

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
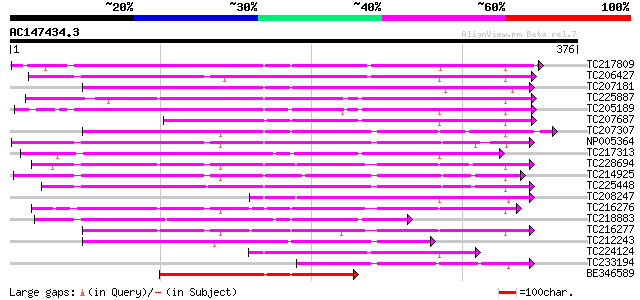
Query= AC147434.3 - phase: 0
(376 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC217809 beta-1,3-glucanase 7 181 5e-46
TC206427 similar to UP|Q8S9I6 (Q8S9I6) AT5g55180/MCO15_13, parti... 173 1e-43
TC207181 168 4e-42
TC225887 similar to UP|Q9M4A9 (Q9M4A9) Beta-1,3 glucanase precur... 160 1e-39
TC205189 similar to UP|Q94G86 (Q94G86) Beta-1,3-glucanase-like p... 159 2e-39
TC207687 143 1e-34
TC207307 homologue to UP|E13B_PHAVU (P23535) Glucan endo-1,3-bet... 135 2e-32
NP005364 beta-1,3-glucanase 134 9e-32
TC217313 weakly similar to UP|Q9LVZ9 (Q9LVZ9) Beta-1,3-glucanase... 132 3e-31
TC228694 UP|Q6S9W0 (Q6S9W0) Endo-1,3-beta-glucanase, complete 132 3e-31
TC214925 UP|E13A_SOYBN (Q03773) Glucan endo-1,3-beta-glucosidase... 131 4e-31
TC225448 similar to UP|Q39900 (Q39900) Beta-1,3-glucanase , part... 127 1e-29
TC208247 similar to GB|BAB01763.1|9280308|AP000603 beta-1,3-gluc... 123 1e-28
TC216276 homologue to UP|E13B_SOYBN (P52395) Glucan endo-1,3-bet... 122 2e-28
TC218883 weakly similar to UP|Q9MAQ2 (Q9MAQ2) CDS, partial (54%) 118 4e-27
TC216277 UP|O49012 (O49012) Beta-1,3-glucanase 3 (Fragment) , co... 118 5e-27
TC212243 UP|O49011 (O49011) Beta-1,3-glucanase 1 (Fragment) , co... 107 9e-24
TC224124 similar to UP|Q8L9D9 (Q8L9D9) Beta-1,3-glucanase-like p... 94 1e-19
TC233194 similar to GB|CAB79832.1|7270016|ATCHRIV74 1, 3-beta-gl... 91 6e-19
BE346589 89 2e-18
>TC217809 beta-1,3-glucanase 7
Length = 1867
Score = 181 bits (459), Expect = 5e-46
Identities = 120/362 (33%), Positives = 199/362 (54%), Gaps = 9/362 (2%)
Frame = +1
Query: 2 FHHSTFMISLTTFSLYLLLSL--LPTTTTSLPTIGVTYSSTTRQDSPPPPSPDRITTAMQ 59
F H+T M T FSL SL L S +GV Y + + P+PD++ +Q
Sbjct: 163 FTHNTAMP--TGFSLIFAASLFLLLLDCCSGSFVGVCYG----RSADDLPTPDKVAQLVQ 324
Query: 60 NLKLTHLRLEEPDPSIIRSLLYTNISLFLTIPNYLVTPIATNRSIARAWIYTHVLPFYPR 119
K+ ++R+ + + ++++ T I L + +PN + + +S A +W+ VLP+YP
Sbjct: 325 LHKIKYVRIYDSNIQVLKAFANTGIELMIGVPNSDLLSFSQFQSNADSWLKNSVLPYYPA 504
Query: 120 AKITTISVGNAFTDVYPESINNLLPAISNVHISLRDLGI-RKISVSTSFSFVTAVTSPFP 178
KI I+VG T+ + + ++PA++NV +L+ LG+ +KI VS++ S + ++ FP
Sbjct: 505 TKIAYITVGAEVTESPNNASSFVVPAMTNVLTALKKLGLHKKIKVSSTHS-LGVLSRSFP 681
Query: 179 PSSAAFQEIPGVNLIGPLLQFLSDTNSSFLINLYPYNLYR-LRPEIPLGIALFQEHPFNF 237
PS+ AF + + P+L+FL++ + F+I++YPY +R R ++ L ALF
Sbjct: 682 PSAGAFNS-SHAHFLKPMLEFLAENQAPFMIDIYPYYAHRDSRSKVSLDYALFDASSEVI 858
Query: 238 RDDFTTGVRYKNLFDIMVDAVVSAMALEGYETIPVIVTETGWPSSGS--ELDANSAYAEI 295
D TG+ Y N+FD +DA+ A+ + TI V+VTETGWPS GS E A A+
Sbjct: 859 --DPNTGLLYTNMFDAQIDAIYFALMALNFRTIKVMVTETGWPSKGSPKETAATPDNAQT 1032
Query: 296 YLKGLVKHLKSGAGTPLLKDGVKGVYLYELFD---KEGAGNGRNWGILYPNGSTKYDKID 352
Y L++H+ + GTP VY++ LF+ K G + RNWG+ YP+ ++ Y +D
Sbjct: 1033YNTNLIRHVINNTGTPAKPGEELDVYIFSLFNENRKPGLESERNWGLFYPDQTSVY-SLD 1209
Query: 353 FS 354
F+
Sbjct: 1210FT 1215
>TC206427 similar to UP|Q8S9I6 (Q8S9I6) AT5g55180/MCO15_13, partial (89%)
Length = 1660
Score = 173 bits (438), Expect = 1e-43
Identities = 108/346 (31%), Positives = 186/346 (53%), Gaps = 9/346 (2%)
Frame = +3
Query: 13 TFSLYLLLSLLPTTTTSLPTIGVTYSSTTRQDSPPPPSPDRITTAMQNLKLTHLRLEEPD 72
TFS + +L ++++ +IG+ Y P+P ++ +++ L ++L + D
