

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147471.10 + phase: 0
(532 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
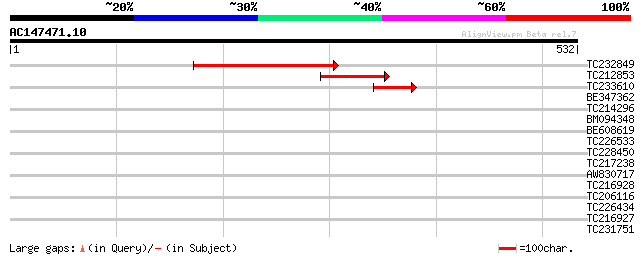
Sequences producing significant alignments: (bits) Value
TC232849 224 1e-58
TC212853 117 2e-26
TC233610 44 2e-04
BE347362 33 0.32
TC214296 similar to UP|Q9VH98 (Q9VH98) CG8348-PA, partial (5%) 32 0.70
BM094348 31 1.2
BE608619 31 1.2
TC226533 similar to GB|AAG53613.1|12407658|AF285832 eukaryotic i... 31 1.6
TC228450 30 2.7
TC217238 30 2.7
AW830717 29 6.0
TC216928 similar to UP|Q9SVA6 (Q9SVA6) GTP-binding-like protein,... 29 6.0
TC206116 UP|CC48_SOYBN (P54774) Cell division cycle protein 48 h... 29 6.0
TC226434 homologue to UP|Q8S3W1 (Q8S3W1) Elongation factor 1-gam... 29 6.0
TC216927 homologue to UP|Q9SVA6 (Q9SVA6) GTP-binding-like protei... 29 6.0
TC231751 homologue to GB|BAB08420.1|9757998|AB025622 cell divisi... 28 7.8
>TC232849
Length = 408
Score = 224 bits (570), Expect = 1e-58
Identities = 112/136 (82%), Positives = 123/136 (90%)
Frame = +1
Query: 173 EERYKKEVQMKHILPDVADKFFLDREGEEKGANILCFDEIQTVDVFAIVALSGILSRLLS 232
EERYK+EVQMK+ILP VAD FFLD E EKGA+ILCFDEIQTVDVFAIVALSGILSRLLS
Sbjct: 1 EERYKQEVQMKNILPAVADMFFLDGEENEKGASILCFDEIQTVDVFAIVALSGILSRLLS 180
Query: 233 SGTIIVATSNRAPKDLNEANMVPEFFQNLLSNLEEHCEKVLVGSEIDYRRFIAQRSENRV 292
SGTIIVATSNRAPKDLNEA M E FQ L+S LEEHCEKVL+GSEIDYRRFIAQ+SEN+V
Sbjct: 181 SGTIIVATSNRAPKDLNEAGMQKEIFQKLVSKLEEHCEKVLIGSEIDYRRFIAQKSENQV 360
Query: 293 NYLWPIERETINKFEK 308
+Y WPIE+E +N+FEK
Sbjct: 361 HYFWPIEKEAMNEFEK 408
>TC212853
Length = 499
Score = 117 bits (292), Expect = 2e-26
Identities = 50/65 (76%), Positives = 60/65 (91%)
Frame = +2
Query: 292 VNYLWPIERETINKFEKKWQDATGRFGGKVISNTISVMFGRTLEVPESCEGVARFTFDYL 351
V+Y WPIE+E +N+FEKKW D TGRFGG++ISNTISVMFGRTLEVP+S +GVARFTF+YL
Sbjct: 80 VHYFWPIEKEAMNEFEKKWHDVTGRFGGRIISNTISVMFGRTLEVPQSFDGVARFTFEYL 259
Query: 352 CGRPL 356
CGRP+
Sbjct: 260 CGRPV 274
>TC233610
Length = 629
Score = 43.9 bits (102), Expect = 2e-04
Identities = 17/40 (42%), Positives = 26/40 (64%)
Frame = +1
Query: 342 GVARFTFDYLCGRPLGAADYIAVAENYHTVFISDIPMMSM 381
G A F F+ +C RPLGAADY + + +HT+ + IP+ +
Sbjct: 16 GCAYFPFEEICDRPLGAADYFGLFKKFHTLVLDGIPIFGL 135
>BE347362
Length = 448
Score = 33.1 bits (74), Expect = 0.32
Identities = 20/60 (33%), Positives = 28/60 (46%), Gaps = 1/60 (1%)
Frame = +1
Query: 92 LYIYGNVGSGKTMLMDMFYSATEGIVKHRR-RYHFHEAMLRINEHMHKTWKKQMEEKPLQ 150
++ YG G+GKT M EG +HR Y E + RI E H T K ++ L+
Sbjct: 166 IFAYGQTGTGKTFTM-------EGTPEHRGVNYRTLEELFRITEERHGTMKYELSVSMLE 324
>TC214296 similar to UP|Q9VH98 (Q9VH98) CG8348-PA, partial (5%)
