

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC147435.1 - phase: 0 /pseudo
(346 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
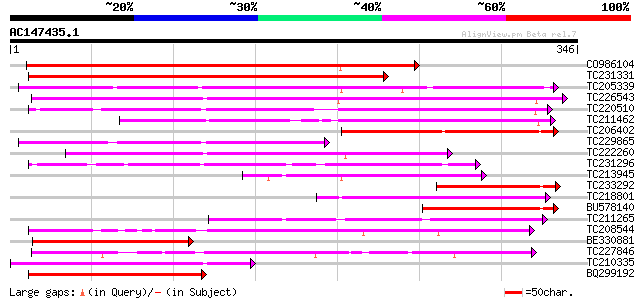
Score E
Sequences producing significant alignments: (bits) Value
CO986104 375 e-104
TC231331 weakly similar to UP|Q8S995 (Q8S995) Glucosyltransferas... 241 3e-64
TC205339 weakly similar to UP|Q6QDB6 (Q6QDB6) UDP-glucose glucos... 229 1e-60
TC226543 weakly similar to UP|Q6VAA9 (Q6VAA9) UDP-glycosyltransf... 208 3e-54
TC220510 similar to UP|Q7XZD0 (Q7XZD0) Isoflavonoid glucosyltran... 194 5e-50
TC211462 similar to UP|Q7XZD0 (Q7XZD0) Isoflavonoid glucosyltran... 168 4e-42
TC206402 similar to UP|Q8S995 (Q8S995) Glucosyltransferase-14, p... 164 5e-41
TC229865 weakly similar to UP|P93789 (P93789) UDP-glucose glucos... 137 7e-33
TC222260 weakly similar to UP|Q7XZD0 (Q7XZD0) Isoflavonoid gluco... 136 2e-32
TC231296 similar to UP|Q7XZD0 (Q7XZD0) Isoflavonoid glucosyltran... 134 8e-32
TC213945 similar to UP|Q6QDB6 (Q6QDB6) UDP-glucose glucosyltrans... 124 8e-29
TC233292 similar to UP|Q8S9A0 (Q8S9A0) Glucosyltransferase-9, pa... 122 3e-28
TC218801 similar to UP|Q8W3P8 (Q8W3P8) ABA-glucosyltransferase, ... 119 2e-27
BU578140 113 1e-25
TC211265 similar to UP|Q9ZWQ5 (Q9ZWQ5) UDP-glycose:flavonoid gly... 113 1e-25
TC208544 weakly similar to PIR|D96766|D96766 protein glucosyltra... 113 1e-25
BE330881 similar to GP|19911201|dbj glucosyltransferase-9 {Vigna... 110 1e-24
TC227846 weakly similar to UP|Q947K4 (Q947K4) Thiohydroximate S-... 103 1e-22
TC210335 98 5e-21
BQ299192 similar to PIR|T03747|T037 glucosyltransferase IS5a (EC... 91 8e-19
>CO986104
Length = 797
Score = 375 bits (963), Expect = e-104
Identities = 181/243 (74%), Positives = 208/243 (85%), Gaps = 3/243 (1%)
Frame = +1
Query: 11 PHFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIV 70
PHFVLFPL+AQGHIIPM+DIA+LLA+RGVIVTIFTTPKNASRF SVLSRAVSSGLQI++V
Sbjct: 64 PHFVLFPLMAQGHIIPMMDIARLLARRGVIVTIFTTPKNASRFNSVLSRAVSSGLQIRLV 243
Query: 71 TLNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIISDFF 130
L+FP+K+AGLPEGCENFDM+ S+DM +FHAI++LQK AEELF+AL PKPSCIISDF
Sbjct: 244 QLHFPSKEAGLPEGCENFDMLTSMDMMYKVFHAISMLQKSAEELFEALIPKPSCIISDFC 423
Query: 131 IPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPGIPDQIQVT 190
IPWT Q+AE H IPRISFHGFSCFCLHC+L + TS I E + SESEYFT+PGIP QIQ T
Sbjct: 424 IPWTAQVAEKHHIPRISFHGFSCFCLHCLLMVHTSNICESITSESEYFTIPGIPGQIQAT 603
Query: 191 KEQIPGILKG---ELKEFGEKMHDAEMKSYGEIINTFEELEKAYVKDYKKEKNGKVWFVG 247
KEQIP ++ E+K FG++M DAEMKSYG IINTFEEL AYV DYKK +N KVW +G
Sbjct: 604 KEQIPMMISNSDEEMKHFGDQMRDAEMKSYGLIINTFEELXXAYVTDYKKVRNDKVWCIG 783
Query: 248 PVS 250
PVS