Sbjct: 30 TFSFFFILISYISSSSEAGSIGINYGRIAND----LPTPAKVVELLKSQGLNRVKLYDTD 197
Query: 73 PSIIRSLLYTNISLFLTIPNYLVTPIATNRSIARAWIYTHVLPFYPRAKITTISVGNAFT 132
+++ + + + + + +PN L+ A +S AW+ ++ +YP +I I+VGN
Sbjct: 198 ATVLTAFANSGMKVVVAMPNELLANAAAEQSFTDAWVQANISSYYPATQIEAIAVGN--- 368
Query: 133 DVYPESINN---LLPAISNVHISLRDLGIRKISVSTSFSFVTAVTSPFPPSSAAFQEIPG 189
+V+ + N L+PA+ NVH SL + K +S ++A+ + FP SS +F+
Sbjct: 369 EVFVDPNNTTKFLVPAMKNVHASLVKYSLDKNIKISSPIALSALQNSFPASSGSFKTELL 548
Query: 190 VNLIGPLLQFLSDTNSSFLINLYPYNLYRLRPE-IPLGIALFQEHPFNFRDDFTTGVRYK 248
+I P+L L T S ++N YP+ Y + I L ALF+E+P D G++Y
Sbjct: 549 EPVIKPMLDLLRQTGSYLMVNAYPFFAYAANSDKISLDYALFKENPGVV--DSGNGLKYT 722
Query: 249 NLFDIMVDAVVSAMALEGYETIPVIVTETGWPSSG--SELDANSAYAEIYLKGLVKHLKS 306
NLFD +DAV +AM+ Y+ + + V+ETGWPS+G +E+ A+ A Y LVK + S
Sbjct: 723 NLFDAQIDAVFAAMSALKYDDVKIAVSETGWPSAGDSNEIGASPDNAASYNGNLVKRVLS 902
Query: 307 GAGTPLLKDGVKGVYLYELFD---KEGAGNGRNWGILYPNGSTKYD 349
G+GTPL ++ V+L+ LF+ K G + RN+G+ YP YD
Sbjct: 903 GSGTPLKQNESLDVFLFALFNENQKTGPTSERNYGLFYPTEKKVYD 1040
>TC207181
Length = 1691
Score = 168 bits (425), Expect = 4e-42
Identities = 105/308 (34%), Positives = 171/308 (55%), Gaps = 8/308 (2%)
Frame = +3
Query: 49 PSPDRITTAMQNLKLTHLRLEEPDPSIIRSLLYTNISLFLTIPNYLVTPIATNRSIARAW 108
PS + +Q K+TH+R+ + + I+++L T I + +++PN + I ++ S A +W
Sbjct: 21 PSASDLVAFLQLQKITHVRIYDANSDILKTLSGTKIRVIISVPNNQLLAIGSSNSTAASW 200
Query: 109 IYTHVLPFYPRAKITTISVGNAFTDVYPESINNLLPAISNVHISLRDLGI-RKISVSTSF 167
I +V+ +YP+ I+ ISVG+ P S +LPA+ +++ +L + ++I VST
Sbjct: 201 IDRNVVAYYPQTLISGISVGDEVLTTVPSSAPLILPAVESLYNALVASNLHQQIKVSTPH 380
Query: 168 SFVTAVTSPFPPSSAAFQEIPGVNLIGPLLQFLSDTNSSFLINLYPYNLY-RLRPEIPLG 226
+ + + PFPPS A F + V++I PLLQFLS T S ++NLYPY ++ + + +PL
Sbjct: 381 A-ASIILDPFPPSQAYFNQ-SLVSVILPLLQFLSRTGSPLMMNLYPYYVFMQNKGVVPLD 554
Query: 227 IALFQE-HPFNFRDDFTTGVRYKNLFDIMVDAVVSAMALEGYETIPVIVTETGWPSSGSE 285
ALF+ P D T + Y N+ D MVDA +M + V+VTETGWP+ G
Sbjct: 555 NALFKPLTPNKEMVDPNTLLHYTNVLDAMVDAAYFSMKNLNITDVAVLVTETGWPAKGDS 734
Query: 286 LD--ANSAYAEIYLKGLVKHLKSGAGTPLLKDGVKGVYLYELFDKEGAG---NGRNWGIL 340
+ A A+ Y L++H+ GTPL + V++YELF+++ + NWG+
Sbjct: 735 KEPYATKDNADTYNSNLIRHVFDRTGTPLHPETTSSVFIYELFNEDLRAPPVSEANWGLF 914
Query: 341 YPNGSTKY 348
Y N S Y
Sbjct: 915 YGNTSPAY 938
>TC225887 similar to UP|Q9M4A9 (Q9M4A9) Beta-1,3 glucanase precursor ,
partial (95%)
Length = 1692
Score = 160 bits (404), Expect = 1e-39
Identities = 117/349 (33%), Positives = 181/349 (51%), Gaps = 10/349 (2%)
Frame = +3
Query: 11 LTTFSLYLLLSLLPTTTTSLPTIGVTYSSTTRQDSPPPPSPDRITTAMQNLKLT-----H 65
+ T S +LLL LL ++ G+ + T D+ PPP+ A+ N T
Sbjct: 72 MATTSAFLLLPLLLLLHLAIAAHGIGINYGTLGDNLPPPA------AVANFLKTKTTIDR 233
Query: 66 LRLEEPDPSIIRSLLYTNISLFLTIPNYLVTPIATNRSIARAWIYTHVLPFYPRAKITTI 125
+++ + +P I+R+ + IS+ +T PN + + T AR W+ TH+ PF+P+ KI I
Sbjct: 234 VKIYDVNPDILRAFAGSGISVTVTAPNGDIAAL-TKIDSARQWVATHIKPFHPQTKINYI 410
Query: 126 SVGNAFTDVYPES-INNLLPAISNVHISLRDLGIRKISVSTSFSFVTAVTSPFPPSSAAF 184
VG+ + I L+PA+ +H +L GI I V+T+ S + + S PPS F
Sbjct: 411 LVGSEVLHWGDTNMIRGLVPAMRTLHSALLAEGITDIKVTTAHS-LAIMRSSIPPSMGRF 587
Query: 185 QEIPGVNLIGPLLQFLSDTNSSFLINLYPYNLYRLRPEIPLGIALFQEHPFNFRDDFTTG 244
+ +++GP+L+FL +T + ++N YPY Y + + LF+ P D T
Sbjct: 588 RPGYAKHVLGPMLKFLRETRTPLMVNPYPYFGYNGKN---VNFLLFR--PNRGLYDRYTK 752
