6%)
Frame = +3
Query: 68 IPQFLHSIVTFHPHFLVSPLVAALSSFD-----DEPIAEQLVHLILNLSASSDPSVCREF 122
+P+F++ ++ F PHFL P + + + + AE L+ + + SS P EF
Sbjct: 75 LPRFVNLLIQFIPHFLFQPNLFSDNQREIIMDWGNVTAEDLIDALREVDWSSPPRPLSEF 254
Query: 123 VARVSDRVSSGALGWSSR 140
+R + V + W+SR
Sbjct: 255 FSRFT--VPRSSSKWNSR 302
>TC208690 similar to GB|AAD49977.1|5734712|F24J5 ESTs gb|T41781 and
gb|AA586096 come from this gene. {Arabidopsis thaliana;}
, partial (70%)
Length = 585
Score = 30.4 bits (67), Expect = 2.0
Identities = 10/31 (32%), Positives = 20/31 (64%)
Frame = -1
Query: 145 LHCLGVLLNCEKEDDLHVHIKDVYSLISVLV 175
LH + ++NC LH H+ +++SL+ V++
Sbjct: 219 LHDVNRIINCHSRYHLHQHLHNIFSLVEVVI 127
>TC203867 similar to UP|FK22_ARATH (Q38936) FK506-binding protein 2-2
precursor (Peptidyl-prolyl cis-trans isomerase)
(PPIase) (Rotamase) (15 kDa FKBP) (FKBP-15-2) , partial
(12%)
Length = 938
Score = 29.6 bits (65), Expect = 3.4
Identities = 17/34 (50%), Positives = 18/34 (52%)
Frame = +3
Query: 455 KEAYAYSHDENSTHISKLELRSGILDICRTHLLP 488
KE A H E S+H S L LR GI C LLP
Sbjct: 48 KEFPALFHHERSSHSSLLPLRFGIRYCCSLPLLP 149
>TC208784 similar to UP|Q9FYC8 (Q9FYC8) Non-phototropic hypocotyl 3-like
protein, partial (16%)
Length = 1068
Score = 28.9 bits (63), Expect = 5.9
Identities = 30/109 (27%), Positives = 46/109 (41%), Gaps = 14/109 (12%)
Frame = -3
Query: 19 SQGHRSSLN---LQTNQGG--------TICLVCFSNLISNPSSPTIHVSYALSQLSRSLS 67
S G +SL+ L TN+G T+ + CF L S+ SSPT+ + ++S S
Sbjct: 472 SDGRGTSLHPIFLATNRGELREL*DFVTLFISCFILLNSSSSSPTLELILSISTFKT*FS 293
Query: 68 IPQFLH-SIVTFHPHFLVSPLV--AALSSFDDEPIAEQLVHLILNLSAS 113
H S+ T P L ++ A + LV+ +L S S
Sbjct: 292 RTTVTHPSLCTISPAVLPRAVIGTGAAGAISPSNALRHLVNAVLKCSCS 146
>BI700836
Length = 420
Score = 28.9 bits (63), Expect = 5.9
Identities = 11/30 (36%), Positives = 17/30 (56%)
Frame = +1
Query: 336 LFEILRLSVPFHPVQYETLKLIYECISECP 365
L+ +L L +PFH + + KL C +CP
Sbjct: 190 LYPMLFLPIPFHSTHHSSQKLFLFC*KDCP 279
>CO984054
Length = 817
Score = 25.4 bits (54), Expect(2) = 6.6
Identities = 19/93 (20%), Positives = 41/93 (43%), Gaps = 1/93 (1%)
Frame = +3
Query: 21 GHRSSLNLQTNQGGTICLVCFSNLISNPSSPTIHVSYALSQLSRSLSIPQFLHSIVTFHP 80
GH + + + ++C + S ++ S ++ +++ + P FLHS
Sbjct: 297 GHHLCRSGKEARPASVCFLILSRSATSDCSSYSSITLCFRKITGAS*SPDFLHSFAKPCS 476
Query: 81 HFLVSPL-VAALSSFDDEPIAEQLVHLILNLSA 112
+L P V S+ +P++ L ++LN S+
Sbjct: 477 IYLQLPQEVRKKSTTTKDPVSLALSRILLNSSS 575
Score = 21.6 bits (44), Expect(2) = 6.6
Identities = 8/33 (24%), Positives = 21/33 (63%)
Frame = +1
Query: 144 SLHCLGVLLNCEKEDDLHVHIKDVYSLISVLVT 176
+LH +LL+ +++ D H+H ++ ++ L++
Sbjct: 565 TLHQW*ILLHIQRDRDHHLHQTNLLDILQQLLS 663
>TC226391 homologue to GB|AAN31107.1|23506191|AY149953 At5g61670/k11j9_190
{Arabidopsis thaliana;} , partial (25%)
Length = 595
Score = 28.5 bits (62), Expect = 7.7
Identities = 23/79 (29%), Positives = 34/79 (42%), Gaps = 11/79 (13%)
Frame = +3
Query: 16 WCCSQGHRSSLNLQTNQGGTICLVCFSNLISNPSSPTIHVSYAL-----------SQLSR 64
W S G ++ + GGT + + P +PT+ + L S LS
Sbjct: 129 WVDSAGSLTTQSNPVGVGGTTPITGPDGV*WPPRNPTLLLHPLLTPTKPPLLGPFSILSL 308
Query: 65 SLSIPQFLHSIVTFHPHFL 83
SLS+ FLHS + F+P L
Sbjct: 309 SLSLFSFLHSSILFYPQIL 365
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.137 0.397
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,386,704
Number of Sequences: 63676
Number of extensions: 387057
Number of successful extensions: 2182
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 2167
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2182
length of query: 526
length of database: 12,639,632
effective HSP length: 102
effective length of query: 424
effective length of database: 6,144,680
effective search space: 2605344320
effective search space used: 2605344320
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC147484.3