Sbjct: 784 PVS 792
>TC231331 weakly similar to UP|Q8S995 (Q8S995) Glucosyltransferase-14,
partial (27%)
Length = 713
Score = 241 bits (616), Expect = 3e-64
Identities = 116/221 (52%), Positives = 157/221 (70%), Gaps = 1/221 (0%)
Frame = +1
Query: 12 HFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIVT 71
HFVLFPL++ GH++PM D+A +LAQ +IVT+ TTP NASR + SRA SGL +++V
Sbjct: 46 HFVLFPLMSPGHLLPMTDLATILAQHNIIVTVVTTPHNASRLSETFSRASDSGLNLRLVQ 225
Query: 72 LNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAIT-LLQKPAEELFDALTPKPSCIISDFF 130
L FP++ AG PEGCENFDM+ S+ M +N F A L +PAE++F+ LTPKP+CIISD
Sbjct: 226 LQFPSQDAGFPEGCENFDMLPSMGMGLNFFLAANNFLHEPAEKVFEELTPKPNCIISDVG 405
Query: 131 IPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPGIPDQIQVT 190
+ +T IA IPRISF+G SCFCL K+ TS +LE ++++SEYF +P IPD+I++T
Sbjct: 406 LAYTAHIATKFNIPRISFYGVSCFCLSWQQKLVTSNLLESIETDSEYFLIPDIPDKIEIT 585
Query: 191 KEQIPGILKGELKEFGEKMHDAEMKSYGEIINTFEELEKAY 231
KEQ + EF +KM AE +YG ++N+FEELE AY
Sbjct: 586 KEQTSRPMHENWSEFVDKMAAAEAVTYGVVVNSFEELEPAY 708
>TC205339 weakly similar to UP|Q6QDB6 (Q6QDB6) UDP-glucose
glucosyltransferase, partial (55%)
Length = 1537
Score = 229 bits (585), Expect = 1e-60
Identities = 125/340 (36%), Positives = 192/340 (55%), Gaps = 10/340 (2%)
Frame = +2
Query: 6 NINDVPHFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGL 65
N++ H +LFP QGH+IPM D+A+ RGV TI TTP N + + + +
Sbjct: 14 NMDGELHIMLFPFPGQGHLIPMSDMARAFNGRGVRTTIVTTPLNVATIRGTIGKETET-- 187
Query: 66 QIKIVTLNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCI 125
I+I+T+ FP+ +AGLPEGCEN + + S D+ + AI +L+ P E L L +P C+
Sbjct: 188 DIEILTVKFPSAEAGLPEGCENTESIPSPDLVLTFLKAIRMLEAPLEHLL--LQHRPHCL 361
Query: 126 ISDFFIPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPGIPD 185
I+ F PW A KIPR+ FHG F L ++ + + V S+++ F +P +P
Sbjct: 362 IASAFFPWASHSATKLKIPRLVFHGTGVFALCASECVRLYQPHKNVSSDTDPFIIPHLPG 541
Query: 186 QIQVTKEQIPGILKGE------LKEFGEKMHDAEMKSYGEIINTFEELEKAYVKDYKKE- 238
IQ+T+ +P K + L +++ ++E+ SYG I+N+F ELE+ Y Y K+
Sbjct: 542 DIQMTRLLLPDYAKTDGDGETGLTRVLQEIKESELASYGMIVNSFYELEQVYADYYDKQL 721
Query: 239 ---KNGKVWFVGPVSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCN 295
+ + W++GP+SLCN+D K +RG AS+ + LKWLD + SVVY C GS+ N
Sbjct: 722 LQVQGRRAWYIGPLSLCNQD---KGKRGKQASVDQGDILKWLDSKKANSVVYVCFGSIAN 892
Query: 296 LIPSQLMELALALEATNRPFIWVIREGNKSSEELEKWISE 335
+QL E+A LE + + FIWV+R +K + W+ E
Sbjct: 893 FSETQLREIARGLEDSGQQFIWVVRRSDKDD---KGWLPE 1003
>TC226543 weakly similar to UP|Q6VAA9 (Q6VAA9) UDP-glycosyltransferase 73E1,
partial (21%)
Length = 1237
Score = 208 bits (529), Expect = 3e-54
Identities = 123/336 (36%), Positives = 179/336 (52%), Gaps = 9/336 (2%)
Frame = +2