Query: 245 VRYKNLFDIMVDAVVSAMALEGYETIPVIVTETGWPSSGSELDANS-AYAEIYLKGLVKH 303
Y N FD ++DAV SAM GY + + V ETGWPS DA S A A+ + + LVKH
Sbjct: 753 RSYTNQFDALMDAVHSAMNALGYGDVDIAVGETGWPSVCDGWDACSVANAQSFNRELVKH 932
Query: 304 LKSGAGTPLLKDGVKGVYLYELFD---KEGAGNGRNWGILYPNGSTKYD 349
L +G GTPL+ + Y++ LF+ K G RNWG+ P+ + YD
Sbjct: 933 LATGKGTPLMPNRSFETYIFALFNENQKPGPIAERNWGLFQPDFTPVYD 1079
>TC205189 similar to UP|Q94G86 (Q94G86) Beta-1,3-glucanase-like protein,
partial (91%)
Length = 1715
Score = 159 bits (402), Expect = 2e-39
Identities = 119/356 (33%), Positives = 188/356 (52%), Gaps = 10/356 (2%)
Frame = +1
Query: 4 HSTFMISLTTFSLYLLLSLLPTTTTSLPTIGVTYSSTTRQDSPPPPSPDRITTAMQNLKL 63
+S+ ++SL+ LL+SL + S IGV Y Q + P+P+ +++ +
Sbjct: 136 NSSILLSLSL----LLISLSLAESQSF--IGVNYG----QVADNLPAPEDTANLLKSTTI 285
Query: 64 THLRLEEPDPSIIRSLLYTNISLFLTIPNYLVTPIATNRSIARAWIYTHVLPFYPRAKIT 123
+RL DP+II++L + I + + N + +A + + A W+ +VLP+YP + IT
Sbjct: 286 GKVRLYGADPAIIKALANSGIGIVIGAANGDIPSLAADPNAATQWVNANVLPYYPASNIT 465
Query: 124 TISVGNAFTDVYPESI-NNLLPAISNVHISLRDLGI-RKISVSTSFSFVTAVTSPFPPSS 181
I+VGN + + + + L+PA+ NV +L + KI VST S + +T PPSS
Sbjct: 466 LITVGNEILTLADQGLKSQLVPAMRNVQNALGAASLGGKIKVSTVHS-MAVLTQSDPPSS 642
Query: 182 AAFQEIPGV-NLIGPLLQFLSDTNSSFLINLYPYNLYRL--RPEIPLGIALFQEHPFNFR 238
F P + + + LL L D S F IN YP+ Y+ RPE L LFQ P + R
Sbjct: 643 GLFN--PALQDTLKQLLALLKDNKSPFTINPYPFFAYQSDPRPE-TLAFCLFQ--PNSGR 807
Query: 239 DDFTTGVRYKNLFDIMVDAVVSAMALEGYETIPVIVTETGWPSSG--SELDANSAYAEIY 296
D G Y N+FD VDAV SA++ G++ + ++V ETGWPS G +E+ + A+ Y
Sbjct: 808 VDSGNGKLYTNMFDAQVDAVHSALSAMGFQDVEIVVAETGWPSRGDSNEVGPSVENAKAY 987
Query: 297 LKGLVKHLKSGAGTPLLKDGVKGVYLYELFD---KEGAGNGRNWGILYPNGSTKYD 349
L+ HL+S GTPL+ Y++ L+D K G G+ R +G+ + + YD
Sbjct: 988 NGNLIAHLRSLVGTPLMPGKSVDTYIFALYDEDLKPGPGSERAFGMFKTDRTVLYD 1155
>TC207687
Length = 1380
Score = 143 bits (361), Expect = 1e-34
Identities = 91/255 (35%), Positives = 145/255 (56%), Gaps = 8/255 (3%)
Frame = +1
Query: 103 SIARAWIYTHVLPFYPRAKITTISVGNAFTDVYPESIN-NLLPAISNVHISLRDLGI-RK 160
S A++W+ HV P+ + +IT I+VGN + + NLLPA+ +V+ +L +LG+ ++
Sbjct: 103 SKAQSWVQQHVQPYISQTRITCITVGNEVFNYNDTQLTANLLPAMQSVYNALVNLGLAQQ 282
Query: 161 ISVSTSFSFVTAVTSPFPPSSAAFQEIPGVNLIGPLLQFLSDTNSSFLINLYPYNLYRLR 220
++V+T+ SF + + FPPSS AF++ + I PLL F + S FLIN YP+ Y+
Sbjct: 283 VTVTTAHSF-NILANSFPPSSGAFRQ-DLIQYIQPLLSFHAQIKSPFLINAYPFFAYKDN 456
Query: 221 P-EIPLGIALFQEHPFNFRDDFTTGVRYKNLFDIMVDAVVSAMALEGYETIPVIVTETGW 279
P +I L LFQ P D T + Y N+ +DAV +A+ G+ + V ++ETGW
Sbjct: 457 PNQISLNYVLFQ--PNQGATDPNTNLHYDNMLYAQIDAVYAAIKALGHTDVEVRISETGW 630
Query: 280 PSSG--SELDANSAYAEIYLKGLVKHLKSGAGTPLLKDGVKGVYLYELFD---KEGAGNG 334
PS G E+ A AEIY L+K ++ GTP ++++ LF+ K G +
Sbjct: 631 PSKGDPDEVGATPQNAEIYNSNLLKRIEQKQGTPANPSVPIDIFVFALFNENLKPGPVSE 810
Query: 335 RNWGILYPNGSTKYD 349
RN+G+ YP+G+ Y+
Sbjct: 811 RNYGLYYPDGTPVYN 855
>TC207307 homologue to UP|E13B_PHAVU (P23535) Glucan
endo-1,3-beta-glucosidase, basic isoform
((1->3)-beta-glucan endohydrolase)
((1->3)-beta-glucanase) (Beta-1,3-endoglucanase) ,
partial (91%)
Length = 1539
Score = 135 bits (341), Expect = 2e-32
Identities = 101/321 (31%), Positives = 166/321 (51%), Gaps = 6/321 (1%)
Frame = +2
Query: 49 PSPDRITTAMQNLKLTHLRLEEPDPSIIRSLLYTNISLFLTIPNYLVTPIATNRSIARAW 108
PS + + ++ + +RL +P+ + + +L + I L L +PN + +ATN +R W