Length = 1491
Score = 32.0 bits (71), Expect = 0.70
Identities = 21/81 (25%), Positives = 32/81 (38%)
Frame = +2
Query: 12 EKKRENERRRILMDEVEKQQNDKDWWKRLNNKITERWTNSRKRPENVDPGVGKWVSYLKR 71
E++RE ER R E E+++ + +R+ + + P P G VSY
Sbjct: 341 ERERERERERERERERERERERERERERITSHSISMYQQQGSDPTKQSPATGFPVSYSNS 520
Query: 72 EKKLDSLVGRRPTAPPAPKGL 92
+ P PP PK L
Sbjct: 521 TTYSTNEASYAPVPPPQPKPL 583
>BM094348
Length = 414
Score = 31.2 bits (69), Expect = 1.2
Identities = 29/113 (25%), Positives = 47/113 (40%), Gaps = 24/113 (21%)
Frame = -3
Query: 388 SLSYFEQQKKKRSLLFRHGDLSLLLTNYTIIILVYVA---WHPLQLMNYF----KELRRA 440
S S E KKK+ + H D++ + N ++I + +A +HPL L + E RR+
Sbjct: 409 SFSPPEYPKKKKKQVLLHSDIT*KVINLSLIDDMMLA*RNYHPLTLFAFLLVSKSEDRRS 230
Query: 441 HFLTWREAEGSK-----------------LRRDVLAEGNVGSGGTPVGITSIL 476
HF SK L + + + SG T G++SI+
Sbjct: 229 HFFLICFNSSSKSTTVCFNSVSFSLYSSHLSKSFMFSAQISSGDTDTGVSSIV 71
>BE608619
Length = 256
Score = 31.2 bits (69), Expect = 1.2
Identities = 15/59 (25%), Positives = 29/59 (48%)
Frame = +1
Query: 9 SNWEKKRENERRRILMDEVEKQQNDKDWWKRLNNKITERWTNSRKRPENVDPGVGKWVS 67
S WE+K R L ++V K + +K W K+ ++ E ++ E +D +W++
Sbjct: 64 SPWEEKSAVAR---LANKVNKSKRNKLWRKKKRKRVVEMLAKEHEQFEQIDREADEWIA 231
>TC226533 similar to GB|AAG53613.1|12407658|AF285832 eukaryotic initiation
factor 3E subunit {Arabidopsis thaliana;} , partial
(98%)
Length = 1619
Score = 30.8 bits (68), Expect = 1.6
Identities = 25/101 (24%), Positives = 41/101 (39%), Gaps = 18/101 (17%)
Frame = +1
Query: 92 LYIYGNVGSGKTMLMDMF------------------YSATEGIVKHRRRYHFHEAMLRIN 133
L+I+ N +G+T ++D+F Y AT IV RRR F + + I
Sbjct: 745 LFIFFNHDNGRTQIIDLFNQDKYLNAIQTSAPHLLRYLATAFIVNKRRRPQFKDFIKVIQ 924
Query: 134 EHMHKTWKKQMEEKPLQSGISSWIMNLPFDTKAKEWLAAEE 174
+ H + P+ ++ +N FD K+ EE
Sbjct: 925 QEQHS------YKDPITEFLACVYVNYDFDGAQKKMRECEE 1029
>TC228450
Length = 739
Score = 30.0 bits (66), Expect = 2.7
Identities = 16/55 (29%), Positives = 28/55 (50%), Gaps = 3/55 (5%)
Frame = -1
Query: 5 HVNLSNWE---KKRENERRRILMDEVEKQQNDKDWWKRLNNKITERWTNSRKRPE 56
H N+ +WE ++ ENE+ L E++K K W+ + RW + ++R E
Sbjct: 208 HRNVRDWEGLFRRSENEKHLKLKWEMKKAMKMKTNWEWRASASARRWKSRQRRGE 44
>TC217238
Length = 1268
Score = 30.0 bits (66), Expect = 2.7
Identities = 14/37 (37%), Positives = 22/37 (58%), Gaps = 1/37 (2%)
Frame = -2
Query: 116 IVKHRRRYHFHEAMLRINEHM-HKTWKKQMEEKPLQS 151
+VKH +HF +A L+ +H+ H Q+E K LQ+
Sbjct: 385 VVKHNLHFHFPQATLKYYKHLEHVNCATQLEVKSLQT 275
>AW830717
Length = 306
Score = 28.9 bits (63), Expect = 6.0
Identities = 12/29 (41%), Positives = 17/29 (58%), Gaps = 1/29 (3%)
Frame = +1
