

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
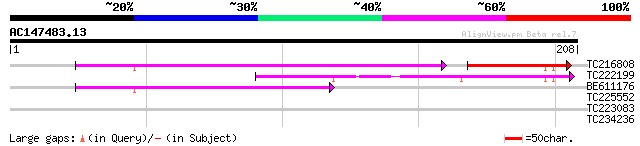
Query= AC147483.13 - phase: 0
(208 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC216808 60 3e-15
TC222199 62 2e-10
BE611176 45 2e-05
TC225552 similar to UP|Q9AYT8 (Q9AYT8) Mangrin, partial (68%) 30 0.78
TC223083 27 5.1
TC234236 similar to UP|Q9LW21 (Q9LW21) Arabidopsis thaliana geno... 27 5.1
>TC216808
Length = 1144
Score = 60.1 bits (144), Expect(2) = 3e-15
Identities = 38/138 (27%), Positives = 66/138 (47%), Gaps = 2/138 (1%)
Frame = +2
Query: 25 KKTKSYKFKEVDLVSLRELA--LKVKNQTGFRLQYGGLLTILRTNVEEKLVHTLVQFYDP 82
K+ K K +++ SL EL + + FR YG + + R V + + +L Q+YD
Sbjct: 146 KRFYQVKIKSLEVTSLGELGQLMGQLQR*AFRKAYGKIWDLARVEVSMEPIASLSQYYDQ 325
Query: 83 SFRCFTFPDFQVVPTLEAYSHLLDSPTAEKTPFTGPGTSLTPLVVAKDLYLKTSDVSNHL 142
RCFTF DFQ+VPT++ + + P+ G + VAK + + +++
Sbjct: 326 PLRCFTFEDFQLVPTVKELEEIRGCQLGGRKPYLFSGFYPSTTRVAKVVKISAQELNRVK 505
Query: 143 TTKSHIRGFTSKYLLEQA 160
++ + G K L E+A
Sbjct: 506 QNRNGVVGVPRKCLEERA 559
Score = 38.1 bits (87), Expect(2) = 3e-15
Identities = 21/43 (48%), Positives = 30/43 (68%), Gaps = 5/43 (11%)
Frame = +3
Query: 169 ILALLIYGLILFPSLDNFVDMNAIKIF----HSK-NPVPTLLA 206
ILALLI+G++LFP+++ VD+ AI F H K +PV +LA
Sbjct: 603 ILALLIFGVVLFPNMEGLVDLAAIDAFLAFHHGKESPVVAILA 731
>TC222199
Length = 691
Score = 62.0 bits (149), Expect = 2e-10
Identities = 48/132 (36%), Positives = 68/132 (51%), Gaps = 15/132 (11%)
Frame = +2
Query: 91 DFQVVPTLEAYSHLLDSPTAEKTPFTG----PGTSLTPLVVAKDLYLKTSDVSNHLTTKS 146
DFQ+VPT+E + +L P E+ P+ P S P +V KD V T++
Sbjct: 5 DFQLVPTIEEFEEILGCPLGERKPYLSSGCLPSLSRIPTMV-KDSARGLDRVKQ---TRN 172
Query: 147 HIRGFTSKYLLEQATCQD------ALEAILALLIYGLILFPSLDNFVDMNAIKIF----H 196
I G KYL ++A +LALLI+G++LFP++D VD+ AI F H
Sbjct: 173 GIAGLPRKYLEDKARGIANQGDWVPFMDVLALLIFGVVLFPNVDGLVDLAAIDAFLAYHH 352
Query: 197 SK-NPVPTLLAD 207
SK +PV +LAD
Sbjct: 353 SKESPVVAVLAD 388
>BE611176
Length = 424
Score = 45.4 bits (106), Expect = 2e-05
Identities = 28/97 (28%), Positives = 43/97 (43%), Gaps = 2/97 (2%)
Frame = -2
Query: 25 KKTKSYKFKEVDLVSLRELA--LKVKNQTGFRLQYGGLLTILRTNVEEKLVHTLVQFYDP 82
K+ K K +D+ S++EL + FR Y +L + + + + +L Q+YD
Sbjct: 360 KRFYQVKIKGLDITSIKELGRLMGPLQMQAFRKAYEKILDLTIAEISTEAIASLTQYYD* 181
Query: 83 SFRCFTFPDFQVVPTLEAYSHLLDSPTAEKTPFTGPG 119
RCFTF DFQ+ L+ D E PG
Sbjct: 180 PLRCFTFGDFQLDRPLKNLRRF*DVLLGEGNRIFSPG 70
>TC225552 similar to UP|Q9AYT8 (Q9AYT8) Mangrin, partial (68%)
Length = 1660
Score = 30.0 bits (66), Expect = 0.78
Identities = 21/62 (33%), Positives = 25/62 (39%)
Frame = +2
Query: 82 PSFRCFTFPDFQVVPTLEAYSHLLDSPTAEKTPFTGPGTSLTPLVVAKDLYLKTSDVSNH 141
P+F T F V P S L S T PFT +SL + L L TS H
Sbjct: 116 PTFALRTISSFSVKPVTTTRSSLTTSSTTTLLPFTSTNSSL-----PRTLKLNTSSPHPH 280
Query: 142 LT 143
L+
Sbjct: 281 LS 286
>TC223083
Length = 243
Score = 27.3 bits (59), Expect = 5.1
Identities = 16/38 (42%), Positives = 20/38 (52%)
Frame = +3
Query: 87 FTFPDFQVVPTLEAYSHLLDSPTAEKTPFTGPGTSLTP 124
F FP +V+ ++ S LLDSPT TPF T P
Sbjct: 72 FLFPS*RVLVVVD--SVLLDSPTTTTTPFAPTCTESNP 179
>TC234236 similar to UP|Q9LW21 (Q9LW21) Arabidopsis thaliana genomic DNA,
chromosome 3, P1 clone: MDJ14, partial (4%)
Length = 586
Score = 27.3 bits (59), Expect = 5.1
Identities = 15/39 (38%), Positives = 19/39 (48%)
Frame = -1
Query: 8 IHRRIGQIKNFVNITGMKKTKSYKFKEVDLVSLRELALK 46
I R QIKN NIT K +YK ++ R+L K
Sbjct: 439 IMRAETQIKNVTNITNEKNKNAYKHSTAYMLGRRDLESK 323
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.323 0.139 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,613,942
Number of Sequences: 63676
Number of extensions: 106094
Number of successful extensions: 514
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 513
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 513
length of query: 208
length of database: 12,639,632
effective HSP length: 93
effective length of query: 115
effective length of database: 6,717,764
effective search space: 772542860
effective search space used: 772542860
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 57 (26.6 bits)
Medicago: description of AC147483.13