Sbjct: 380 PSANDVIGLYRSNNIKRMRLYDPNQAALEALRNSGIELILGVPNSDLQGLATNPDTSRQW 559
Query: 109 IYTHVLPFYPRAKITTISVGNAFTDVYPES--INNLLPAISNVHISLRDLGIR-KISVST 165
+ +VL F+P KI ++VGN + V S +LPAI NV+ ++R G+ +I VST
Sbjct: 560 VQKNVLNFWPSVKIKYVAVGNEVSPVGGSSSVAQYVLPAIQNVYQAIRAQGLHDQIKVST 739
Query: 166 SFSFVTAVTSPFPPSSAAFQEIPGVNLIGPLLQFLSDTNSSFLINLYPYNLYRLRP-EIP 224
S +T + + FPPS +F+ + + P++ +L N+ L+N+YPY Y P +I
Sbjct: 740 SID-MTLIGNSFPPSQGSFRG-DVRSYLDPIIGYLVYANAPLLVNVYPYFSYTGNPRDIS 913
Query: 225 LGIALFQEHPFNFRDDFTTGVRYKNLFDIMVDAVVSAMALEGYETIPVIVTETGWPSSGS 284
L ALF D Y+NLFD M+D+V +A+ + V+V+E+GWPS G
Sbjct: 914 LPYALFTAPNVVVWDG---QYGYQNLFDAMLDSVHAAIDNTKIGYVEVVVSESGWPSDGG 1084
Query: 285 ELDANSAYAEIYLKGLVKHLKSGAGTPLLKDGVKGVYLYELFD--KEGAGNGRNWGILYP 342
A A +YL LV+ ++ G+P Y++ +FD ++ +++G+ P
Sbjct: 1085-FAATYDNARVYLDNLVR--RANRGSPRRPSKPTETYIFAMFDENQKNPEIEKHFGLFNP 1255
Query: 343 NGSTKYDKIDFSSGWKRLVNV 363
N KY F G KRL V
Sbjct: 1256NKQKKY---PFGFGGKRLGKV 1309
>NP005364 beta-1,3-glucanase
Length = 1047
Score = 134 bits (336), Expect = 9e-32
Identities = 101/358 (28%), Positives = 172/358 (47%), Gaps = 11/358 (3%)
Frame = +1
Query: 2 FHHSTFMISLTTFSLYLLLSLLPTTTTSLPTIGVTYSSTTRQDSPPPPSPDRITTAMQNL 61
+H S S+T+ + +L L+ T + GV Y PSP + +
Sbjct: 10 YHLSGKSSSITSIAFLFILLLITNTGKAGAQSGVCYGRIGNN----LPSPQEVVALFKQY 177
Query: 62 KLTHLRLEEPDPSIIRSLLYTNISLFLTIPNYLVTPIATNRSIARAWIYTHVLPFYPRAK 121
+R+ +P ++ +L +NI L L IPN + +A ++ A W+ ++ + +
Sbjct: 178 DFRRMRIYDPSQEVLEALRGSNIELLLDIPNDNLQNLAFSQDNANKWLQDNIKNYANNVR 357
Query: 122 ITTISVGNAFTDVYPES--INNLLPAISNVHISLRDLGI-RKISVSTSFSFVTAVTSPFP 178
ISVGN +V PE L+PA+ N+ ++ + G+ +I VST+ A+ +P
Sbjct: 358 FRYISVGN---EVKPEHSFAQFLVPAMQNIQRAISNAGLGNQIKVSTAIE-TGALADSYP 525
Query: 179 PSSAAFQEIPGVNLIGPLLQFLSDTNSSFLINLYPYNLYRLRPE-IPLGIALFQEHPFNF 237
PS +F+ + +++ L + N+ L+N+YPY Y P I L ALF+
Sbjct: 526 PSMGSFRSDYRTAYLDGVIRHLVNNNTPLLVNVYPYFAYINDPRNISLDYALFRSPSVVV 705
Query: 238 RDDFTTGVRYKNLFDIMVDAVVSAMALEGYETIPVIVTETGWPSSGSELDANSAYAEIYL 297
+D + Y+NLFD MVDAV +A+ G ++ ++V+E+GWPSSG + A Y
Sbjct: 706 QDG---SLGYRNLFDAMVDAVYAALEKAGGGSVSIVVSESGWPSSGGTA-TSLDNARTYN 873
Query: 298 KGLVKHLKSG-----AGTPLLKDGVKGVYLYELFDK--EGAGNGRNWGILYPNGSTKY 348
LV+++K G AG PL Y++ +F++ + + WG+ PN KY
Sbjct: 874 TNLVRNVKQGTPKRPAGRPL------ETYVFAMFNENHKQPEYEKFWGVFLPNKQPKY 1029
>TC217313 weakly similar to UP|Q9LVZ9 (Q9LVZ9) Beta-1,3-glucanase-like
protein, partial (67%)
Length = 1142
Score = 132 bits (332), Expect = 3e-31
Identities = 97/329 (29%), Positives = 160/329 (48%), Gaps = 8/329 (2%)
Frame = +2
Query: 8 MISLTTFSLYLLLSLLPTTTTSL--PTIGVTYSSTTRQDSPPPPSPDRITTAMQNLKLTH 65
M L F + +L P + T G+ Y PSPD + T ++ K+ +
Sbjct: 155 MTILRFFCPFFILFFTPIASVQAFTGTYGINYGRIANNI----PSPDEVVTLLRAAKIRN 322
Query: 66 LRLEEPDPSIIRSLLYTNISLFLTIPNYLVTPIATNRSIARAWIYTHVLPFYPRAKITTI 125
+R+ + D S++++ T + + + +PN + +++N A W+ +V F P +I I
Sbjct: 323 VRIYDADHSVLKAFSGTGLEIVVGLPNGQLQDMSSNPDHALNWVKENVQSFLPDTRIRGI 502
Query: 126 SVGNAFTDVYPESI-NNLLPAISNVHISLRDLGI-RKISVSTSFSFVTAVTSPFPPSSAA 183
+VGN S+ LL A+ N++ + + L + + + +ST+ SF S +PPSS
Sbjct: 503 AVGNEVLGGTDYSLWGVLLGAVKNIYNATKKLHLDQLVQISTANSFAVFAVS-YPPSSGK 679
Query: 184 FQEIPGVN-LIGPLLQFLSDTNSSFLINLYPYNLYRLRPE-IPLGIALFQEHPFNFRDDF 241
F VN + PLL+F S F +N YP+ Y PE I + ALF+ P D