Query: 14 VLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIVTLN 73
+ P ++ HIIP++D+A+L A V VTI TT NA+ F + S G I+ +N
Sbjct: 56 IFLPFLSTSHIIPLVDMARLFALHDVDVTIITTAHNATVFQKSIDLDASRGRPIRTHVVN 235
Query: 74 FPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIISDFFIPW 133
FP Q GLP G E F++ +M ++ ++LLQ+ E+LF L +P I++D F PW
Sbjct: 236 FPAAQVGLPVGIEAFNVDTPREMTPRIYMGLSLLQQVFEKLFHDL--QPDFIVTDMFHPW 409
Query: 134 TIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPGIPDQIQVTKEQ 193
++ A IPRI FHG S ++ +++ F +PG+PD +++T+ Q
Sbjct: 410 SVDAAAKLGIPRIMFHGASYLARSAAHSVEQYAPHLEAKFDTDKFVLPGLPDNLEMTRLQ 589
Query: 194 IPGILK--GELKEFGEKMHDAEMKSYGEIINTFEELEKAYVKDYKKEKNGKVWFVGPVSL 251
+P L+ + E + +E KSYG + N+F +LE AY + YK K W +GPVSL
Sbjct: 590 LPDWLRSPNQYTELMRTIKQSEKKSYGSLFNSFYDLESAYYEHYKSIMGTKSWGIGPVSL 769
Query: 252 -CNKDGLDKAQRGIIASISEHH-CLKWLDLHQPKSVVYACLGSLCNLIPSQLMELALALE 309
N+D DKA RG E LKWL+ SV+Y GS+ SQL+E+A ALE
Sbjct: 770 WANQDAQDKAARGYAKEEEEKEGWLKWLNSKAESSVLYVSFGSINKFPYSQLVEIARALE 949
Query: 310 ATNRPFIWVIR-----EGNKSSEELEKWISEERNGY 340
+ FIWV+R EG+ EE EK + E GY
Sbjct: 950 DSGHDFIWVVRKNDGGEGDNFLEEFEKRMKESNKGY 1057
>TC220510 similar to UP|Q7XZD0 (Q7XZD0) Isoflavonoid glucosyltransferase,
partial (62%)
Length = 925
Score = 194 bits (493), Expect = 5e-50
Identities = 111/323 (34%), Positives = 175/323 (53%), Gaps = 3/323 (0%)
Frame = +3
Query: 12 HFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIVT 71
+F+ F +A+GH+IP+ D+A L + RG VTI TTP NA +L +++ S +++ T
Sbjct: 18 YFIHF--LAEGHMIPLCDMATLFSTRGHHVTIITTPSNAQ----ILRKSLPSHPLLRLHT 179
Query: 72 LNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIISDFFI 131
+ FP+ + GLP+G EN +D + A +LQ P E+L + P CI++D+
Sbjct: 180 VQFPSHEVGLPDGIENIYADSDLDSLRKVCSATAMLQPPIEDLVEQ--QPPDCIVADYLF 353
Query: 132 PWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPGIPDQIQVTK 191
PW +A+ IPR++F+G S F + + S + S T+ P +
Sbjct: 354 PWVDDLAKKLHIPRLAFNGLSLFTICAIHSSSESSDSTIIQSLPHPITLNATPPK----- 518
Query: 192 EQIPGILKGELKEFGEKMHDAEMKSYGEIINTFEELE-KAYVKDYKKEKNGKVWFVGPVS 250
EL +F E + + E+KSYG I+N+F EL+ + Y + Y+K K W +GP S
Sbjct: 519 ---------ELTKFLETVLETELKSYGLIVNSFTELDGEEYTRYYEKTTGHKAWHLGPAS 671
Query: 251 LCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQLMELALALEA 310
L + +KA+RG + +S H C+ WLD + SVVY C GSLC QL E+A ++A
Sbjct: 672 LIGRTAQEKAERGQKSVVSMHECVAWLDSKRENSVVYICFGSLCYFQDKQLYEIACGIQA 851
Query: 311 TNRPFIWVI--REGNKSSEELEK 331
+ FIWV+ ++G + +E EK
Sbjct: 852 SGHDFIWVVPEKKGKEHEKEEEK 920
>TC211462 similar to UP|Q7XZD0 (Q7XZD0) Isoflavonoid glucosyltransferase,
partial (64%)
Length = 1066
Score = 168 bits (425), Expect = 4e-42
Identities = 96/271 (35%), Positives = 147/271 (53%), Gaps = 5/271 (1%)