Query: 132 INEHMHKTWKKQME-EKPLQSGISSWIMN 159
++ H+H+ WK Q E KPL I SW +
Sbjct: 13 VDHHIHRIWKWQSEPNKPLSRLIKSWFFS 99
>TC216928 similar to UP|Q9SVA6 (Q9SVA6) GTP-binding-like protein, partial
(25%)
Length = 354
Score = 28.9 bits (63), Expect = 6.0
Identities = 16/48 (33%), Positives = 24/48 (49%)
Frame = +1
Query: 435 KELRRAHFLTWREAEGSKLRRDVLAEGNVGSGGTPVGITSILSGQEEL 482
K AH L +A+ +KLRR++L + G+GG G SG +
Sbjct: 139 KNKATAHHLGLLKAKLAKLRRELLTPSSKGAGGAGEGFDVTKSGDSRV 282
>TC206116 UP|CC48_SOYBN (P54774) Cell division cycle protein 48 homolog
(Valosin containing protein homolog) (VCP), complete
Length = 2583
Score = 28.9 bits (63), Expect = 6.0
Identities = 14/35 (40%), Positives = 21/35 (60%)
Frame = +1
Query: 71 REKKLDSLVGRRPTAPPAPKGLYIYGNVGSGKTML 105
R +L +G +P PKG+ +YG GSGKT++
Sbjct: 706 RHPQLFKSIGVKP-----PKGILLYGPPGSGKTLI 795
>TC226434 homologue to UP|Q8S3W1 (Q8S3W1) Elongation factor 1-gamma, complete
Length = 1517
Score = 28.9 bits (63), Expect = 6.0
Identities = 14/50 (28%), Positives = 26/50 (52%)
Frame = +1
Query: 397 KKRSLLFRHGDLSLLLTNYTIIILVYVAWHPLQLMNYFKELRRAHFLTWR 446
K +L RH +L + I+ L+ P++L+N+ + + A FL W+
Sbjct: 91 KPTKILTRHSLQPSMLASKLILPLILRWVSPIKLLNFSRSILLARFLCWK 240
>TC216927 homologue to UP|Q9SVA6 (Q9SVA6) GTP-binding-like protein, complete
Length = 1771
Score = 28.9 bits (63), Expect = 6.0
Identities = 16/48 (33%), Positives = 24/48 (49%)
Frame = +2
Query: 435 KELRRAHFLTWREAEGSKLRRDVLAEGNVGSGGTPVGITSILSGQEEL 482
K AH L +A+ +KLRR++L + G+GG G SG +
Sbjct: 266 KNKATAHHLGLLKAKLAKLRRELLTPSSKGAGGAGEGFDVTKSGDSRV 409
>TC231751 homologue to GB|BAB08420.1|9757998|AB025622 cell division protein
FtsH protease-like {Arabidopsis thaliana;} , partial
(30%)
Length = 733
Score = 28.5 bits (62), Expect = 7.8
Identities = 16/40 (40%), Positives = 21/40 (52%)
Frame = +1
Query: 66 VSYLKREKKLDSLVGRRPTAPPAPKGLYIYGNVGSGKTML 105
V YLK K L G+ PKG+ + G G+GKT+L
Sbjct: 139 VEYLKNPAKFTRLGGK------LPKGILLTGPPGTGKTLL 240
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.135 0.402
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 24,042,374
Number of Sequences: 63676
Number of extensions: 333540
Number of successful extensions: 1841
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 1826
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1840
length of query: 532
length of database: 12,639,632
effective HSP length: 102
effective length of query: 430
effective length of database: 6,144,680
effective search space: 2642212400
effective search space used: 2642212400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC147471.10