Sbjct: 680 FDN--NVNQYMKPLLEFFQQIGSPFCLNAYPFLAYAGDPEHIDINYALFE--PTKGIYDP 847
Query: 242 TTGVRYKNLFDIMVDAVVSAMALEGYETIPVIVTETGWPSSG--SELDANSAYAEIYLKG 299
+ Y N+ D +DA +A+ G++ + VI+TETGW S+G +E AN+ A Y
Sbjct: 848 MYHLHYDNMLDAQIDAAYAALEDAGFDKMEVIITETGWASNGDQTEAGANATNARTYNHN 1027
Query: 300 LVKHLKSGAGTPLLKDGVKGVYLYELFDK 328
L + L GTP V Y++ LF++
Sbjct: 1028LRRRLAKRKGTPHRPKNVVKAYIFALFNE 1114
>TC228694 UP|Q6S9W0 (Q6S9W0) Endo-1,3-beta-glucanase, complete
Length = 1251
Score = 132 bits (331), Expect = 3e-31
Identities = 98/344 (28%), Positives = 170/344 (48%), Gaps = 10/344 (2%)
Frame = +3
Query: 15 SLYLLLSLLPTTT---TSLPTIGVTYSSTTRQDSPPPPSPDRITTAMQNLKLTHLRLEEP 71
++ LLL +L +T T ++GV Y P+ + ++ ++ +RL P
Sbjct: 93 AILLLLGILSSTGVEFTGAQSVGVCYGGNGNN----LPTKQAVVDLYKSNRIGKIRLYYP 260
Query: 72 DPSIIRSLLYTNISLFLTIPNYLVTPIATNRSIARAWIYTHVLPFYPRAKITTISVGNAF 131
D ++++L +NI + L +PN + + TN A W+ +V + K I+VGN
Sbjct: 261 DEGVLQALRGSNIEVILGVPNDQLQSL-TNAGAATNWVNKYVKAYSQNVKFKYIAVGN-- 431
Query: 132 TDVYPES--INNLLPAISNVHISLRDLGIR-KISVSTSFSFVTAVTSPFPPSSAAFQEIP 188
+++P ++LPA+ N+ ++ ++ ++ VST+ T + + +PP F
Sbjct: 432 -EIHPGDSLAGSVLPALENIQKAISAANLQGQMKVSTAID-TTLLGNSYPPKDGVFSSSA 605
Query: 189 GVNLIGPLLQFLSDTNSSFLINLYPYNLY-RLRPEIPLGIALFQEHPFNFRDDFTTGVRY 247
+ I P++ FL+ + L N+YPY Y + I L ALF +H N V Y
Sbjct: 606 S-SYIRPIVNFLARNGAPLLANVYPYFAYVNNQQSIGLDYALFTKHGNN-------EVGY 761
Query: 248 KNLFDIMVDAVVSAMALEGYETIPVIVTETGWPSSGSELDANSAYAEIYLKGLVKHLKSG 307
+NLFD ++D++ +A+ G + V+V+E+GWPS G + A A Y + L+ H K
Sbjct: 762 QNLFDALLDSLYAALEKVGAPNVKVVVSESGWPSEGG-VGATVQNAGTYYRNLINHAK-- 932
Query: 308 AGTPLLKDGVKGVYLYELFD---KEGAGNGRNWGILYPNGSTKY 348
GTP G YL+ +FD K+G R++G+ P+ S KY
Sbjct: 933 GGTPKRPSGPIETYLFAMFDENQKDGPEIERHFGLFRPDKSPKY 1064
>TC214925 UP|E13A_SOYBN (Q03773) Glucan endo-1,3-beta-glucosidase precursor
((1->3)-beta-glucan endohydrolase)
((1->3)-beta-glucanase) (Beta-1,3-endoglucanase) ,
complete
Length = 1299
Score = 131 bits (330), Expect = 4e-31
Identities = 98/347 (28%), Positives = 171/347 (49%), Gaps = 7/347 (2%)
Frame = +1
Query: 3 HHSTFMISLTTFSLYLLLSLLPTTTTSLPTIGVTYSSTTRQDSPPPPSPDRITTAMQNLK 62
+HS+ S T +L + L+ T T+ GV Y P+P +
Sbjct: 79 YHSSGKSSSMTAIAFLFILLITYTGTTDAQSGVCYGRLGNN----LPTPQEVVALYNQAN 246
Query: 63 LTHLRLEEPDPSIIRSLLYTNISLFLTIPNYLVTPIATNRSIARAWIYTHVLPFYPRAKI 122
+ +R+ P P ++ +L +NI L L IPN + +A+++ A W+ ++ + +
Sbjct: 247 IRRMRIYGPSPEVLEALRGSNIELLLDIPNDNLRNLASSQDNANKWVQDNIKNYANNVRF 426
Query: 123 TTISVGNAFTDVYPES--INNLLPAISNVHISLRDLGI-RKISVSTSFSFVTAVTSPFPP 179
+SVGN +V PE L+PA+ N+ ++ + G+ ++ VST+ A+ FPP
Sbjct: 427 RYVSVGN---EVKPEHSFAQFLVPALENIQRAISNAGLGNQVKVSTAID-TGALAESFPP 594
Query: 180 SSAAFQ-EIPGVNLIGPLLQFLSDTNSSFLINLYPYNLYRLRP-EIPLGIALFQEHPFNF 237
S +F+ + G L G +++FL + N+ ++N+Y Y Y P +I L ALF+
Sbjct: 595 SKGSFKSDYRGAYLDG-VIRFLVNNNAPLMVNVYSYFAYTANPKDISLDYALFRSPSVVV 771
Query: 238 RDDFTTGVRYKNLFDIMVDAVVSAMALEGYETIPVIVTETGWPSSGSELDANSAYAEIYL 297
+D + Y+NLFD VDAV +A+ G ++ ++V+E+GWPSSG + A Y
Sbjct: 772 QDG---SLGYRNLFDASVDAVYAALEKAGGGSLNIVVSESGWPSSGGTA-TSLDNARTYN 939
Query: 298 KGLVKHLKSGAGTPLLKDGVKGVYLYELFD--KEGAGNGRNWGILYP 342
LV+++K GTP Y++ +FD ++ + WG+ P
Sbjct: 940 TNLVRNVKQ--GTPKRPGAPLETYVFAMFDENQKQPEFEKFWGLFSP 1074
>TC225448 similar to UP|Q39900 (Q39900) Beta-1,3-glucanase , partial (93%)
Length = 1098
Score = 127 bits (318), Expect = 1e-29