Frame = +3
Query: 68 KIVTLNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIIS 127
++ T +FP+++ GLP+G EN V ++ ++ A T L + E F P P CI++
Sbjct: 30 RVHTFDFPSEEVGLPDGVENLSAVTDLEKSYRIYIAATTLLREPIESFVERDP-PDCIVA 206
Query: 128 DFFIPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPGIPDQI 187
DF W +A+ +IP + F+GFS F + M ++ +I + F +P PD +
Sbjct: 207 DFLYCWVEDLAKKLRIPWLDFNGFSLFSICAMESVKKHRIGDGP------FVIPDFPDHV 368
Query: 188 QVTKEQIPGILKGELKEFGEKMHDAEMKSYGEIINTFEELE-KAYVKDYKKEKNGKVWFV 246
+ K P +++EF E + A +KS G IIN F EL+ + Y++ Y+K K W +
Sbjct: 369 TI-KSTPPK----DMREFLEPLLTAALKSNGFIINNFAELDGEEYLRHYEKTTGHKAWHL 533
Query: 247 GPVSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQLMELAL 306
GP SL + ++KA+RG + +S H CL WLD + SVVY GSLC QL E+A
Sbjct: 534 GPASLVRRTEMEKAERGQKSVVSTHECLSWLDSKRVNSVVYVSFGSLCYFPDKQLYEIAC 713
Query: 307 ALEATNRPFIWVIRE----GNKSSEELEKWI 333
+EA+ FIWV+ E +S EE EKW+
Sbjct: 714 GMEASGYEFIWVVPEKKGKEEESEEEKEKWL 806
>TC206402 similar to UP|Q8S995 (Q8S995) Glucosyltransferase-14, partial (59%)
Length = 1152
Score = 164 bits (415), Expect = 5e-41
Identities = 83/133 (62%), Positives = 98/133 (73%)
Frame = +2
Query: 203 KEFGEKMHDAEMKSYGEIINTFEELEKAYVKDYKKEKNGKVWFVGPVSLCNKDGLDKAQR 262
K+ E++ +AEM SYG ++N+FEELE AY YKK + K+W +GPVSL NKD LDKAQR
Sbjct: 14 KQINEEIREAEMSSYGVVMNSFEELEPAYATGYKKIRGDKLWCIGPVSLINKDHLDKAQR 193
Query: 263 GIIASISEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQLMELALALEATNRPFIWVIREG 322
G ASI +KWLD +P +V+YACLGSLCNL QL EL LALEA+ RPFIWVIREG
Sbjct: 194 GT-ASIDVSQHIKWLDCQKPGTVIYACLGSLCNLTTPQLKELGLALEASKRPFIWVIREG 370
Query: 323 NKSSEELEKWISE 335
SEELEKWI E
Sbjct: 371 G-HSEELEKWIKE 406
>TC229865 weakly similar to UP|P93789 (P93789) UDP-glucose
glucosyltransferase , partial (15%)
Length = 647
Score = 137 bits (345), Expect = 7e-33
Identities = 77/191 (40%), Positives = 102/191 (53%), Gaps = 1/191 (0%)
Frame = +1
Query: 6 NINDVPHFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGL 65
N N H + FP A GHIIP ID+A++ A RG+ T+ TTP N + + +A
Sbjct: 91 NENRELHVLFFPFPANGHIIPSIDLARVFASRGIKTTVVTTPLNVPLISRTIGKA----- 255
Query: 66 QIKIVTLNFPT-KQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSC 124
IKI T+ FP+ ++ GLPEGCEN D S D+ M A LL+ P E L P C
Sbjct: 256 NIKIKTIKFPSHEETGLPEGCENSDSALSSDLIMTFLKATVLLRDPLENLMQQ--EHPDC 429
Query: 125 IISDFFIPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPGIP 184
+I+D F PW A IPR+ FHG F ++T K + V S SE F VP +P
Sbjct: 430 VIADMFYPWATDSAAKFGIPRVVFHGMGFFPTCVSACVRTYKPQDNVSSWSEPFAVPELP 609
Query: 185 DQIQVTKEQIP 195
+I +TK Q+P
Sbjct: 610 GEITITKMQLP 642
>TC222260 weakly similar to UP|Q7XZD0 (Q7XZD0) Isoflavonoid
glucosyltransferase, partial (5%)
Length = 717