Identities = 91/331 (27%), Positives = 161/331 (48%), Gaps = 4/331 (1%)
Frame = +2
Query: 22 LLPTTTTSLPTIGVTYSSTTRQDSPPPPSPDRITTAMQNLKLTHLRLEEPDPSIIRSLLY 81
L+ T T+ GV Y PSP + + +R+ +P ++++L
Sbjct: 11 LITNTGTTGAQSGVCYGRVGNN----LPSPQEVVALYKQYDFRRMRIYDPSQQVLQALRG 178
Query: 82 TNISLFLTIPNYLVTPIATNRSIARAWIYTHVLPFYPRAKITTISVGNAFTDVYPESINN 141
+NI L L +PN + +A+++ A W+ +V + + ISVGN +
Sbjct: 179 SNIELLLDLPNVNLQSVASSQDNANRWVQDNVRNYANNVRFRYISVGNE-VKPWDSFARF 355
Query: 142 LLPAISNVHISLRDLGI-RKISVSTSFSFVTAVTSPFPPSSAAFQEIPGVNLIGPLLQFL 200
++PAI N+ ++ G+ +I VST+ A+ +PPS +F+ + + +++ L
Sbjct: 356 VVPAIQNIQRAVSAAGLGNQIKVSTAIE-TGALAESYPPSRGSFRSDYLTSYLDGVIRHL 532
Query: 201 SDTNSSFLINLYPYNLYRLRP-EIPLGIALFQEHPFNFRDDFTTGVRYKNLFDIMVDAVV 259
+ N+ L+N+YPY Y P +I L ALF+ +D + Y+NLF+ MVDAV
Sbjct: 533 VNNNAPLLVNVYPYFAYIGNPRDISLDYALFRSPSVVVQDG---SLGYRNLFNAMVDAVY 703
Query: 260 SAMALEGYETIPVIVTETGWPSSGSELDANSAYAEIYLKGLVKHLKSGAGTPLLKDGVKG 319
+A+ G ++ ++V+E+GWPSSG + A Y LV+++K GTP
Sbjct: 704 AALEKAGGGSLNIVVSESGWPSSGGTA-TSLDNARTYNTNLVRNVKQ--GTPKRPGRPLE 874
Query: 320 VYLYELFD--KEGAGNGRNWGILYPNGSTKY 348
Y++ +F+ ++ + WG+ PN KY
Sbjct: 875 TYVFAMFEENQKQPEYEKFWGLFLPNKQLKY 967
>TC208247 similar to GB|BAB01763.1|9280308|AP000603 beta-1,3-glucanase-like
protein {Arabidopsis thaliana;} , partial (62%)
Length = 1563
Score = 123 bits (309), Expect = 1e-28
Identities = 76/196 (38%), Positives = 112/196 (56%), Gaps = 7/196 (3%)
Frame = +1
Query: 160 KISVSTSFSFVTAVTSPFPPSSAAFQEIPGVNLIGPLLQFLSDTNSSFLINLYPYNLYRL 219
++ VST S + + PFPPS+A F + I LLQFL +TNSS+++N YPY Y
Sbjct: 7 RVKVSTPQS-MDIIPKPFPPSTATFNSSWN-STIYQLLQFLKNTNSSYMLNAYPYYGYTK 180
Query: 220 RPEI-PLGIALFQEHPFNFRD-DFTTGVRYKNLFDIMVDAVVSAMALEGYETIPVIVTET 277
I P+ ALF+ P + D T Y ++FD MVDA ++ + IP++VTET
Sbjct: 181 GDGIFPIEYALFRPLPSVKQIVDPNTLFHYNSMFDAMVDATYYSIEALNFNNIPIVVTET 360
Query: 278 GWPSSG--SELDANSAYAEIYLKGLVKHLKSGAGTPLLKDGVKGVYLYELFDKE---GAG 332
GWPS G +E DA AE+Y+ +++ + + +G P + Y+YELF+++ G
Sbjct: 361 GWPSFGGANEPDATEENAELYINNMIQRVMNDSGPPSQPNIAINTYIYELFNEDKRNGPV 540
Query: 333 NGRNWGILYPNGSTKY 348
+ +NWGI Y NGST Y
Sbjct: 541 SEKNWGIFYTNGSTVY 588
>TC216276 homologue to UP|E13B_SOYBN (P52395) Glucan
endo-1,3-beta-glucosidase ((1->3)-beta-glucan
endohydrolase) ((1->3)-beta-glucanase)
(Beta-1,3-endoglucanase) (Fragment) , partial (95%)
Length = 1180
Score = 122 bits (307), Expect = 2e-28
Identities = 91/331 (27%), Positives = 159/331 (47%), Gaps = 6/331 (1%)
Frame = +3
Query: 15 SLYLLLSLLPTTTTSLPTIGVTYSSTTRQDSPPPPSPDRITTAMQNLKLTHLRLEEPDPS 74
+++LL+ +L + T + +IGV Y PS + + + +R+ PD
Sbjct: 48 AIFLLVGMLSSITVA-QSIGVCYGVI----GDNLPSRQEVVDLYKTNGIGRMRIYYPDEE 212
Query: 75 IIRSLLYTNISLFLTIPNYLVTPIATNRSIARAWIYTHVLPFYPRAKITTISVGNAFTDV 134
+++L + I L + + + + T+ + A W+ +V P+ I+VGN ++
Sbjct: 213 ALQALRGSGIELIMDVAKETLQSL-TDSNAATDWVNKYVTPYSQDVNFKYIAVGN---EI 380
Query: 135 YPES--INNLLPAISNVHISLRDLGIRKISVSTSFSFVTAVTSPFPPSSAAFQEIPGVNL 192
+P + +L A++N+ ++ ++ I VST+ T +T+ +PP+ F
Sbjct: 381 HPNTNEAQYILSAMTNIQNAISSANLQ-IKVSTAIDS-TLITNSYPPNDGVFTS-DAEPY 551
Query: 193 IGPLLQFLSDTNSSFLINLYPYNLYRLRPEIPLGIALFQEHPFNFRDDFTTGVRYKNLFD 252
I P++ FL + L N+YPY Y IPL ALF + N V Y+NLFD
Sbjct: 552 IKPIINFLVSNGAPLLANVYPYFAYANDQSIPLAYALFTQQGNN-------DVGYQNLFD 710