Score = 136 bits (342), Expect = 2e-32
Identities = 81/239 (33%), Positives = 129/239 (53%), Gaps = 3/239 (1%)
Frame = +1
Query: 35 AQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIVTLNFPTKQAGLPEGCENFDMVDSI 94
A+ GV VTI TP AS F + + + G I+ + FP+ Q GL +G EN ++
Sbjct: 1 AKHGVSVTILNTPAIASTFQNAIDSDFNCGYHIRTQVVPFPSAQVGLIDGLENMKDATTL 180
Query: 95 DMRMNLFHAITLLQKPAEELFDALTPKPSCIISDFFIPWTIQIAENHKIPRISFHGFSCF 154
+M + + + ++ LQ E F L +P CI++D PWT++ AE IPRI F+ S F
Sbjct: 181 EMLVKIGYGLSTLQDEIELRFQDL--QPDCIVTDMMYPWTVESAEKLGIPRIFFYSSSYF 354
Query: 155 CLHCMLKIQTSKILERVDSESEYFTVPGIPDQIQVTKEQIPGILKGELK--EFGEKMHDA 212
I+ + E + S+S FT+PG+P +I++T Q+ ++ + + + E ++
Sbjct: 355 SNCASHFIRKHRPHESLVSDSHKFTIPGLPHRIEMTPSQLADWIRSKTRATAYLEPTFES 534
Query: 213 EMKSYGEIINTFEELEKAYVKDYKKEKNGKVWFVGPVSL-CNKDGLDKAQRGIIASISE 270
E +SYG + N+F ELE Y + +K K W +GPVS NKD +KA RG ++E
Sbjct: 535 ESRSYGALYNSFHELESEYEQLHKNTLGIKSWNIGPVSAWVNKDDGEKANRGHKEDLAE 711
>TC231296 similar to UP|Q7XZD0 (Q7XZD0) Isoflavonoid glucosyltransferase,
partial (41%)
Length = 808
Score = 134 bits (336), Expect = 8e-32
Identities = 85/277 (30%), Positives = 138/277 (49%), Gaps = 1/277 (0%)
Frame = +1
Query: 12 HFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIVT 71
HF+ P ++ GH+IP+ IA L A RG VT+ TTP + + + R S LQ+ +V
Sbjct: 43 HFI--PYLSPGHVIPLCGIATLFASRGQHVTVITTP-----YYAQILRKSSPSLQLHVV- 198
Query: 72 LNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIISDFFI 131
+FP K GLP+G E V + + A LL++P D P CI++D
Sbjct: 199 -DFPAKDVGLPDGVEIKSAVTDLADTAKFYQAAMLLRRPISHFMD--QHPPDCIVADTMY 369
Query: 132 PWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPGIPDQIQVTK 191
W +A N +IPR++F+G+ F M + + L S++ F +P P ++
Sbjct: 370 SWADDVANNLRIPRLAFNGYPLFSGAAMKCVISHPELH---SDTGPFVIPDFPHRV---- 528
Query: 192 EQIPGILKGELKEFGEKMHDAEMKSYGEIINTFEELE-KAYVKDYKKEKNGKVWFVGPVS 250
+P F + + E+KS+G I+N+F EL+ + ++ Y+K K W +GP
Sbjct: 529 -TMPSRPPKMATAFMDHLLKIELKSHGLIVNSFAELDGEECIQHYEKSTGHKAWHLGPAC 705
Query: 251 LCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVY 287
L K ++ ++ + +S++ CL WLD SVVY
Sbjct: 706 LVGKRDQERGEKSV---VSQNECLTWLDPKPTNSVVY 807
>TC213945 similar to UP|Q6QDB6 (Q6QDB6) UDP-glucose glucosyltransferase,
partial (23%)
Length = 475
Score = 124 bits (310), Expect = 8e-29
Identities = 64/156 (41%), Positives = 93/156 (59%), Gaps = 7/156 (4%)
Frame = +2
Query: 143 IPRISFHGFSCFCL---HCMLKIQTSKILERVDSESEYFTVPGIPDQIQVTKEQIPGILK 199
IPR+ FHG S F L CM + + S+S+ F +P P +I++ K +IP K
Sbjct: 14 IPRLVFHGTSFFSLCVTTCMPFYEPHD--KYASSDSDSFLIPNFPGEIRIEKTKIPPYSK 187
Query: 200 GE----LKEFGEKMHDAEMKSYGEIINTFEELEKAYVKDYKKEKNGKVWFVGPVSLCNKD 255
+ L + E+ ++E++SYG ++N+F ELEK Y ++ K W +GP+SLCNKD
Sbjct: 188 SKEKAGLAKLLEEAKESELRSYGVVVNSFYELEKVYADHFRNVLGRKAWHIGPLSLCNKD 367
Query: 256 GLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLG 291
+KA+RG ASI EH CLKWL+ +P SV+Y C G
Sbjct: 368 AEEKARRGKEASIDEHECLKWLNTKKPNSVIYICXG 475
>TC233292 similar to UP|Q8S9A0 (Q8S9A0) Glucosyltransferase-9, partial (46%)
Length = 993
Score = 122 bits (305), Expect = 3e-28
Identities = 59/76 (77%), Positives = 66/76 (86%)
Frame = +1
Query: 261 QRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQLMELALALEATNRPFIWVIR 320
+RG ASI+EHHCLKWLDL + KSVVY C GSLCNLIPSQL+ELALALE T RPF+WVIR
Sbjct: 211 KRGDQASINEHHCLKWLDLQKSKSVVYVCFGSLCNLIPSQLVELALALEDTKRPFVWVIR 390
Query: 321 EGNKSSEELEKWISEE 336
EG+K +ELEKWISEE
Sbjct: 391 EGSK-YQELEKWISEE 435
>TC218801 similar to UP|Q8W3P8 (Q8W3P8) ABA-glucosyltransferase, partial
(60%)
Length = 1281
Score = 119 bits (298), Expect = 2e-27
Identities = 59/144 (40%), Positives = 86/144 (58%), Gaps = 1/144 (0%)
Frame = +1
Query: 188 QVTKEQIPGILKGELKEFGEKMHDAEMKSYGEIINTFEELEKAYVKDYKKEKNGKVWFVG 247
++T+ Q+P L+ +F +++ E KS+G +N+F +LE AY + K + K W +G
Sbjct: 1 EMTRSQLPVFLRTP-SQFPDRVRQLEEKSFGTFVNSFHDLEPAYAEQVKNKWGKKAWIIG 177
Query: 248 PVSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQLMELALA 307
PVSLCN+ DK +RG + +I E CL WL+ +P SV+Y GSL L QL E+A
Sbjct: 178 PVSLCNRTAEDKTERGKLPTIDEEKCLNWLNSKKPNSVLYVSFGSLLRLPSEQLKEIACG 357
Query: 308 LEATNRPFIWVIRE-GNKSSEELE 330
LEA+ + FIWV+R N SE E
Sbjct: 358 LEASEQSFIWVVRNIHNNPSENKE 429
>BU578140
Length = 434
Score = 113 bits (283), Expect = 1e-25
Identities = 54/83 (65%), Positives = 65/83 (78%)
Frame = +1
Query: 253 NKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQLMELALALEATN 312
N++ LDKAQRG AS H C+KWLDL +P SVVY CLGS+CNLIP QL+EL LALEA+
Sbjct: 7 NRNQLDKAQRGNKASSDAHSCMKWLDLQKPNSVVYVCLGSICNLIPLQLIELGLALEASE 186
Query: 313 RPFIWVIREGNKSSEELEKWISE 335
+PFIW RE N+ +EEL KWI+E
Sbjct: 187 KPFIWFFRERNQ-TEELNKWINE 252
>TC211265 similar to UP|Q9ZWQ5 (Q9ZWQ5) UDP-glycose:flavonoid
glycosyltransferase, partial (41%)
Length = 616
Score = 113 bits (282), Expect = 1e-25
Identities = 67/207 (32%), Positives = 98/207 (46%)
Frame = +3
Query: 122 PSCIISDFFIPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVP 181
P I D WT ++ I R+ F+ S F + + I+T E S+S F +P
Sbjct: 21 PDVFIPDILFTWTKDFSQKLSISRLVFNPISIFDVCMIHAIKTHP--EAFASDSGPFLIP 194
Query: 182 GIPDQIQVTKEQIPGILKGELKEFGEKMHDAEMKSYGEIINTFEELEKAYVKDYKKEKNG 241
+P + + + PG E + D E S+G I+N+F +L+ Y + Y+K
Sbjct: 195 DLPHPLTLPVKPSPGFAA-----LTESLLDGEQDSHGVIVNSFADLDAEYTQHYQKLTGR 359
Query: 242 KVWFVGPVSLCNKDGLDKAQRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQL 301
KVW VGP SL + K + S H CL WLD + SV+Y C GSL + QL