Query: 253 IMVDAVVSAMALEGYETIPVIVTETGWPSSGSELDANSAYAEIYLKGLVKHLKSGAGTPL 312
M+D++ +A+ G + ++V+E+GWPS G A+ A Y L++H SG GTP
Sbjct: 711 AMLDSIYAALEKVGASNLQIVVSESGWPSEGG-AGASIDNAGTYYANLIRHASSGNGTPK 887
Query: 313 LKDGVKGVYLYELFD---KEGAGNGRN-WGI 339
YL+ +FD K+GA R+ W +
Sbjct: 888 RPGESIETYLFAMFDENQKQGADTERHLWSL 980
>TC218883 weakly similar to UP|Q9MAQ2 (Q9MAQ2) CDS, partial (54%)
Length = 875
Score = 118 bits (296), Expect = 4e-27
Identities = 78/254 (30%), Positives = 136/254 (52%), Gaps = 3/254 (1%)
Frame = +1
Query: 17 YLLLSLLPTTTTSLPTIGVTYSSTTRQDSPPPPSPDRITTAMQNLKLTHLRLEEPDPSII 76
+++L + + ++G+ Y PS D ++++ T ++L + DP ++
Sbjct: 133 FIMLFITAAAIGLVSSLGINYGQIANN----LPSQDDAVALVKSIGATKVKLYDADPRVL 300
Query: 77 RSLLYTNISLFLTIPNYLVTPIATNRSIARAWIYTHVLPFYPRAKITTISVGNAFTDVYP 136
++ T + L + + N ++ + + A+AWI ++ P+ P KIT+I VGN
Sbjct: 301 KAFANTGVELMVGLGNEYLSRMKDPKQ-AQAWIKANLQPYLPATKITSIFVGNEVLTFND 477
Query: 137 ESI-NNLLPAISNVHISLRDLGI-RKISVSTSFSFVTAVTSPFPPSSAAFQEIPGVNLIG 194
S+ +NLLPA+ +VH +L +LG+ ++I+V+T+ S TS +PPS+ AF+ +
Sbjct: 478 TSLTSNLLPAMQSVHAALINLGLDKQITVTTTHSLAVLQTS-YPPSAGAFRP-DLAPCLA 651
Query: 195 PLLQFLSDTNSSFLINLYPYNLYRLRP-EIPLGIALFQEHPFNFRDDFTTGVRYKNLFDI 253
P+L F + T S FLIN YPY Y+ P ++PL LFQ P D ++ + Y N+
Sbjct: 652 PILSFQAKTGSPFLINAYPYFAYKANPKQVPLEYVLFQ--PNEGMVDPSSNLHYDNMLFA 825
Query: 254 MVDAVVSAMALEGY 267
+DAV SA+ GY
Sbjct: 826 QIDAVYSALDSLGY 867
>TC216277 UP|O49012 (O49012) Beta-1,3-glucanase 3 (Fragment) , complete
Length = 1087
Score = 118 bits (295), Expect = 5e-27
Identities = 83/307 (27%), Positives = 145/307 (47%), Gaps = 7/307 (2%)
Frame = +3
Query: 49 PSPDRITTAMQNLKLTHLRLEEPDPSIIRSLLYTNISLFLTIPNYLVTPIATNRSIARAW 108
PS + + + +R+ PD +++L + I L + + + + T+ + A W
Sbjct: 42 PSRQEVVDLYKTNGIGRMRIYYPDEEALQALRGSGIELIMDVAKETLQSM-TDPNAATDW 218
Query: 109 IYTHVLPFYPRAKITTISVGNAFTDVYPES--INNLLPAISNVHISLRDLGIRKISVSTS 166
+ +V + I+VGN +++P + +L A++N+ ++ ++ I VST+
Sbjct: 219 VNKYVTAYSQDVNFKYIAVGN---EIHPNTNEAQYILSAMTNIQNAISSANLQ-IKVSTA 386
Query: 167 FSFVTAVTSPFPPSSAAFQEIPGVNLIGPLLQFLSDTNSSFLINLYPYNLYR--LRPEIP 224
+PP+ A F + P++ FL + L N+YPY Y + IP
Sbjct: 387 IDSTFIAPPSYPPNDAVFTS-DAEPYVKPIIDFLVRNEAPLLANVYPYFAYANDQQNSIP 563
Query: 225 LGIALFQEHPFNFRDDFTTGVRYKNLFDIMVDAVVSAMALEGYETIPVIVTETGWPSSGS 284
L ALF + N Y+NLFD M+D++ +A+ G + ++V+E+GWPS G
Sbjct: 564 LAYALFTQQGNN-------DAGYQNLFDAMLDSIYAAVEKVGASNLQIVVSESGWPSEGG 722
Query: 285 ELDANSAYAEIYLKGLVKHLKSGAGTPLLKDGVKGVYLYELFD---KEGAGNGRNWGILY 341
A+ A Y L++H SG GTP YL+ +FD K+GA R++G+
Sbjct: 723 -AGASIDNAGTYYANLIRHASSGNGTPKRPGESIETYLFAMFDENQKQGADTDRHFGLFN 899
Query: 342 PNGSTKY 348
P+ S KY
Sbjct: 900 PDKSPKY 920
>TC212243 UP|O49011 (O49011) Beta-1,3-glucanase 1 (Fragment) , complete
Length = 741
Score = 107 bits (267), Expect = 9e-24
Identities = 71/237 (29%), Positives = 124/237 (51%), Gaps = 3/237 (1%)
Frame = +3
Query: 49 PSPDRITTAMQNLKLTHLRLEEPDPSIIRSLLYTNISLFLTIPNYLVTPIATNRSIARAW 108
P + + + ++ + +R+ PD + +++L + I L L + + + +ATN S A+ W
Sbjct: 42 PPANEVVSLYKSNDIMRMRIYNPDQAALQALGNSGIELILGVLHQDLQGLATNASTAQQW 221
Query: 109 IYTHVLPFYPRAKITTISVGNAFTDV--YPESINNLLPAISNVHISLRDLGIRKISVSTS 166
+ ++VL F+P KI + VGN V E +LPAI N++ ++R G++ + T+
Sbjct: 222 VQSNVLNFWPSVKIKHVVVGNEINPVGSSSEFAQYVLPAIQNIYQAIRAQGLQDLIKVTT 401
Query: 167 FSFVTAVTSPFPPSSAAFQEIPGVNLIGPLLQFLSDTNSSFLINLYPYNLYRLRP-EIPL 225