Sbjct: 360 KVWHVGPSSLM----VQKTVKSSTVDESRHDCLTWLDSKKESSVLYICFGSLSLISDEQL 527
Query: 302 MELALALEATNRPFIWVIREGNKSSEE 328
++A LE + F+WV+ NK EE
Sbjct: 528 YQIATGLEGSGHCFLWVVHRKNKDGEE 608
>TC208544 weakly similar to PIR|D96766|D96766 protein glucosyltransferase
F2P9.25 [imported] - Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (16%)
Length = 937
Score = 113 bits (282), Expect = 1e-25
Identities = 94/318 (29%), Positives = 146/318 (45%), Gaps = 9/318 (2%)
Frame = +1
Query: 12 HFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIVT 71
H + +P GH+IP++D K L RGV VT+ TP N + +L + S LQ T
Sbjct: 31 HVLAYPFPTSGHVIPLLDFTKTLVSRGVHVTVLVTPYNEA----LLPKNYSPLLQ----T 186
Query: 72 LNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPSCIISDFFI 131
L P Q P +N +V + + + I + A+ + P+ IISDFF+
Sbjct: 187 LLLPEPQ--FPNPKQN-RLVSMVTFMRHHHYPIIMDWAQAQPI------PPAAIISDFFL 339
Query: 132 PWTIQIAENHKIPRISFHGFSCFCLHCMLKI-QTSKILERVDSESEYFTVPGIPDQIQVT 190
WT +A + +PR+ F F L + + + + + + + P +P+
Sbjct: 340 GWTHLLARDLHVPRVVFSPSGAFALSVSYSLWRDAPQNDNPEDPNGVVSFPNLPNSPFYP 519
Query: 191 KEQIPGILKGELKEFGE-KMHDAEM----KSYGEIINTFEELEKAYVKDYKKEK-NGKVW 244
QI + + E K H M S+G +INTF ELE+ Y+ KKE + +V+
Sbjct: 520 WWQITHLFHDTERGGPEWKFHRENMLLNIDSWGVVINTFTELEQVYLNHLKKELGHERVF 699
Query: 245 FVGPVSLCNKDGLDKA--QRGIIASISEHHCLKWLDLHQPKSVVYACLGSLCNLIPSQLM 302
VGPV + +RG +++S H ++WLD SVVY C GS L SQ+
Sbjct: 700 AVGPVLPIQTGSISTKPEERGGNSTVSRHDIMEWLDARDKGSVVYVCFGSRTFLTSSQME 879
Query: 303 ELALALEATNRPFIWVIR 320
L ALE + F+ +R
Sbjct: 880 VLTRALEISGVNFVLSVR 933
>BE330881 similar to GP|19911201|dbj glucosyltransferase-9 {Vigna angularis},
partial (10%)
Length = 328
Score = 110 bits (274), Expect = 1e-24
Identities = 58/98 (59%), Positives = 73/98 (74%)
Frame = +2
Query: 15 LFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIVTLNF 74
L L+AQGHIIPM+DIA+L + G VTIFTTPKNASR+ +VLS AVS LQI ++ L+F
Sbjct: 35 LSALMAQGHIIPMMDIARL*TRSGATVTIFTTPKNASRYNAVLSHAVS*SLQIPLL*LHF 214
Query: 75 PTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAE 112
+K+AGLP G N M+ S+DM HAI++LQKPAE
Sbjct: 215 TSKEAGLP*GSHNCYMLTSMDMMYKALHAISMLQKPAE 328
>TC227846 weakly similar to UP|Q947K4 (Q947K4) Thiohydroximate
S-glucosyltransferase, partial (49%)
Length = 1605
Score = 103 bits (256), Expect = 1e-22
Identities = 91/322 (28%), Positives = 143/322 (44%), Gaps = 14/322 (4%)
Frame = +2
Query: 14 VLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTS--VLSRAVSSGLQIKIVT 71
++ P AQGHI P++ AK LA +GV T+ TT A+ + + A+S G
Sbjct: 2 LVLPYPAQGHINPLVQFAKRLASKGVKATVATTHYTANSINAPNITVEAISDGFD----- 166
Query: 72 LNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTPKPS-CIISDFF 130
QAG + N + + R N ++ L + ++ TP PS CI+ D F