+T + + +PPS + F+ + + P++ +L N+ L N+ PY Y P +I L
Sbjct: 402 AIDMTLLGNSYPPSQSYFR-TDVRSYLDPIIGYLVYANAPLLANVLPYFSYSNNPIDISL 578
Query: 226 GIALFQEHPFNFRDDFTTGVRYKNLFDIMVDAVVSAMALEGYETIPVIVTETGWPSS 282
ALF D Y+NLFD M+DAV A+ G + V+V+E GWP+S
Sbjct: 579 SYALFNSTNVVVWDG---QYGYQNLFDAMLDAVHVAIDNTGIGYVEVVVSERGWPNS 740
>TC224124 similar to UP|Q8L9D9 (Q8L9D9) Beta-1,3-glucanase-like protein,
partial (38%)
Length = 532
Score = 94.0 bits (232), Expect = 1e-19
Identities = 56/157 (35%), Positives = 92/157 (57%), Gaps = 3/157 (1%)
Frame = +3
Query: 159 RKISVSTSFSFVTAVTSPFPPSSAAFQEIPGVNLIGPLLQFLSDTNSSFLINLYPYNLYR 218
+ I VS+ + ++A+ + +P S+ +F+ + P+L FL +T S ++N+YP+ Y
Sbjct: 39 KDIKVSSPIA-LSALANSYPSSAGSFRPELVEPVFKPMLDFLRETGSYLMVNVYPFFAYE 215
Query: 219 LRPE-IPLGIALFQEHPFNFRDDFTTGVRYKNLFDIMVDAVVSAMALEGYETIPVIVTET 277
+ I L ALF+++P D G+RY NLFD +DAV SA++ Y+ + ++VTET
Sbjct: 216 SNADVISLDYALFRDNPGVV--DPGNGLRYYNLFDAQIDAVFSALSALKYDDVKIVVTET 389
Query: 278 GWPSSG--SELDANSAYAEIYLKGLVKHLKSGAGTPL 312
GWPS G +E+ A+ A Y LV+ + + GTPL
Sbjct: 390 GWPSKGDSNEVGASVDNAAAYNGNLVRKILTAGGTPL 500
>TC233194 similar to GB|CAB79832.1|7270016|ATCHRIV74 1, 3-beta-glucanase-like
protein {Arabidopsis thaliana;} , partial (40%)
Length = 611
Score = 91.3 bits (225), Expect = 6e-19
Identities = 54/163 (33%), Positives = 87/163 (53%), Gaps = 5/163 (3%)
Frame = +2
Query: 191 NLIGPLLQFLSDTNSSFLINLYPYNLYRLRPEIPLGIALFQEHPFNFRDDFTTGVRYKNL 250
+L+ +++FLS N+ F +N+YP+ P P+ A F D+ G Y N+
Sbjct: 50 DLMVQIVKFLSQNNAPFTVNIYPFISLYSDPNFPVDYAFFNGFQSPISDN---GRIYDNV 220
Query: 251 FDIMVDAVVSAMALEGYETIPVIVTETGWPSSGSELDANSAYAEIYLKGLVKHLKSGAGT 310
FD D +V A+ G+ +P+IV E GWP+ G + +AN YA+ + +G + +G GT
Sbjct: 221 FDANHDTLVWALQKNGFGNMPIIVGEVGWPTDG-DRNANLQYAQRFNQGFMSRYIAGKGT 397
Query: 311 PLLKDGVKGVYLYELFDKE----GAGN-GRNWGILYPNGSTKY 348
P ++ G YL+ L D++ GN R+WG+ Y +G KY
Sbjct: 398 P-MRPGPMDAYLFSLIDEDFKSIQPGNFERHWGLFYYDGQPKY 523
>BE346589
Length = 478
Score = 89.4 bits (220), Expect = 2e-18
Identities = 57/135 (42%), Positives = 82/135 (60%), Gaps = 3/135 (2%)
Frame = +3
Query: 100 TNRSIARAWIYTHVLPFYPRAKITTISVGN-AFTDVYPESINNLLPAISNVHISLRDLGI 158
TN A+ WI HV P+ + KIT I+VGN F + + NLLPA+ VH +L +LG+
Sbjct: 45 TNPYKAQTWIQQHVQPYLSQTKITCITVGNEVFNSNDTQQMLNLLPAMQTVHDALVNLGL 224
Query: 159 -RKISVSTSFSFVTAVTSPFPPSSAAFQEIPGVNLIGPLLQFLSDTNSSFLINLYPYNLY 217
++++V+T+ SF +++ +PPSS AF+E V I LL F + NS FLIN YP+ Y
Sbjct: 225 DQQVTVTTAHSF-NILSNSYPPSSGAFRE-DLVQYIQALLDFHAQINSPFLINAYPFFAY 398
Query: 218 RLRP-EIPLGIALFQ 231
+ P E+ L LFQ
Sbjct: 399 KDNPDEVSLNYVLFQ 443
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.139 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,372,429
Number of Sequences: 63676
Number of extensions: 275510
Number of successful extensions: 3436
Number of sequences better than 10.0: 129
Number of HSP's better than 10.0 without gapping: 2905
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3293
length of query: 376
length of database: 12,639,632
effective HSP length: 99
effective length of query: 277
effective length of database: 6,335,708
effective search space: 1754991116
effective search space used: 1754991116
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 60 (27.7 bits)
Medicago: description of AC147434.3