Sbjct: 167 ------QAGFAQTNNNVQLFLA-SFRTNGSRTLSELIRKHQQ-----TPSPSTCIVYDSF 310
Query: 131 IPWTIQIAENHKIPRISFHGFSCFCLHCMLKIQTSKILERVDSESEYFTVPGIPD----- 185
PW + +A+ H I +F S + ++ I V E VPG+P
Sbjct: 311 FPWVLDVAKQHGIYGAAFFTNSAAVCNIFCRLHHGFIQLPVKMEHLPLRVPGLPPLDSRA 490
Query: 186 --QIQVTKEQIPGILKGELKEFGEKMHDAEMKSYGEIINTFEELEKAYVKDYKKEKNGKV 243
E P + +L +F +++A+ +NTFE LE +K + K+
Sbjct: 491 LPSFVRFPESYPAYMAMKLSQFSN-LNNADWM----FVNTFEALESEVLKGLTELFPAKM 655
Query: 244 WFVGP-VSLCNKDGLDKAQRGIIASISE---HHCLKWLDLHQPKSVVYACLGSLCNLIPS 299
+GP V DG K +G AS+ + C WL+ P+SVVY GS+ +L
Sbjct: 656 --IGPMVPSGYLDGRIKGDKGYGASLWKPLTEECSNWLESKPPQSVVYISFGSMVSLTEE 829
Query: 300 QLMELALALEATNRPFIWVIRE 321
Q+ E+A L+ + F+WV+RE
Sbjct: 830 QMEEVAWGLKESGVSFLWVLRE 895
>TC210335
Length = 652
Score = 98.2 bits (243), Expect = 5e-21
Identities = 52/150 (34%), Positives = 82/150 (54%)
Frame = +2
Query: 1 MVLPANINDVPHFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRA 60
+++P + V P ++ H+IP++DIA+L A GV VTI TT A+ F S + R
Sbjct: 68 LIVPGEHDHKLKLVSLPFVSTSHLIPVVDIARLFAIHGVDVTIITTTATAAIFQSSIDRD 247
Query: 61 VSSGLQIKIVTLNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTP 120
G I+ + FP +Q GLPEG E+F+ D+ ++ +T+LQ ++LF L
Sbjct: 248 RDRGHAIRTHVVKFPCEQVGLPEGVESFNSNTPRDLVPKIYQGLTILQDQYQQLFHDL-- 421
Query: 121 KPSCIISDFFIPWTIQIAENHKIPRISFHG 150
+P + +D F PWT+ A IPR+ + G
Sbjct: 422 QPDFLFTDMFYPWTVDAAAKLGIPRLIYVG 511
>BQ299192 similar to PIR|T03747|T037 glucosyltransferase IS5a (EC 2.4.1.-)
salicylate-induced - common tobacco, partial (14%)
Length = 421
Score = 90.9 bits (224), Expect = 8e-19
Identities = 44/109 (40%), Positives = 66/109 (60%)
Frame = +1
Query: 12 HFVLFPLIAQGHIIPMIDIAKLLAQRGVIVTIFTTPKNASRFTSVLSRAVSSGLQIKIVT 71
H FP +A GH+IP +D+AKL A++GV TI TTP N + + ++ ++G +I I T
Sbjct: 85 HIFFFPFLAHGHMIPTVDMAKLFAEKGVKATIITTPLNEPFIYNAIGKSKTNGNKIHIQT 264
Query: 72 LNFPTKQAGLPEGCENFDMVDSIDMRMNLFHAITLLQKPAEELFDALTP 120
+ FP+ +AGL +GCEN + V S ++ F A LQ+P E+L P
Sbjct: 265 IEFPSAEAGLLDGCENTESVPSPELLNPFFMATHFLQEPLEQLLQKQLP 411
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.322 0.139 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,583,637
Number of Sequences: 63676
Number of extensions: 247868
Number of successful extensions: 1573
Number of sequences better than 10.0: 136
Number of HSP's better than 10.0 without gapping: 1512
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1519
length of query: 346
length of database: 12,639,632
effective HSP length: 98
effective length of query: 248
effective length of database: 6,399,384
effective search space: 1587047232
effective search space used: 1587047232
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC147435.1


